
Contents:
- Introduction
- Key Steps
- Importing Libraries
- Reading Input Data
- Exploratory Data Analysis (EDA)
- Above codes will help to give us information about it’s unique values and count of each value.
- Helps to plot a count plot which will help us to see count of values in each unique category.
- This plot will help to analyze how gender will affect chances of stroke.
- Returns number of unique values in this attribute
- This will plot a distribution plot of variable age
- Above code will plot a boxplot of variable age with respect of target attribute stroke
- Above codes will help to give us information about it’s unique values and count of each value.
- Helps to plot a count plot which will help us to see count of values in each unique category.
- This plot will help to analyze how gender will affect chances of stroke.
- Above code will return unique value for heart disease attribute and its value counts
- Will plot a counter plot of variable heart diseases
- This plot will help to analyze how gender will affect chances of stroke.
- Above code will show us number unique values of attribute and its count
- Counter plot of ever married attribute
- Ever married with respect to stroke
- Above code will return unique values of attributes and its count
- Above code will create a count plot
- Above code will create a count plot with respect to stroke
- Above code will return unique values of variable and its count
- This will create a counter plot
- Residence Type with respect to stroke
- Number of unique values
- Distribution of avg_glucose_level
- Avg_glucose_level and Stroke
- Filling null values with average value
- Returns number of unique values in that attribute
- Distribution of BMI
- BMI with respect to Stroke
- Returns unique values and its count
- Returns unique values and its count
- Smoking Status with respect to Stroke
- Count Plot of Stroke
- Count Plot of Stroke
- Feature Engineering
- Data Preparation
- Model Building, Testing and X-Validation
- Model Predictions and Performance QC
- TF Performance Analysis
- Binary Classification Summary Report
- Conclusions
- References
Introduction
Stroke, also known as brain attack, happens when blood flow to the brain is blocked, preventing it from getting oxygen and nutrients from it and causing the death of brain cells within minutes. According to the World Health Organization (WHO) stroke is the 2nd leading cause of death globally after ischemic heart disease, responsible for approximately 11% of total deaths. Stroke victims can experience paralysis, impaired speed, or loss of vision. While some of the Stroke risk factors cannot be modified, such as family history of cerebrovascular diseases, age, gender and race, others can and are estimated to account for 60% -80% of stroke risk in the general population.
Therefore, predicting stroke outcome for new cases can be determining for them to be treated early enough and avoid disabling and mortal consequences.
The use of AI-powered Machine Learning (ML) techniques in medical records have great impact on the fields of healthcare and bio-medicine [1-4]. This assists the medical practitioners to identify the onset of disease at an earlier stage. Here, we are particularly interested in stroke, and to identify the key factors that are associated with its occurrence.
In this blog, we formulate the ML stroke prediction process as a binary classification problem and provide a detailed understanding of the various risk factors for stroke prediction. Specifically, we analyse various features present in Electronic Health Record (EHR) records of patients, and identify the most important factors necessary for stroke prediction [1,2]. In doing so, we use dimensionality reduction techniques to identify patterns in low-dimension subspace of the feature space [1-4]. We benchmark popular ML/AI binary classification models for stroke prediction in a publicly available dataset.
Key Steps
Our ML stroke prediction workflow [1-3] in Python consists of the 7 steps
that implement ETL Preparation + ML Execution + Performance QC + Prediction:
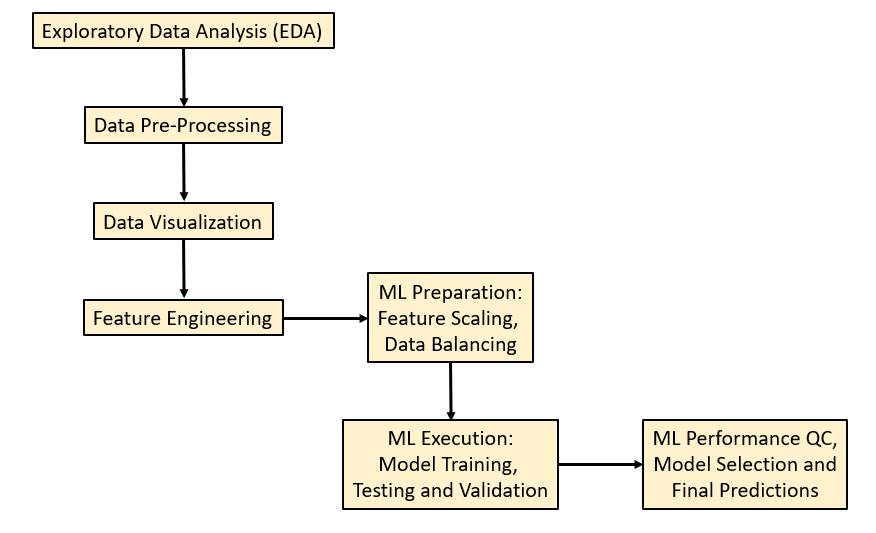
This boils down to the following sequence:
- Install the Anaconda IDE with the Jupyter notebook and Python 3.9.
- Install and import the required Python ML libraries.
- Read input EMR Kaggle dataset healthcare-dataset-stroke-data.csv
- Input data preparation, editing, transformation and visualization
- Feature engineering, correlations and impact factors
- Train/test data splitting (test_size=0.25) and SMOTE balancing
- Binary classification model training, ML benchmarking, testing and X-validation
- Scikit-Plot: Visualizing ML Algorithm Results & Performance
- Output the final classification report, the best training model and predictions
Importing Libraries
Let’s begin with importing/installing the required libraries:
!pip install imblearn
!pip install xgboost
!pip install tensorflow
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, f1_score,classification_report,precision_score,recall_score
from imblearn.over_sampling import SMOTE/SMOTENC
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from xgboost import XGBClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
import tensorflow as tf
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import cross_val_score
Reading Input Data
data=pd.read_csv(‘YOURPATH/healthcare-dataset-stroke-data.csv’)
df=data
data.head(10)
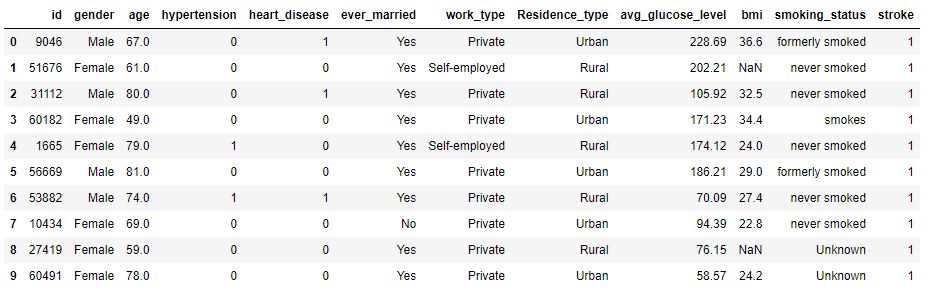
Displaying top 10 rows
data.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 5110 entries, 0 to 5109 Data columns (total 12 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 id 5110 non-null int64 1 gender 5110 non-null object 2 age 5110 non-null float64 3 hypertension 5110 non-null int64 4 heart_disease 5110 non-null int64 5 ever_married 5110 non-null object 6 work_type 5110 non-null object 7 Residence_type 5110 non-null object 8 avg_glucose_level 5110 non-null float64 9 bmi 4909 non-null float64 10 smoking_status 5110 non-null object 11 stroke 5110 non-null int64 dtypes: float64(3), int64(4), object(5) memory usage: 479.2+ KB
Showing information about dataset
data.describe()
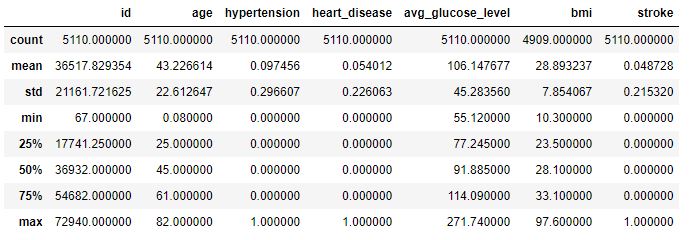
Showing data’s statistical features.
We can print the table out as follows:
print(stroke_df.shape)
print(stroke_df.head(10))
print(stroke_df.describe().T)
(5110, 12)
id gender age hypertension heart_disease ever_married \
0 9046 Male 67.0 0 1 Yes
1 51676 Female 61.0 0 0 Yes
2 31112 Male 80.0 0 1 Yes
3 60182 Female 49.0 0 0 Yes
4 1665 Female 79.0 1 0 Yes
5 56669 Male 81.0 0 0 Yes
6 53882 Male 74.0 1 1 Yes
7 10434 Female 69.0 0 0 No
8 27419 Female 59.0 0 0 Yes
9 60491 Female 78.0 0 0 Yes
work_type Residence_type avg_glucose_level bmi smoking_status \
0 Private Urban 228.69 36.6 formerly smoked
1 Self-employed Rural 202.21 NaN never smoked
2 Private Rural 105.92 32.5 never smoked
3 Private Urban 171.23 34.4 smokes
4 Self-employed Rural 174.12 24.0 never smoked
5 Private Urban 186.21 29.0 formerly smoked
6 Private Rural 70.09 27.4 never smoked
7 Private Urban 94.39 22.8 never smoked
8 Private Rural 76.15 NaN Unknown
9 Private Urban 58.57 24.2 Unknown
stroke
0 1
1 1
2 1
3 1
4 1
5 1
6 1
7 1
8 1
9 1
count mean std min 25% \
id 5110.0 36517.829354 21161.721625 67.00 17741.250
age 5110.0 43.226614 22.612647 0.08 25.000
hypertension 5110.0 0.097456 0.296607 0.00 0.000
heart_disease 5110.0 0.054012 0.226063 0.00 0.000
avg_glucose_level 5110.0 106.147677 45.283560 55.12 77.245
bmi 4909.0 28.893237 7.854067 10.30 23.500
stroke 5110.0 0.048728 0.215320 0.00 0.000
50% 75% max
id 36932.000 54682.00 72940.00
age 45.000 61.00 82.00
hypertension 0.000 0.00 1.00
heart_disease 0.000 0.00 1.00
avg_glucose_level 91.885 114.09 271.74
bmi 28.100 33.10 97.60
stroke 0.000 0.00 1.00
As we can see, the Kaggle stroke dataset has 11 variables representing clinical features for predicting stroke events plus 1 record ID (columns) and 5110 observations (rows). This dataset is used to predict whether a patient is likely to get stroke based on the input parameters like gender, age, various diseases, and smoking status. Each row in the data provides relavant information about the patient.
Exploratory Data Analysis (EDA)
The Gender attribute states the gender of the patient. Let’s see how does Gender affects and Gender wise comparison of stroke rate.
print(‘Unique values\n’,data[‘gender’].unique())
print(‘Value Counts\n’,data[‘gender’].value_counts())
Above codes will help to give us information about it’s unique values and count of each value.
sns.countplot(data=data,x=’gender’)
Helps to plot a count plot which will help us to see count of values in each unique category.
sns.countplot(data=data,x=’gender’,hue=’stroke’)
This plot will help to analyze how gender will affect chances of stroke.
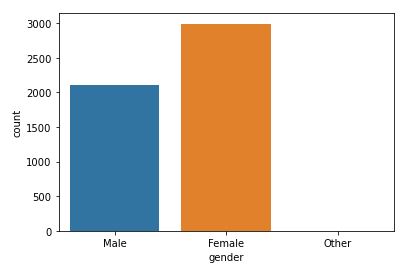
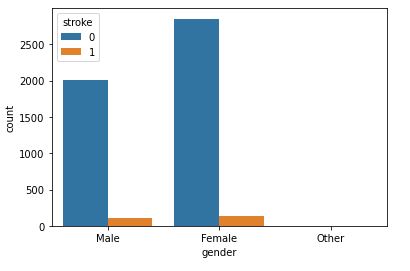
The dataset appears to be imbalanced. As we can there is not much difference between stroke rate concerning gender.
Let’s look at the Age attribute.
data[‘age’].nunique()
Returns number of unique values in this attribute
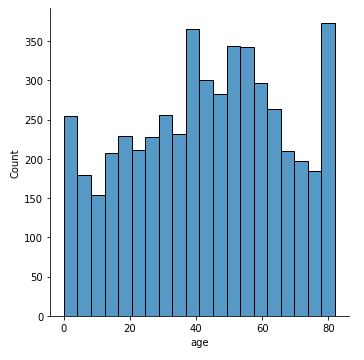
sns.displot(data[‘age’])
This will plot a distribution plot of variable age
plt.figure(figsize=(15,7))
sns.boxplot(data=data,x=’stroke’,y=’age’)
Above code will plot a boxplot of variable age with respect of target attribute stroke
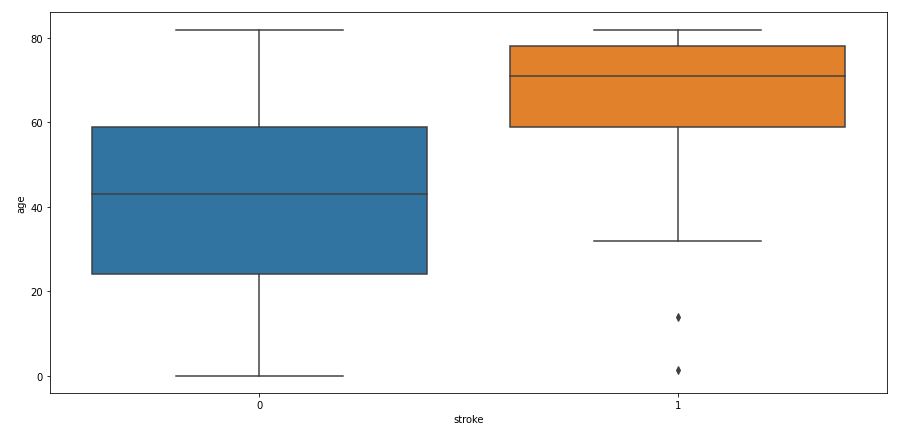
One can see that 60+ seniors tend to have a high risk of stroke. Some outliers can be seen for young people who suffer from stroke.
Since hypertension (high blood pressure) might lead to stroke, we consider the Hypertension attribute as well.
print(‘Unique values\n’,data[‘hypertension’].unique())
print(‘Value Counts\n’,data[‘hypertension’].value_counts())
Above codes will help to give us information about it’s unique values and count of each value.
sns.countplot(data=data,x=’hypertension’)
Helps to plot a count plot which will help us to see count of values in each unique category.
sns.countplot(data=data,x=’hypertension’,hue=’stroke’)
This plot will help to analyze how gender will affect chances of stroke.
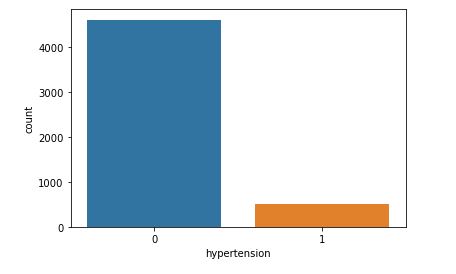
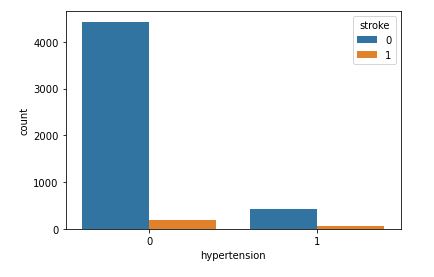
Even though hypertension can cause a stroke, our dataset does not contain a large amount clinical records of patients with high blood pressure.
Let’s look at heart disease associated with a higher risk of stroke.
print(‘Unique Value\n’,data[‘heart_disease’].unique())
print(‘Value Counts\n’,data[‘heart_disease’].value_counts())
Above code will return unique value for heart disease attribute and its value counts
sns.countplot(data=data,x=’heart_disease’)
Will plot a counter plot of variable heart diseases
sns.countplot(data=data,x=’heart_disease’,hue=’stroke’)
This plot will help to analyze how gender will affect chances of stroke.
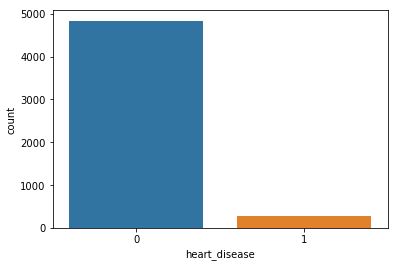
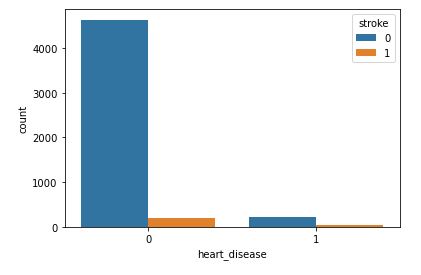
As with hypertension, it’s rather difficult to estimate the impact of heart desease because the number of recorded stroke patients with heart desease is relatively small.
The “ever_married” attribute will tell us whether or not the patient was ever married.
print(‘Unique Values\n’,data[‘ever_married’].unique())
print(‘Value Counts\n’,data[‘ever_married’].value_counts())
Above code will show us number unique values of attribute and its count
sns.countplot(data=data,x=’ever_married’)
Counter plot of ever married attribute
sns.countplot(data=data,x=’ever_married’,hue=’stroke’)
Ever married with respect to stroke
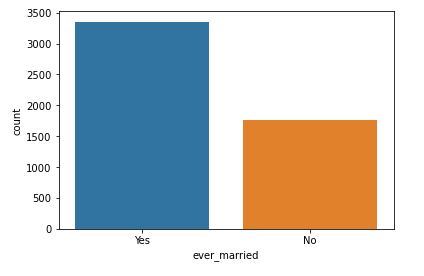
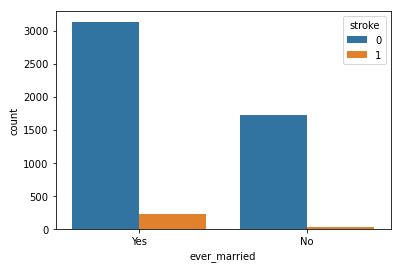
It is clear that people who are married have a higher stroke rate.
Let’s check the relationship between working conditions and the risk of stroke.
print(‘Unique Value\n’,data[‘work_type’].unique())
print(‘Value Counts\n’,data[‘work_type’].value_counts())
Above code will return unique values of attributes and its count
sns.countplot(data=data,x=’work_type’)
Above code will create a count plot
sns.countplot(data=data,x=’work_type’,hue=’stroke’)
Above code will create a count plot with respect to stroke
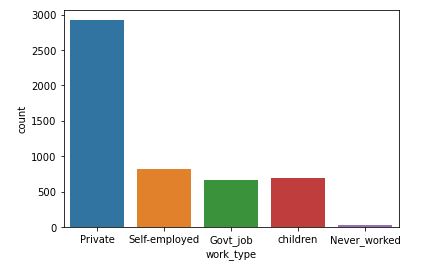
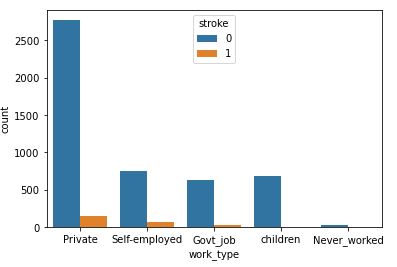
People working in the Private sector have a highest risk of stroke.
Let’s look at the residence type of stroke patients (urban or rural).
print(‘Unique Values\n’,data[‘Residence_type’].unique())
print(“Value Counts\n”,data[‘Residence_type’].value_counts())
Above code will return unique values of variable and its count
sns.countplot(data=data,x=’Residence_type’)
This will create a counter plot
sns.countplot(data=data,x=’Residence_type’,hue=’stroke’)
Residence Type with respect to stroke
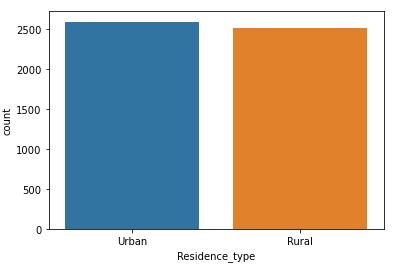
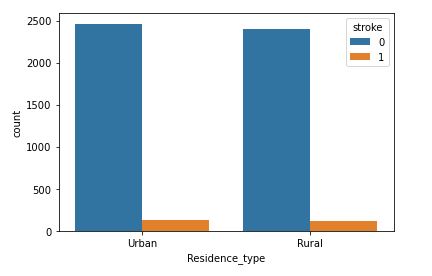
This attribute is of no use for stroke prediction. It would be best to discard it.
Let’s examine the average glucose level of stroke patients.
data[‘avg_glucose_level’].nunique()
Number of unique values
sns.displot(data[‘avg_glucose_level’])
Distribution of avg_glucose_level
sns.boxplot(data=data,x=’stroke’,y=’avg_glucose_level’)
Avg_glucose_level and Stroke
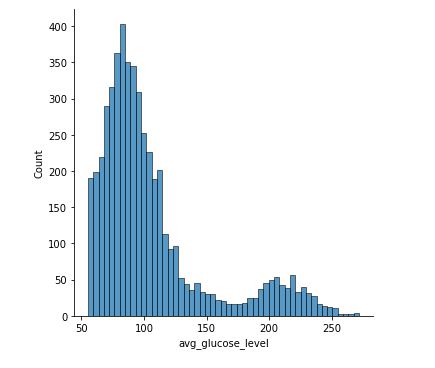
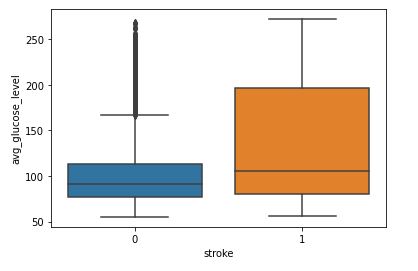
We can see that people having stroke have an average glucose level of more than 100.
Body Mass Index (BMI) is a measure of body fat based on height and weight of adult men and women. Let’s see how does it affect the chances of having a stroke.
data[‘bmi’].fillna(data[‘bmi’].mean(),inplace=True)
Filling null values with average value
data[‘bmi’].nunique()
Returns number of unique values in that attribute
sns.displot(data[‘bmi’])
Distribution of BMI
sns.boxplot(data=data,x=’stroke’,y=’bmi’)
BMI with respect to Stroke
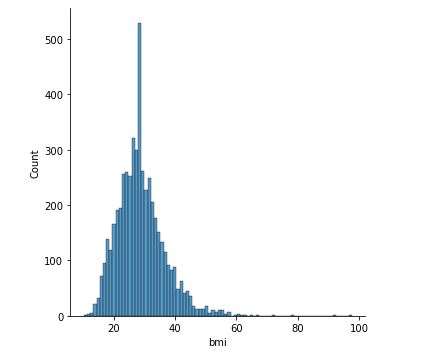
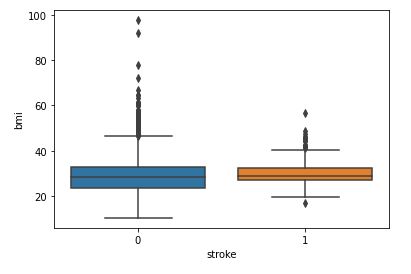
The relationship between BMI and stroke is unclear at this stage due to the overlap of two box plots.
Let’s check the impact of smoking.
print(‘Unique Values\n’,data[‘smoking_status’].unique())
print(‘Value Counts\n’,data[‘smoking_status’].value_counts())
Returns unique values and its count
Returns unique values and its count
sns.countplot(data=data,x=’smoking_status’)
sns.countplot(data=data,x=’smoking_status’,hue=’stroke’)
Smoking Status with respect to Stroke
print(‘Unique Value\n’,data[‘stroke’].unique())
print(‘Value Counts\n’,data[‘stroke’].value_counts())
sns.countplot(data=data,x=’stroke’)
Count Plot of Stroke
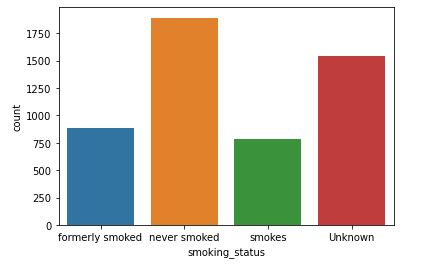
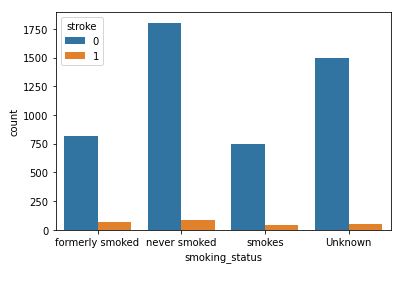
As with BMI, the correlation between the smoking status and the risk of stroke is unclear at this stage.
Finally, let’ plot our target binary variable Stroke.
print(‘Unique Value\n’,data[‘stroke’].unique())
print(‘Value Counts\n’,data[‘stroke’].value_counts())
sns.countplot(data=data,x=’stroke’)
Count Plot of Stroke
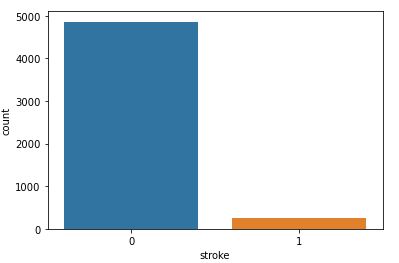
Here is the summary of attribute unique values:
- Gender
Unique values ['Male' 'Female' 'Other'] Value Counts Female 2994 Male 2115 Other 1
- Age
Number of Unique Values:
104
- Hypertension
Value Count [0 1] Value Counts 0 4612 1 498
Unique Values and Value Counts:
- Heart Disease
Unique Value [1 0] Value Counts 0 4834 1 276
- Ever Married
Unique Values ['Yes' 'No'] Value Counts Yes 3353 No 1757
- Work Type
Unique Value ['Private' 'Self-employed' 'Govt_job' 'children' 'Never_worked'] Value Counts Private 2925 Self-employed 819 children 687 Govt_job 657 Never_worked 22
- Residence Type
Unique Values ['Urban' 'Rural'] Value Counts Urban 2596 Rural 2514
- Average Glucose Level
Unique Values and Count:
3979
- BMI
Null Values:
201
Unique Values and Counts:
419
- Smoking Status
Unique Values ['formerly smoked' 'never smoked' 'smokes' 'Unknown'] Value Counts never smoked 1892 Unknown 1544 formerly smoked 885 smokes 789
- Stroke
Unique Value [1 0] Value Counts 0 4861 1 249
Feature Engineering
Let’s encode our categorical data into numeric ones using Label Encoder.
cols=data.select_dtypes(include=[‘object’]).columns
print(cols)
This code will fetch columns whose data type is object.
le=LabelEncoder()
Initializing our Label Encoder object
data[cols]=data[cols].apply(le.fit_transform)
Transfering categorical data into numeric
print(data.head(10))
Index(['gender', 'ever_married', 'work_type', 'Residence_type',
'smoking_status'],
dtype='object')
gender age hypertension heart_disease ever_married work_type \
0 1 67.0 0 1 1 2
1 0 61.0 0 0 1 3
2 1 80.0 0 1 1 2
3 0 49.0 0 0 1 2
4 0 79.0 1 0 1 3
5 1 81.0 0 0 1 2
6 1 74.0 1 1 1 2
7 0 69.0 0 0 0 2
8 0 59.0 0 0 1 2
9 0 78.0 0 0 1 2
Residence_type avg_glucose_level bmi smoking_status stroke
0 1 228.69 36.600000 1 1
1 0 202.21 28.893237 2 1
2 0 105.92 32.500000 2 1
3 1 171.23 34.400000 3 1
4 0 174.12 24.000000 2 1
5 1 186.21 29.000000 1 1
6 0 70.09 27.400000 2 1
7 1 94.39 22.800000 2 1
8 0 76.15 28.893237 0 1
9 1 58.57 24.200000 0 1
Let’s compute the feature correlation matrix
plt.figure(figsize=(15,10))
sns.heatmap(data.corr(),annot=True,fmt=’.2′)
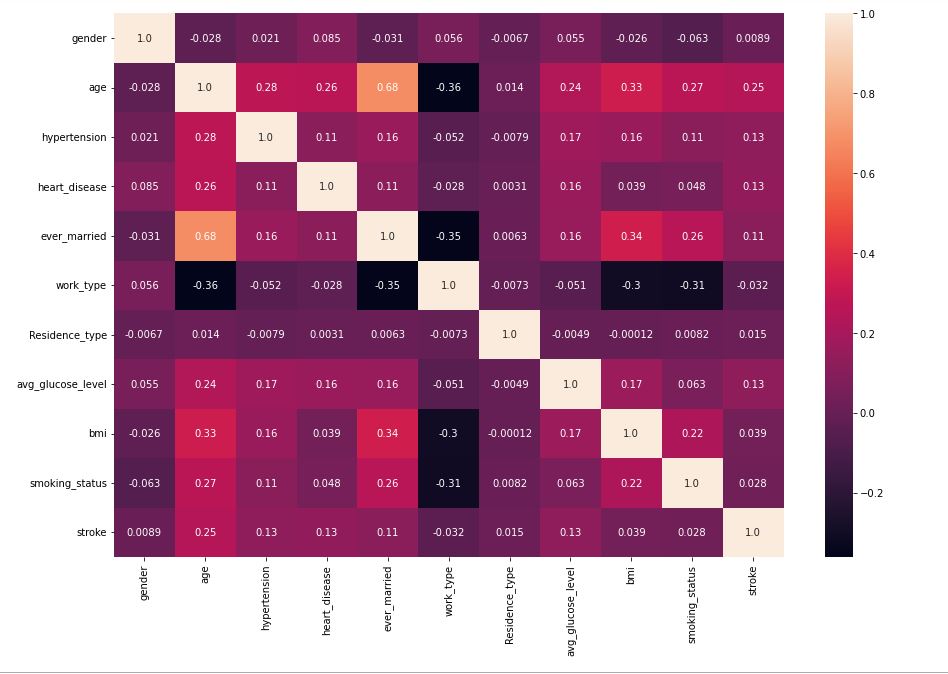
Variables that exhibit weak correlations are:
age, hypertension, heart_disease, ever_married, and avg_glucose_level.
Let’s double check our observations using SelectKBest and F_Classif.
classifier = SelectKBest(score_func=f_classif,k=5)
fits = classifier.fit(data.drop(‘stroke’,axis=1),data[‘stroke’])
x=pd.DataFrame(fits.scores_)
columns = pd.DataFrame(data.drop(‘stroke’,axis=1).columns)
fscores = pd.concat([columns,x],axis=1)
fscores.columns = [‘Attribute’,’Score’]
fscores.sort_values(by=’Score’,ascending=False)
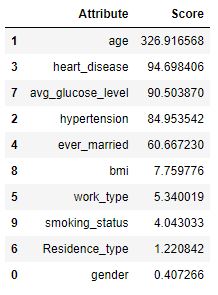
import matplotlib.pyplot as plt
fscores.columns = [‘Attribute’,’Score’]
fscores.sort_values(by=’Score’,ascending=False)
naming the x and y axis
f = plt.figure()
f.set_figwidth(16)
f.set_figheight(4)
plt.xlabel(‘Features’)
plt.ylabel(‘ML/AI Importance Factor’)
plt.plot(fscores[‘Attribute’],fscores[‘Score’])
plt.show()
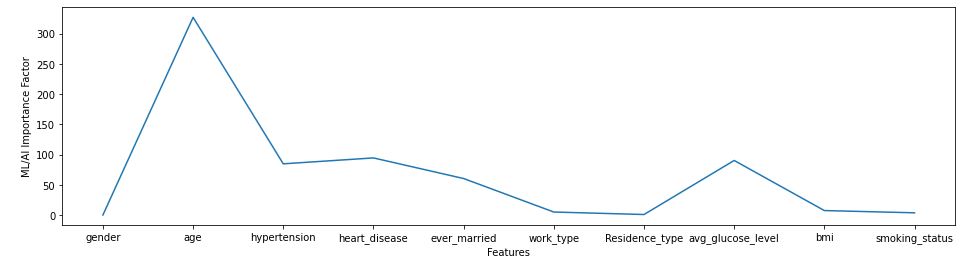
cols=fscores[fscores[‘Score’]>50][‘Attribute’]
print(cols)
1 age 2 hypertension 3 heart_disease 4 ever_married 7 avg_glucose_level Name: Attribute, dtype: object
Data Preparation
ID is nothing but a unique number assigned to every patient to keep track of them and making them unique. So we can remove it.
data.drop("id",inplace=True,axis=1)
I can see there are 201 NaN values for BMI variable. I decide to impute values for BMI since there are 40 patients who had a stroke that have a missing BMI value. The median was chosen over the mean since BMI has a long right tail.
stroke_df.fillna(stroke_df.median(), inplace=True)
stroke_df
The categorical data set is then transformed prior to processing through algorithms. Features with only 2 values were transformed into 1s and 0s using class mapping: ever_married, Residence_type, gender. Features with more than 2 values were transformed using One-Hot Label Encoding with the built-in Pandas get_dummies function: work_type, smoking_status.
# transform nominal variables that only have 2 values
class_mapping = {label: idx for idx, label in enumerate(np.unique(stroke_df['ever_married']))}
print(class_mapping)stroke_df['ever_married'] = stroke_df['ever_married'].map(class_mapping)
class_mapping = {label: idx for idx, label in enumerate(np.unique(stroke_df['Residence_type']))}
print(class_mapping)stroke_df['Residence_type'] = stroke_df['Residence_type'].map(class_mapping)
class_mapping = {label: idx for idx, label in enumerate(np.unique(stroke_df['gender']))}
print(class_mapping)stroke_df['gender'] = stroke_df['gender'].map(class_mapping)# transform nominal variables that have more than 2 values
stroke_df[['work_type','smoking_status']] = stroke_df[['work_type','smoking_status']].astype(str)# concatenate the nominal variables from pd.getdummies and the ordinal variables to form the final datasettranspose = pd.get_dummies(stroke_df[['work_type','smoking_status']])
stroke_dummies_df = pd.concat([stroke_df,transpose],axis=1)[['id','age','hypertension','heart_disease','ever_married','Residence_type','avg_glucose_level','bmi','gender','work_type_Govt_job','work_type_Never_worked','work_type_Private','work_type_Self-employed','work_type_children','smoking_status_Unknown','smoking_status_formerly smoked','smoking_status_never smoked','smoking_status_smokes','stroke']]
stroke_dummies_df
Now, let’s split features into training and testing sets for training and testing our classification models.
train_x,test_x,train_y,test_y=train_test_split(data[cols],data['stroke'],random_state=1255,test_size=0.25) #Splitting data train_x.shape,test_x.shape,train_y.shape,test_y.shape # Shape of data ((3832, 5), (1278, 5), (3832,), (1278,))
Let’s balance our training and test data. We are going to use the SMOTE method
smote=SMOTE()
train_x,train_y=smote.fit_resample(train_x,train_y)
test_x,test_y=smote.fit_resample(test_x,test_y)
print(train_x.shape,train_y.shape,test_x.shape,test_y.shape)
(7296, 5) (7296,) (2426, 5) (2426,)
Model Building, Testing and X-Validation
Let’s models using Logistic Regression, Random Forest Classifier, SVM, and XGBClassifier. We select the best model performer XGBClassifier while comparing other ML algorithms.
xgc=XGBClassifier(objective=’binary:logistic’,n_estimators=100000,max_depth=5,learning_rate=0.001,n_jobs=-1)
xgc.fit(train_x,train_y)
predict=xgc.predict(test_x)
print(‘Accuracy –> ‘,accuracy_score(predict,test_y))
print(‘F1 Score –> ‘,f1_score(predict,test_y))
print(‘Classification Report –> \n’,classification_report(predict,test_y))
Accuracy --> 0.9117889530090684
F1 Score --> 0.9066317626527051
Classification Report -->
precision recall f1-score support
0 0.97 0.87 0.92 1347
1 0.86 0.96 0.91 1079
accuracy 0.91 2426
macro avg 0.91 0.92 0.91 2426
weighted avg 0.92 0.91 0.91 2426
Model Predictions and Performance QC
Let’s examine the model performance using scikit-plot
!pip install scikit-plot
import scikitplot as skplt
mport sklearn
from sklearn.datasets import load_digits, load_boston, load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor, GradientBoostingClassifier, ExtraTreesClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import sys
import warnings
warnings.filterwarnings(“ignore”)
print(“Scikit Plot Version : “, skplt.version)
print(“Scikit Learn Version : “, sklearn.version)
print(“Python Version : “, sys.version)
%matplotlib inline
Scikit Plot Version : 0.3.7 Scikit Learn Version : 1.0.2 Python Version : 3.9.12 (main, Apr 4 2022, 05:22:27) [MSC v.1916 64 bit (AMD64)]
Let’s plot the LogisticRegression learning curve
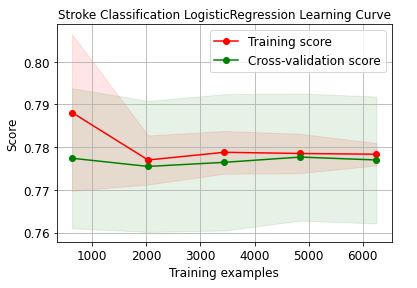
Let’s plot the RandomForest learning curve
skplt.estimators.plot_learning_curve(RandomForestClassifier(),train_x, train_y,
cv=7, shuffle=True, scoring=”r2″, n_jobs=-1,
figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”Stroke Classification RandomForest Learning Curve”);
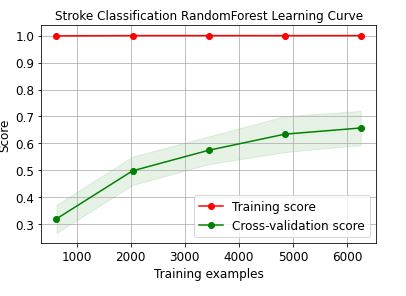
Let’s plot the XGBClassifier learning curve
skplt.estimators.plot_learning_curve(XGBClassifier(),train_x, train_y,
cv=7, shuffle=True, scoring=”r2″, n_jobs=-1,
figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”Stroke Classification XGBClassifier Learning Curve”);
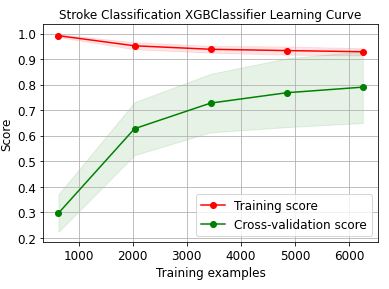
Let’s plot the GradientBoosting learning curve
skplt.estimators.plot_learning_curve(GradientBoostingClassifier(),train_x, train_y,
cv=7, shuffle=True, scoring=”r2″, n_jobs=-1,
figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”Stroke Classification GradientBoostingClassifier Learning Curve”);
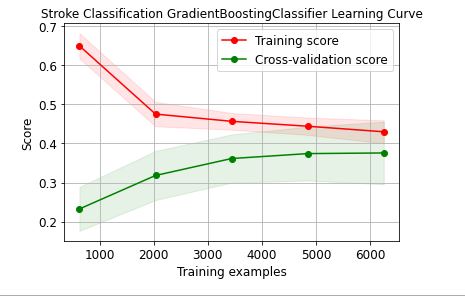
Let’s plot the ExtraTrees learning curve
skplt.estimators.plot_learning_curve(ExtraTreesClassifier(),train_x, train_y,
cv=7, shuffle=True, scoring=”r2″, n_jobs=-1,
figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”Stroke Classification ExtraTreesClassifier Learning Curve”);
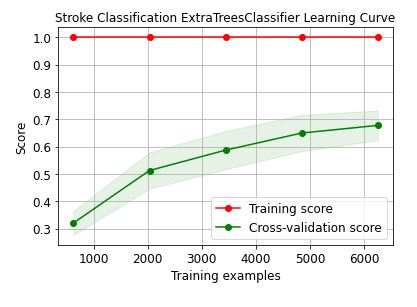
These learning curves confirm that the XGBClassifier is the best performer in terms of the training/X-validation scores and confidence intervals.
TF Performance Analysis
Let’s continue the GradientBoostingClassifier prediction QC analysis by invoking the TensorFlow library.
!pip install tensorflow
import tensorflow as tf
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import cross_val_score
Y_test_probs = xgc.predict_proba(test_x)
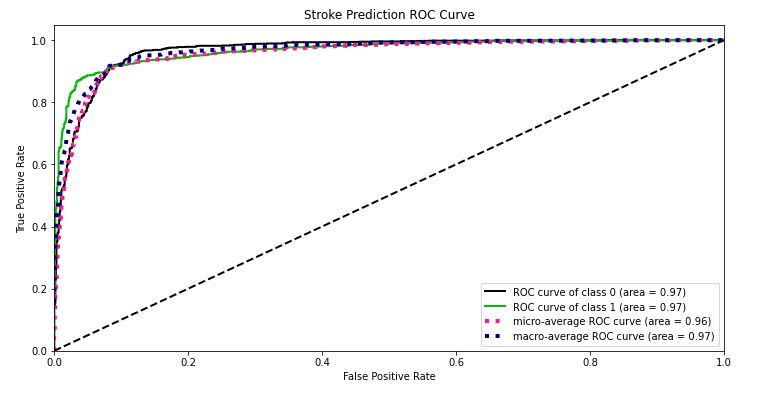
skplt.metrics.plot_roc_curve(test_y, Y_test_probs,
title=”Stroke Prediction ROC Curve”, figsize=(12,6));
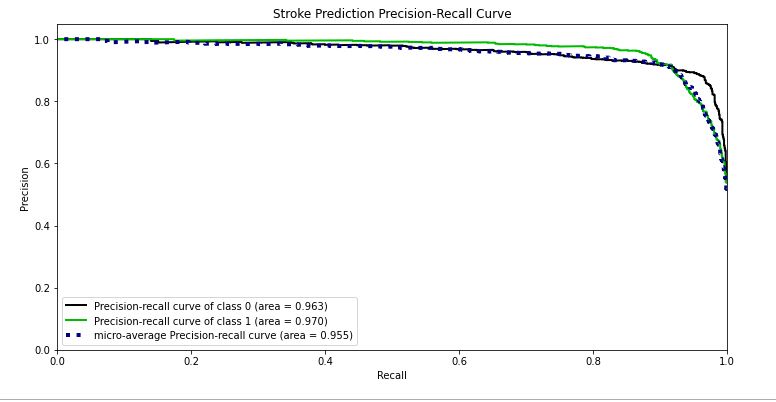
skplt.metrics.plot_precision_recall_curve(test_y, Y_test_probs,
title=”Stroke Prediction Precision-Recall Curve”, figsize=(12,6));
kmeans = KMeans(n_clusters=10, random_state=1)
kmeans.fit(train_x, train_y)
cluster_labels = kmeans.predict(test_x)
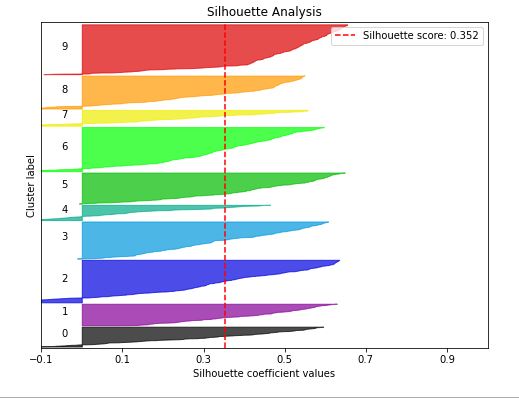
skplt.metrics.plot_silhouette(test_x, cluster_labels,
figsize=(8,6));
lr_probas = LogisticRegression().fit(train_x, train_y).predict_proba(test_x)
rf_probas = RandomForestClassifier().fit(train_x, train_y).predict_proba(test_x)
gb_probas = GradientBoostingClassifier().fit(train_x, train_y).predict_proba(test_x)
et_scores = ExtraTreesClassifier().fit(train_x, train_y).predict_proba(test_x)
probas_list = [lr_probas, rf_probas, gb_probas, et_scores]
clf_names = [‘Logistic Regression’, ‘Random Forest’, ‘Gradient Boosting’, ‘Extra Trees Classifier’]
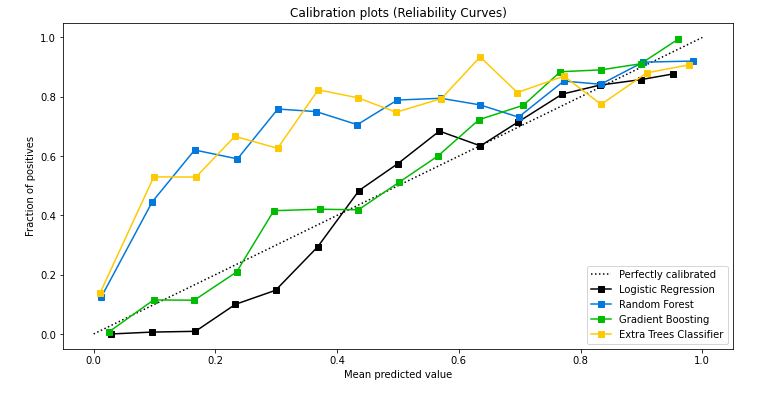
skplt.metrics.plot_calibration_curve(test_y,
probas_list,
clf_names, n_bins=15,
figsize=(12,6)
);
rf = GradientBoostingClassifier()
rf.fit(train_x, train_y)
Y_cancer_probas = rf.predict_proba(test_x)
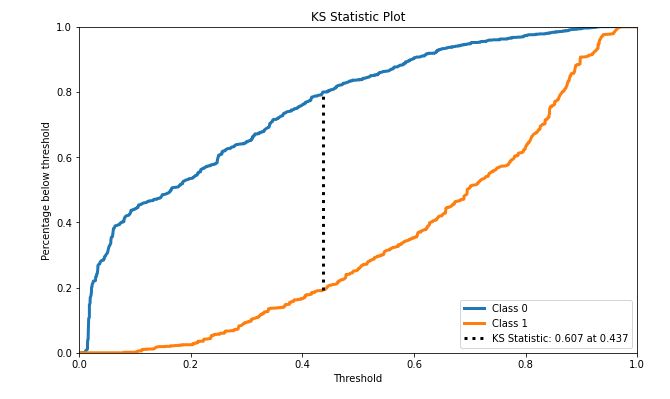
skplt.metrics.plot_ks_statistic(test_y, Y_cancer_probas, figsize=(10,6));
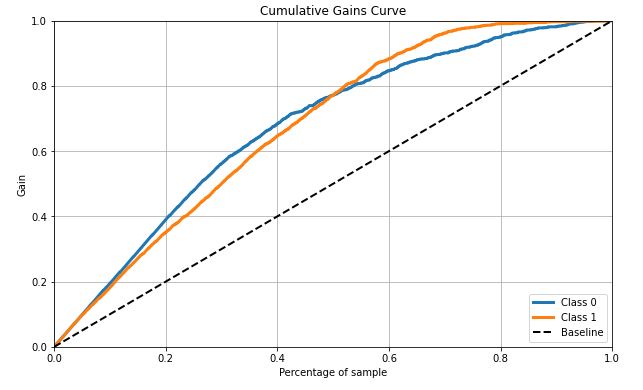
skplt.metrics.plot_cumulative_gain(test_y, Y_cancer_probas, figsize=(10,6));
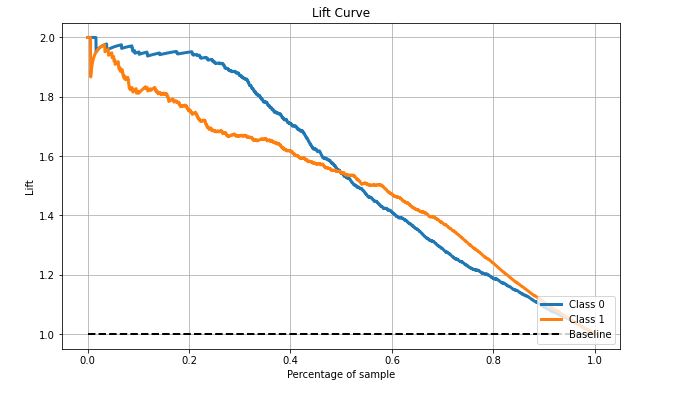
skplt.metrics.plot_lift_curve(test_y, Y_cancer_probas, figsize=(10,6));
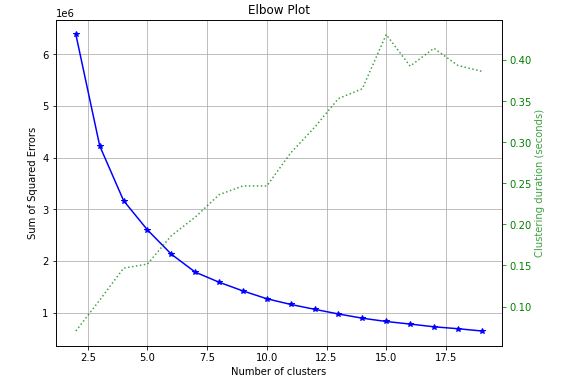
skplt.cluster.plot_elbow_curve(KMeans(random_state=1),
train_x,
cluster_ranges=range(2, 20),
figsize=(8,6));
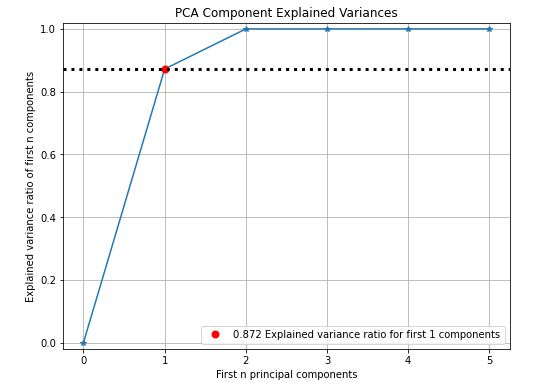
pca = PCA(random_state=1)
pca.fit(train_x)
skplt.decomposition.plot_pca_component_variance(pca, figsize=(8,6));
skplt.decomposition.plot_pca_2d_projection(pca, train_x, train_y,
figsize=(10,10),
cmap=”tab10″);
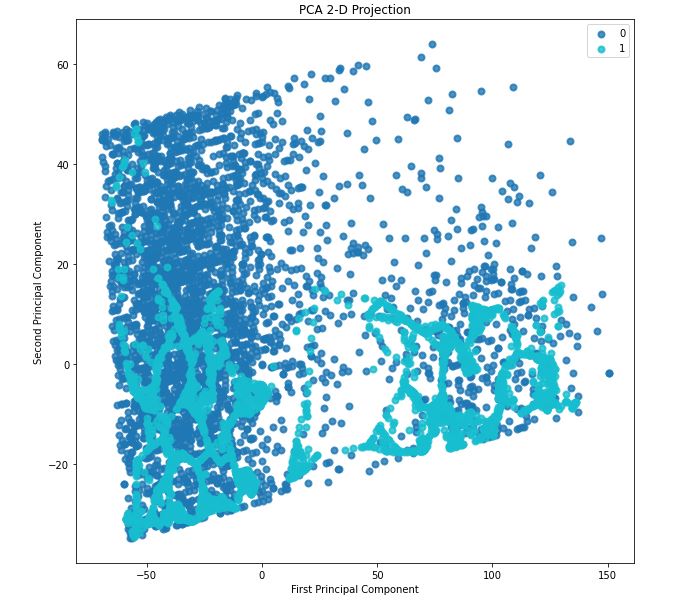
We can see the data is non-linear and also some interesting clustering. This suggests a high-dimension classification model or clustering model will work nicely.
Binary Classification Summary Report
Let’s summarize our best prediction classification results
xgc=XGBClassifier(objective=’binary:logistic’,n_estimators=100000,max_depth=5,learning_rate=0.001,n_jobs=-1)
xgc.fit(train_x,train_y)
predict=xgc.predict(test_x)
print(‘Accuracy –> ‘,accuracy_score(predict,test_y))
print(‘F1 Score –> ‘,f1_score(predict,test_y))
print(‘Classification Report –> \n’,classification_report(predict,test_y))
Accuracy --> 0.9117889530090684
F1 Score --> 0.9066317626527051
Classification Report -->
precision recall f1-score support
0 0.97 0.87 0.92 1347
1 0.86 0.96 0.91 1079
accuracy 0.91 2426
macro avg 0.91 0.92 0.91 2426
weighted avg 0.92 0.91 0.91 2426
skplt.metrics.plot_confusion_matrix(test_y, predict, normalize=True)
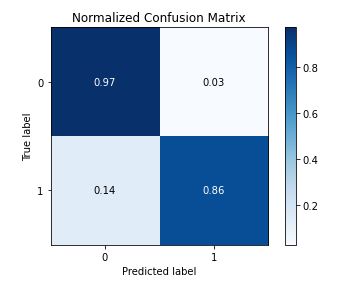
gc=GradientBoostingClassifier(n_estimators=100000,max_depth=5,learning_rate=0.001)
gc.fit(train_x,train_y)
GradientBoostingClassifier(learning_rate=0.001, max_depth=5,
n_estimators=100000)
predict=xgc.predict(test_x)
print(‘Accuracy –> ‘,accuracy_score(predict,test_y))
print(‘F1 Score –> ‘,f1_score(predict,test_y))
print(‘Classification Report –> \n’,classification_report(predict,test_y))
Accuracy --> 0.9117889530090684
F1 Score --> 0.9066317626527051
Classification Report -->
precision recall f1-score support
0 0.97 0.87 0.92 1347
1 0.86 0.96 0.91 1079
accuracy 0.91 2426
macro avg 0.91 0.92 0.91 2426
weighted avg 0.92 0.91 0.91 2426
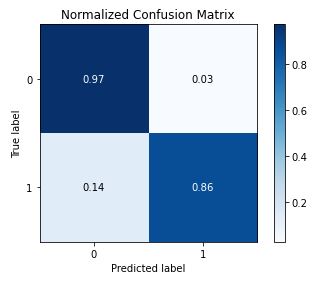
Conclusions
In this study, the best trained GradientBoosting Classifier on the SMOTE-balanced stroke dataset can predict a patient stroke with accuracy=91%, precision = 97%, recall = 96% and f1-score=90%. The summary outcome of our performance QC analysis is as follows:
- learning curves
- average score ~0.4 +/- 0.08 and 0.02 for validation and training data, respectively
- ROC curve area = 0.97
- Precision-recall curve area = 0.963, 0.97 of class 0, 1
- Silhouette score = 0.352
- Calibration plot confirms that the GradientBoosting is the best Classifier
- KS statistic 0.607 at 0.437
- Cumulate gains curve is max 60% deviates from baseline at 50% samples
- Lift is ~ 1.55 at 50% samples
- Elbow plot – the optimal number of clusters is 6
- PCA analysis – 0.872 explained variance ratio for first 1 components
- PCA 2-D projection – we can see the data is non-linear and also some interesting clustering. This suggests a high-dimension classification model or clustering model will work nicely.
References
[1] https://www.researchgate.net/publication/352261064
[2] https://doi.org/10.1016/j.health.2022.100032
[3] https://doi.org/10.1016/j.jstrokecerebrovasdis.2020.105162
[4] https://orfe.princeton.edu/

Leave a comment