A wildfire, forest fire, bushfire, wildland fire or rural fire is an unplanned, uncontrolled and unpredictable fire in an area of combustible vegetation starting in rural and urban areas.
Wildland fire is a widespread and critical element of the Earth’s system. Presently, global annual area burned is estimated to be approximately 420 Mha (Giglio et al. 2018), which is greater in area than the country of India. Wildland fires can result in significant impacts to humans, either directly through loss of life and destruction to communities or indirectly through smoke exposure. Moreover, as the climate warms, we are seeing increasing impacts from wildland fire (Coogan et al. 2019).
Consequently, billions of dollars are spent every year on fire management activities aimed at mitigating or preventing wildfires’ negative effects. Understanding and better predicting wildfires is therefore crucial in several important areas of wildfire management, including emergency response, ecosystem management, land-use planning, and climate adaptation to name a few.
Contents:
- Import Libraries
- Read Input Data
- Exploratory Data Analysis (EDA)
- ML Data Preparation
- ANN Model Training
- Hyper-Parameter Optimization (HPO)
- Conclusions

Wildfires, whether natural or caused by humans, are considered among the most dangerous and devastating disasters around the world. Their complexity comes from the fact that they are hard to predict, hard to extinguish and cause enormous financial losses. To address this issue, many research efforts have been conducted in order to monitor, predict and prevent wildfires using several Artificial Intelligence techniques and strategies such as Big Data, Machine Learning (ML), and Remote Sensing.
Artificial intelligence (AI) has been applied in wildfire science and management since the 1990s, with early applications including neural networks and expert systems. Since then, the field has rapidly progressed congruently with the wide adoption of machine learning (ML) methods in the environmental sciences. The diagram below presents a scoping review of ML applications in wildfire science and management.

In this project, we deploy ML applications in wildfire science and management. The objective is to improve awareness of ML methods among fire researchers and managers and illustrate the diverse and challenging problems in wildfire open to data scientists. Specifically, we develop an early warning detection system of forest fires. We revise the Deep Learning (DL) framework that focuses on understanding the effects of the physical and environmental conditions such as temperature, wind, humidity and relative humidity on predicting the occurrence of fire in a given area.
The dataset used in this project was downloaded from the UCI Machine Learning Repository. This is a public dataset that was used to predict the burned area of forest fires, in the NE region of Portugal (the Montesinho park), by using meteorological and other data.
Following the earlier study, the entire end-to-end ETL ML pipeline is implemented in Python/Jupyter as the following sequence of steps:
- Import/install key libraries
- Download the input dataset
- Data Preparation/Manipulations
- Exploratory Data Analysis (EDA)
- Model Training, Testing and Validation
- Hyperparameter Tuning/Optimization
- Model Performance Evaluation
- Export/Visualization of Prediction Results
Import Libraries
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
Let’s import key libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use(‘seaborn’)
import seaborn as sns
from sklearn.preprocessing import LabelEncoder, StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
import tensorflow as tensorflow
from keras.models import Sequential
from keras.layers import Dense, Dropout
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.utils import to_categorical
from keras.callbacks import EarlyStopping
from keras.callbacks import ModelCheckpoint
from keras.utils.vis_utils import plot_model
%matplotlib inline
Read Input Data
Let’s read the csv file
df = pd.read_csv(‘forestfires.csv’)
df.head(10)

We can see the following attributes in the above table:
- X and Y are the spatial coordinate numbers (1-9)
- month of the year (1-12)
- day of the week (1-7)
- FFMC (Fine Fuel Moisture Code) index from the FWI system (18.7-96.20)
- DMC (Duff Moisture Code) index from the FWI system (1.1-291.3)
- DC (Drought Code) index from the FWI system (7.9-860.6)
- ISI (Initial Spread Index) index from the FWI system (0.0-56.10)
- temperature in Celsius degrees (2.2-33.30)
- RH = relative humidity in % (15.0-100)
- wind speed in km/h (0.40 to 9.40)
- rain in mm/m2 (0.0-6.4)
- area = the burned area of the forest in ha (0.00-1090.84).
Let’s check the data structure
df.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 517 entries, 0 to 516 Data columns (total 13 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 X 517 non-null int64 1 Y 517 non-null int64 2 month 517 non-null object 3 day 517 non-null object 4 FFMC 517 non-null float64 5 DMC 517 non-null float64 6 DC 517 non-null float64 7 ISI 517 non-null float64 8 temp 517 non-null float64 9 RH 517 non-null int64 10 wind 517 non-null float64 11 rain 517 non-null float64 12 area 517 non-null float64 dtypes: float64(8), int64(3), object(2) memory usage: 52.6+ KB
Count null features in the dataset
df.isnull().sum()
X 0 Y 0 month 0 day 0 FFMC 0 DMC 0 DC 0 ISI 0 temp 0 RH 0 wind 0 rain 0 area 0 dtype: int64
Check the number of columns and rows in the dataset
print(df.shape)
(517, 13)
Check statistical information about numerical fields
df.describe()

It appears that
- the structured dataset consists of 13 columns (attributes) and 517 rows (measurements)
- columns X and Y are int indices that do not represent actual coordinates
- the dataset does not contain NaN values.
Exploratory Data Analysis (EDA)
Let’s plot the pie-chart comparing 12 months
plt.figure(figsize= (12,8))
explode_gender=(0.1,0.1,0.1,0.1,0.1,0.1,0.1,0.1,0.2,0.25,0.35,0.55)
colors = [‘#ff9999′,’#66b3ff’,’#99ff99′,’#ffcc99′,’#c2c2f0′,’#ffb3e6′,’#ffebcd’,’#ffefdb’,’#66cdaa’,’#e3cf57′,’#0000ff’,’#ff4040′]
fig = df[“month”].value_counts(normalize = True).plot.pie(autopct=’%1.2f%%’,colors=colors,explode=explode_gender,wedgeprops={‘linewidth’: 3.0, ‘edgecolor’: ‘white’},
textprops={‘size’: ‘x-large’},
startangle=90,labeldistance = 1.1)
fig.legend(title=”month”,
loc=”center left”,
bbox_to_anchor=(1.15, 0, 0.5, 1))
plt.savefig(‘piechartmonth.png’)
plt.show()

According to this chart, August and September are the most wildfire-prone months in the study area.
Let’s plot the pie-chart comparing 7 days of the week
plt.figure(figsize= (10,6))
import plotly.express as px
from matplotlib import cm
import random
import matplotlib.colors as mcolors
number_of_colors=7
#colors = random.choices(list(mcolors.CSS4_COLORS.values()),k = number_of_colors)
colors = [‘#ff9999′,’#66b3ff’,’#99ff99′,’#ffcc99′,’#c2c2f0′,’#ffb3e6′,’#ffebcd’]
fig = df[“day”].value_counts(normalize = True).plot.pie(autopct=’%1.2f%%’,colors=colors,textprops={‘size’: ‘x-large’})
plt.title(“Pie-chart showing Day”, fontdict={‘fontsize’: 20, ‘fontweight’ : 5, ‘color’ : ‘Blue’})
fig.legend(title=”day”,
loc=”center left”,
bbox_to_anchor=(1.1, 0, 0.5, 1))
plt.savefig(‘piechartday.png’)
plt.show()
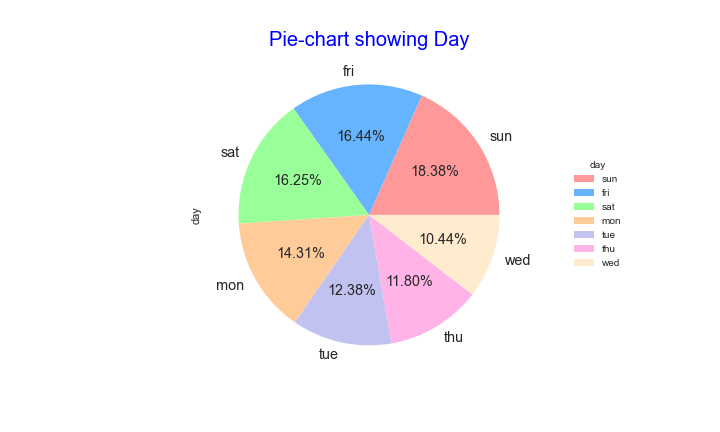
The distribution of wildfires looks pretty uniform throughout the week, with the slight increase during weekends due to a relatively slow response of firefighters.
Let’s plot the attribute list
df.columns
Index(['X', 'Y', 'month', 'day', 'FFMC', 'DMC', 'DC', 'ISI', 'temp', 'RH',
'wind', 'rain', 'area'],
dtype='object')
df.dtypes
X int64 Y int64 month object day object FFMC float64 DMC float64 DC float64 ISI float64 temp float64 RH int64 wind float64 rain float64 area float64 dtype: object
Let’s create the list
X = df[[‘FFMC’, ‘DMC’ , ‘DC’, ‘ISI’, ‘RH’,’wind’,’temp’]]
and plot the cross-pairs
sns_plot=sns.pairplot(X)
plt.savefig(‘snspairplot.png’)
plt.show()
![sns pair plot of the list
X = df[['FFMC', 'DMC' , 'DC', 'ISI', 'RH','wind','temp']]](https://newdigitals.org/wp-content/uploads/2022/07/snspairplot.png?w=1024)
We can see a significant correlation trends in the following X-plots: (temp-RH), (temp-ISI), and (temp-FFMC). At the same time, wind does not appear to exhibit strong correlations with other attributes.
Let’s plot the correlation coefficient heatmap
sns.heatmap(X.corr(), annot=True)
plt.savefig(‘snscorrheatmap.png’)
plt.show()
![X.corr() sns heatmap
where
X = df[['FFMC', 'DMC' , 'DC', 'ISI', 'RH','wind','temp']]](https://newdigitals.org/wp-content/uploads/2022/07/snscorrheatmap.png?w=432)
We can see that the pair DC-DMC has the maximum correlation of 0.68, while the pair wind-RH has the minimum negative correlation of -0.53.
Let’s create the following scatter plot
import matplotlib.pyplot as plt
plt.figure(figsize=(8, 8), dpi=80)
SMALL_SIZE = 14
MEDIUM_SIZE = 14
BIGGER_SIZE = 14
plt.rc(‘font’, size=SMALL_SIZE) # controls default text sizes
plt.rc(‘axes’, titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc(‘axes’, labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc(‘xtick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘ytick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘legend’, fontsize=SMALL_SIZE) # legend fontsize
plt.rc(‘figure’, titlesize=BIGGER_SIZE) # fontsize of the figure title
plt.scatter(df[‘DMC’],df[‘FFMC’],s=df[‘RH’]*2,c=df[‘temp’],cmap=’magma’) plt.colorbar() plt.xlabel(“DMC”) plt.ylabel(“FFMC”) plt.text( 3.2, 35, “Size of marker = RH*2\n” “Color of marker = temp”,
)
plt.savefig(‘pltscatterdmcffmc_rhtemp.png’)
plt.show()
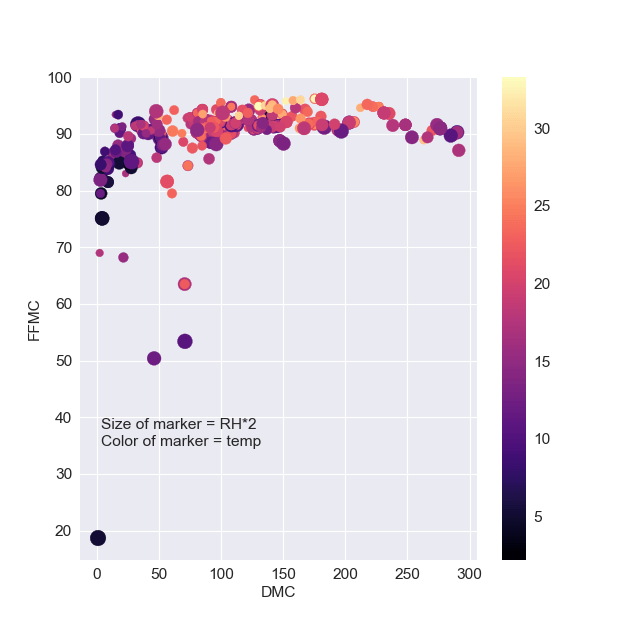
This plot shows a strong correlation between DMC and FFMC, except outliers in the range of lower temp<15 and higher RH.
Let’s create the 2nd scatter plot
plt.figure(figsize=(8, 8), dpi=80)
SMALL_SIZE = 14
MEDIUM_SIZE = 14
BIGGER_SIZE = 14
plt.rc(‘font’, size=SMALL_SIZE) # controls default text sizes
plt.rc(‘axes’, titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc(‘axes’, labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc(‘xtick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘ytick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘legend’, fontsize=SMALL_SIZE) # legend fontsize
plt.rc(‘figure’, titlesize=BIGGER_SIZE) # fontsize of the figure title
plt.scatter(df[‘DC’],df[‘ISI’],s=df[‘RH’]*2,c=df[‘temp’],cmap=’magma’) plt.colorbar() plt.xlabel(“DC”) plt.ylabel(“ISI”) plt.text( 3.2, 35, “Size of marker = RH*2\n” “Color of marker = temp”,
)
plt.savefig(‘pltscatterdcisi_rhtemp.png’)
plt.show()
Let’s create the 3rd scatter plot
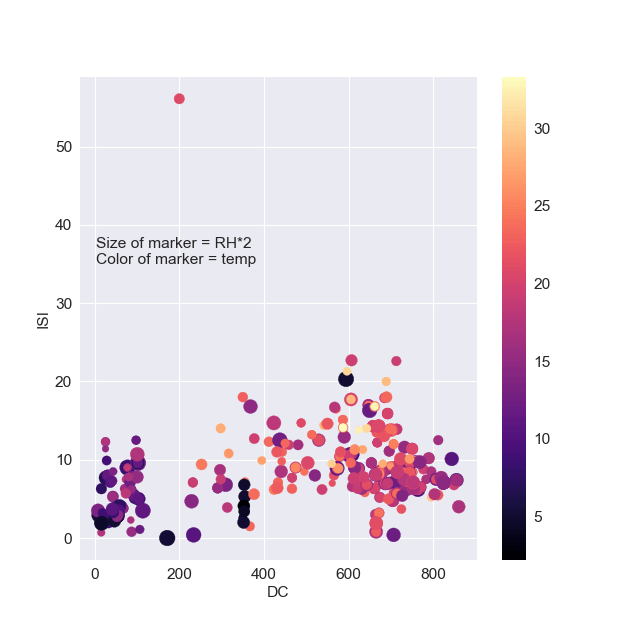
We can observe a weak correlation between ISI and DC, with a single outlier ISI>55 at DC~200.
plt.figure(figsize=(8, 8), dpi=80)
SMALL_SIZE = 14
MEDIUM_SIZE = 14
BIGGER_SIZE = 14
plt.rc(‘font’, size=SMALL_SIZE) # controls default text sizes
plt.rc(‘axes’, titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc(‘axes’, labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc(‘xtick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘ytick’, labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc(‘legend’, fontsize=SMALL_SIZE) # legend fontsize
plt.rc(‘figure’, titlesize=BIGGER_SIZE) # fontsize of the figure title
plt.scatter(df[‘RH’],df[‘FFMC’],s=df[‘ISI’]10,c=df[‘temp’],cmap=’magma’) plt.colorbar() plt.xlabel(“RH”) plt.ylabel(“FFMC”) plt.text( 19.8, 32.5, “Size of marker = ISI10\n” “Color of marker = temp”
)
plt.savefig(‘pltscatterrhffmc_isitemp.png’)
plt.show()
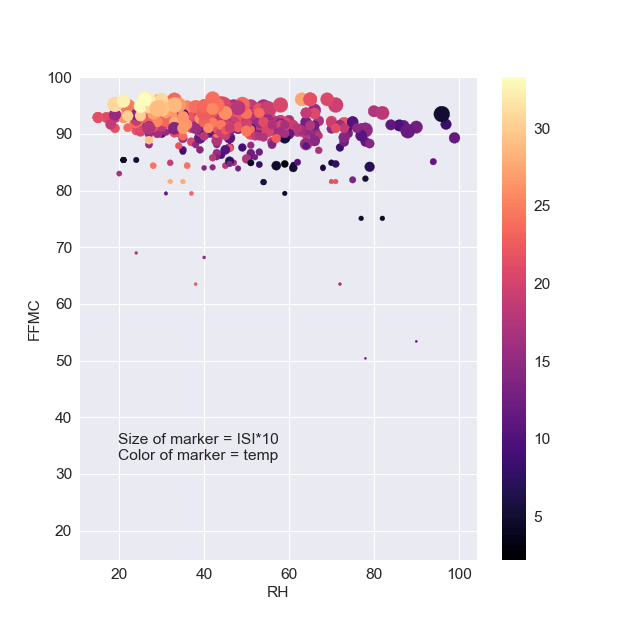
This plot shows a strong correlation between FFMC and RH, excluding outliers at very low values of ISI as FFMC<80.
Let’s look at the monthly FFMC box-plots
fig = px.box(df, x=’month’, y=’FFMC’, points=”all”)
fig.update_layout(
title_text=”Monthly FFMC Spread”)

and the corresponding monthly/weekly violin-plots
fig = px.violin(df, y=”FFMC”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly FFMC”)
fig.show()
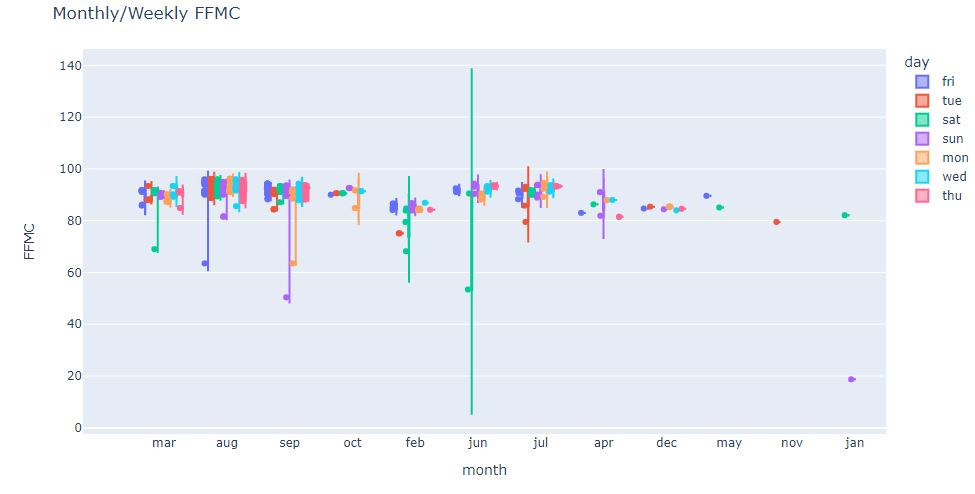
We can see that the maximum spread takes place in jan and jun/sat. Most outliers are recorded on weekends when FFMC<80.
Let’s look at the DMC monthly spread
fig = px.box(df, x=’month’, y=’DMC’, points=”all”)
fig.update_layout(
title_text=”Monthly DMC Spread”)
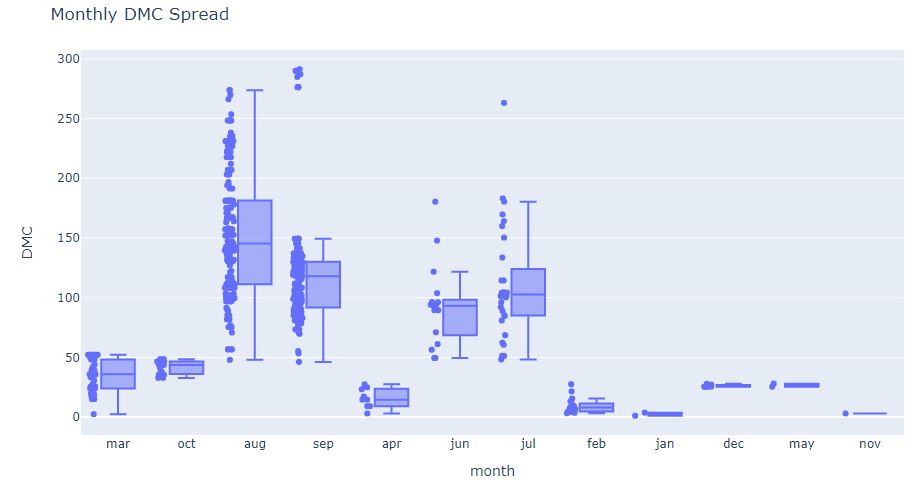
and the corresponding violin plot showing monthly/weekly DMC spread
fig = px.violin(df, y=”DMC”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly DMC”)
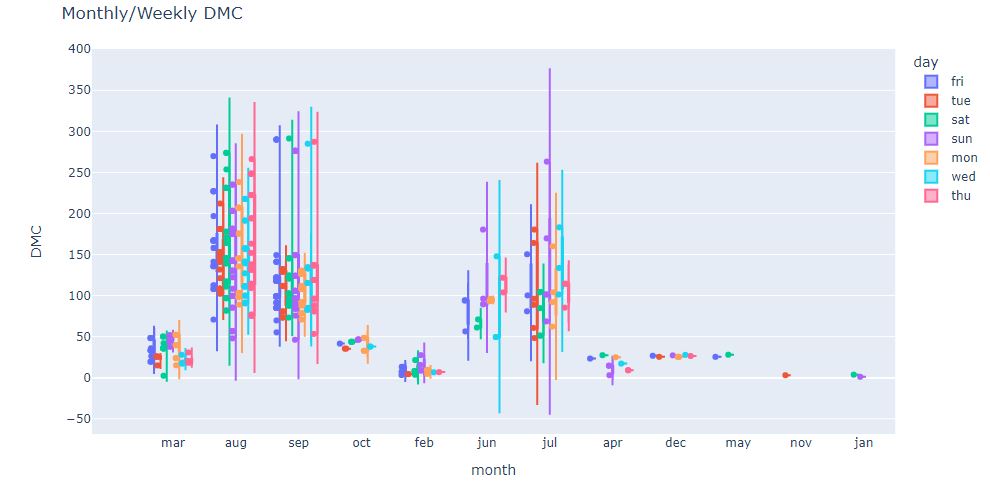
These plots show that jun, jul, aug and sep have seen the highest DMC spreads for both weekdays and weekends.
Let’s look at the DC monthly spread
fig = px.box(df, x=’month’, y=’DC’, points=”all”)
fig.update_layout(
title_text=”Monthly DC Spread”)

and the corresponding violin plot showing monthly/weekly DC spread
fig = px.violin(df, y=”DC”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly DC”)

We can see a few DC outliers in aug feb and jul violin-plots. The data recorded in nov, dec, jan, apr, and may may not be reliable.
Let’s look at the ISI monthly spread
fig = px.box(df, x=’month’, y=’ISI’, points=”all”)
fig.update_layout(
title_text=”Monthly ISI Spread”)

and the corresponding monthly/weekly ISI violin-plot
fig = px.violin(df, y=”ISI”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly ISI”)
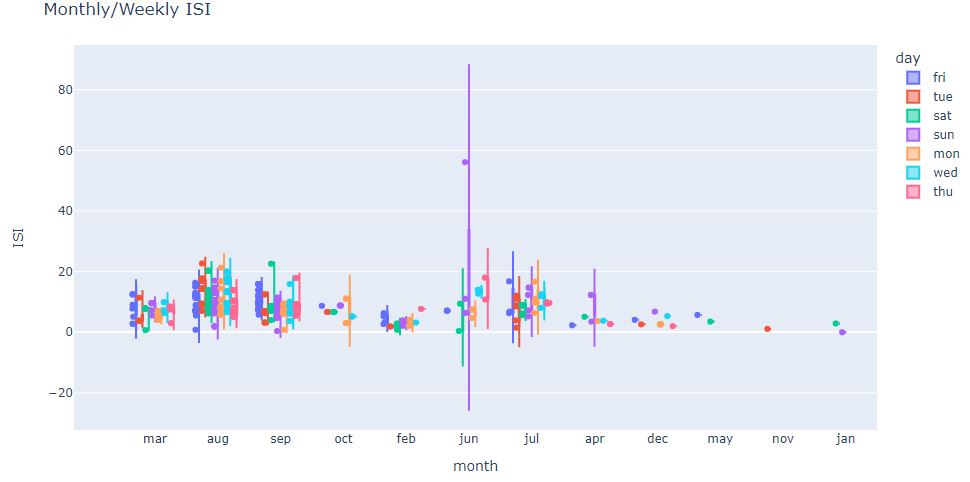
We can see 1 ISI outlier on sun jun and insufficient monthly data samples in dec, may, nov, and jan.
Let’s look at the RH monthly spread
fig = px.box(df, x=’month’, y=’RH’, points=”all”)
fig.update_layout(
title_text=”Monthly RH Spread”)
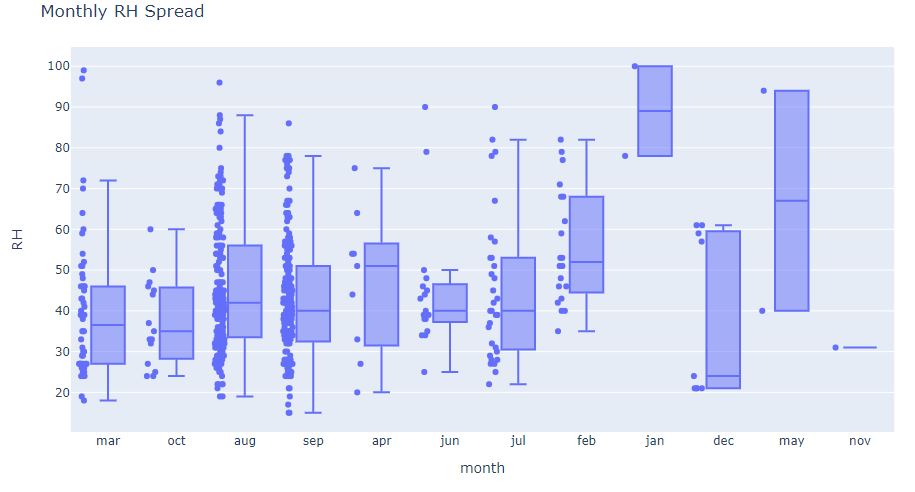
and the corresponding monthly/weekly RH violin-plot
fig = px.violin(df, y=”RH”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly RH”)

We can see that the monthly RH data in apr, dec, may, nov, and jan are not suffifient. Also, notice RH outliers when RH>100 and RH<10.
Let’s look at the temp monthly spread
fig = px.box(df, x=’month’, y=’temp’, points=”all”)
fig.update_layout(
title_text=”Monthly temp Spread”)
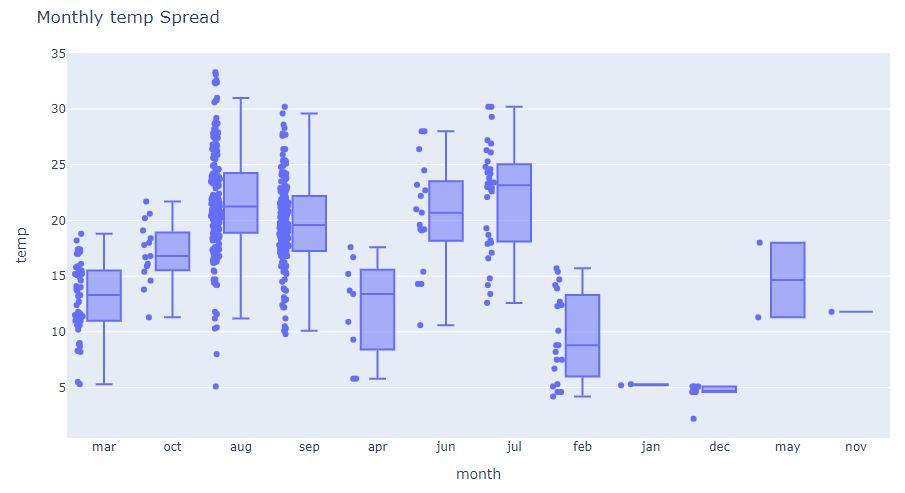
and the corresponding monthly/weekly temp violin-plot
fig = px.violin(df, y=”temp”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly temp”)

We can see 1 temp outlier on sat jun, other outliers as temp<5.0 and sparse temp data sampling in dec, may, nov, and jan.
Let’s look at the wind monthly spread
fig = px.box(df, x=’month’, y=’wind’, points=”all”)
fig.update_layout(
title_text=”Monthly wind Spread”)
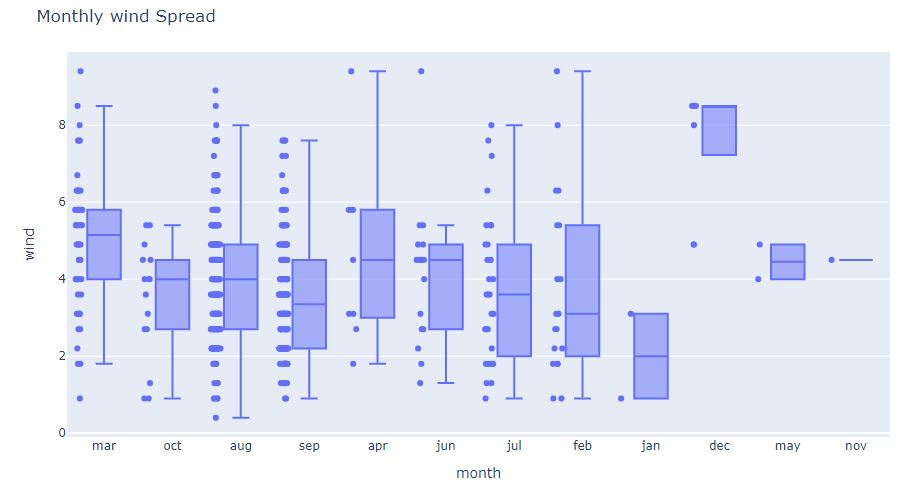
and the corresponding monthly/weekly wind violin-plot
fig = px.violin(df, y=”wind”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly wind”)

We can see several wind outliers as wind>10.0 and wind <2.0. Observe sparse monthly wind data in apr, dec, may, nov, and jan.
Let’s look at the rain monthly spread
fig = px.box(df, x=’month’, y=’rain’, points=”all”)
fig.update_layout(
title_text=”Monthly rain Spread”)

and the corresponding monthly/weekly rain violin-plot
fig = px.violin(df, y=”rain”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly rain”)

We can see that rain ~0.0 except a few outliers rain >1.0 in aug.
Let’s look at the area (target variable) monthly spread
fig = px.box(df, x=’month’, y=’area’, points=”all”)
fig.update_layout(
title_text=”Monthly area Spread”)

and the corresponding monthly/weekly area violin-plot
fig = px.violin(df, y=”area”, x=”month”, color=”day”, box=True, points=”all”, hover_data=df.columns)
fig.update_layout(title_text=”Monthly/Weekly area”)

We can see a few area outliers in jul, aug and sep as area>200. Notice sparse area data in dec, may, nov, and jan.
Let’s look at the following density plots:
sns.distplot(df.FFMC)
plt.savefig(‘distplot_ffmc.png’)
plt.show()

sns.distplot(df.DMC)
plt.savefig(‘distplot_dmc.png’)
plt.show()
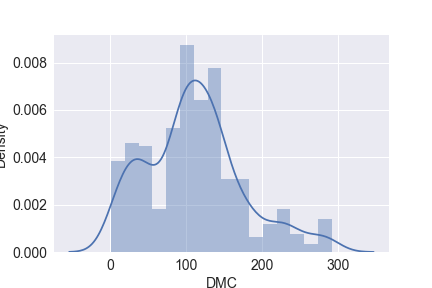
sns.distplot(df.DC)
plt.savefig(‘distplot_dc.png’)
plt.show()

sns.distplot(df.ISI)
plt.savefig(‘distplot_isi.png’)
plt.show()

sns.distplot(df.RH)
plt.savefig(‘distplot_rh.png’)
plt.show()
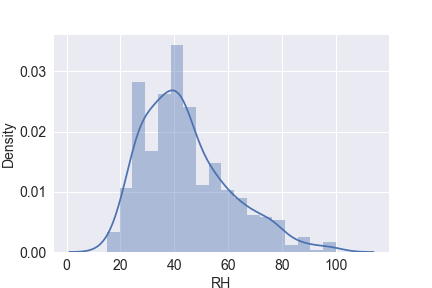
sns.distplot(df.temp)
plt.savefig(‘distplot_temp.png’)
plt.show()

sns.distplot(df.wind)
plt.savefig(‘distplot_wind.png’)
plt.show()
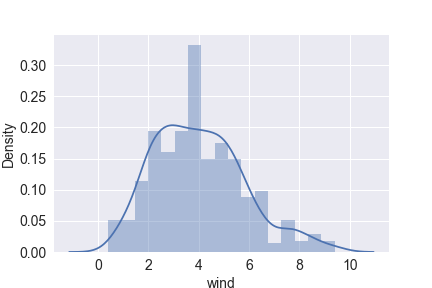
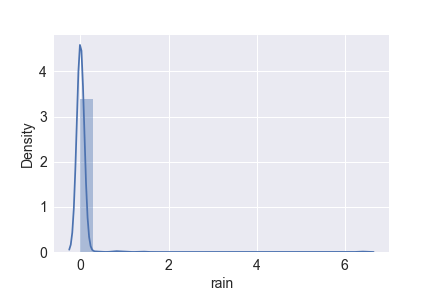
sns.distplot(df.rain)
plt.savefig(‘distplot_rain.png’)
plt.show()
sns.distplot(df.area)
plt.savefig(‘distplot_area.png’)
plt.show()

Let’s look at the pivot tables of interest:
pivot8 = pd.pivot_table(data = df, index = “DMC”, columns = “DC”, values = “temp”)
plt.figure(figsize= (10,6))
sns.heatmap(pivot8, cmap = “Greens”, annot = False,fmt=’.2f’)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “wind”, columns = “temp”, values = “ISI”)
plt.figure(figsize= (10,6))
sns.heatmap(pivot8, cmap = “Greens”, annot = False,fmt=’.2f’)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “ISI”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()
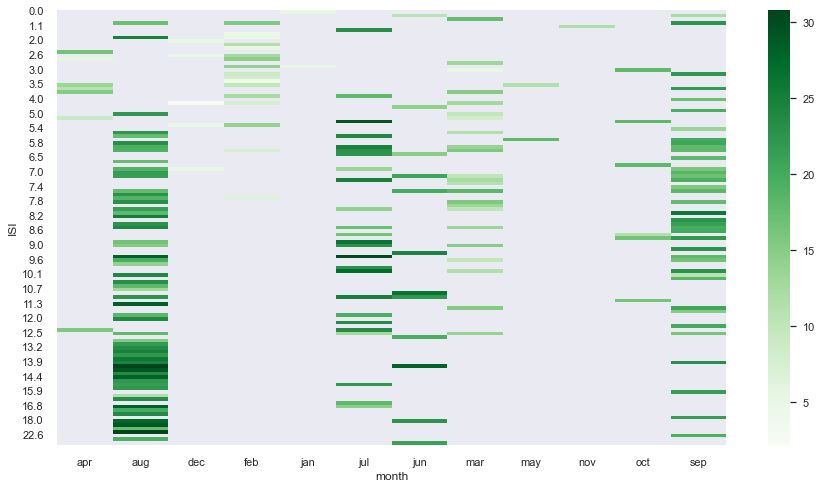
pivot8 = pd.pivot_table(data = df, index = “FFMC”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()
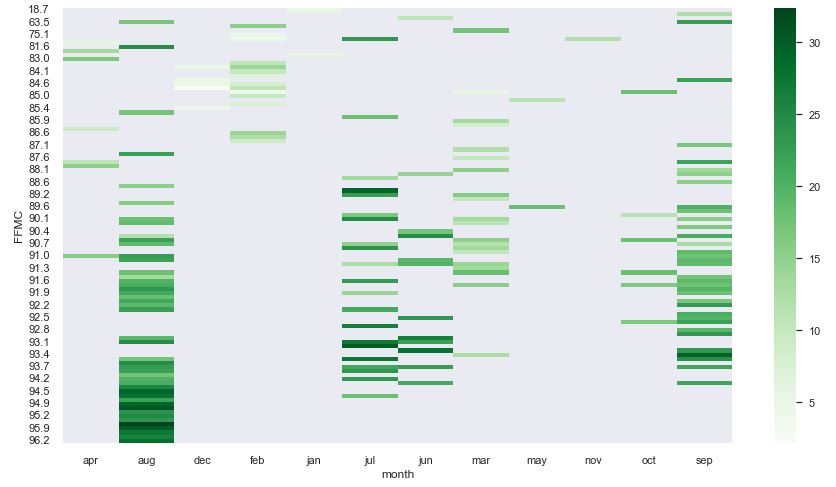
pivot8 = pd.pivot_table(data = df, index = “RH”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()
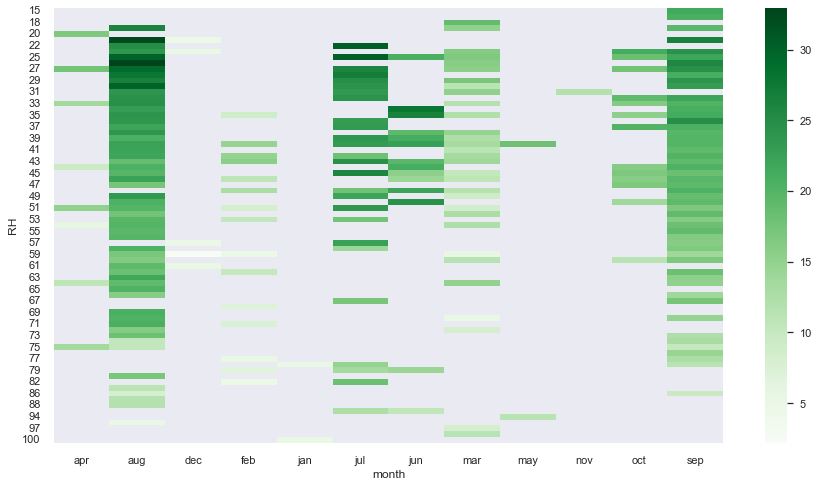
pivot8 = pd.pivot_table(data = df, index = “DMC”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “DC”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “area”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “rain”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()

pivot8 = pd.pivot_table(data = df, index = “wind”, columns = “month”, values = “temp”)
sns.set(rc = {‘figure.figsize’:(15,8)})
sns.heatmap(pivot8, cmap = “Greens”, annot = False)
plt.show()

These pivot table show that the best data quality is observed in jul, aug, and sep.
ML Data Preparation
Let’s transform/rename our weekly data
df[‘day’] = ((df[‘day’] == ‘sun’) | (df[‘day’] == ‘sat’))
df = df.rename(columns = {‘day’ : ‘is_weekend’})
and plot the day count
sns.countplot(df[‘is_weekend’])
plt.title(‘Count plot of weekend vs weekday’)

Let’s scale the rain and area data representing skewed distributions with outliers, as shown above:
df.loc[:, [‘rain’, ‘area’]] = df.loc[:, [‘rain’, ‘area’]].apply(lambda x: np.log(x + 1), axis = 1)
Let’s consider the best quality monthly data, as discussed above:
df[‘month’] = ((df[‘month’] == ‘aug’) | (df[‘month’] == ‘sep’)| (df[‘month’] == ‘jul’))
Let’s split train/test data as 80:20
features = df.drop([‘size_category’], axis = 1)
labels = df[‘size_category’].values.reshape(-1, 1)
X_train, X_test, y_train, y_test = train_test_split(features,labels, test_size = 0.2, random_state = 42)
and apply StandardScaler()
sc_features = StandardScaler()
X_test = sc_features.fit_transform(X_test)
X_train = sc_features.transform(X_train)
Features:
X_test = pd.DataFrame(X_test, columns = features.columns)
X_train = pd.DataFrame(X_train, columns = features.columns)
Labels:
y_test = pd.DataFrame(y_test, columns = [‘size_category’])
y_train = pd.DataFrame(y_train, columns = [‘size_category’])
ANN Model Training
Let’s define our ANN model as follows
model = Sequential()
Input layer + 1st hidden layer:
model.add(Dense(6, input_dim=13, activation=’relu’))
2nd hidden layer:
model.add(Dense(6, activation=’relu’))
Output layer:
model.add(Dense(6, activation=’sigmoid’))
model.add(Dropout(0.2))
model.add(Dense(1, activation = ‘relu’))
model.summary()
Model: "sequential"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
dense (Dense) (None, 6) 84
dense_1 (Dense) (None, 6) 42
dense_2 (Dense) (None, 6) 42
dropout (Dropout) (None, 6) 0
dense_3 (Dense) (None, 1) 7
=================================================================
Total params: 175
Trainable params: 175
Non-trainable params: 0
Let’s compile the model
model.compile(optimizer = ‘adam’, metrics=[‘accuracy’], loss =’binary_crossentropy’)
and train it
history = model.fit(X_train, y_train, validation_data = (X_test, y_test), batch_size = 10, epochs = 100)
Epoch 1/100 42/42 [==============================] - 0s 3ms/step - loss: 3.9964 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 2/100 42/42 [==============================] - 0s 1ms/step - loss: 3.9510 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 3/100 42/42 [==============================] - 0s 1ms/step - loss: 3.9330 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 4/100 42/42 [==============================] - 0s 1ms/step - loss: 3.8950 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 5/100 42/42 [==============================] - 0s 1ms/step - loss: 3.7621 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 6/100 42/42 [==============================] - 0s 1ms/step - loss: 3.7872 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 7/100 42/42 [==============================] - 0s 1ms/step - loss: 3.5116 - accuracy: 0.7312 - val_loss: 4.1529 - val_accuracy: 0.7308 Epoch 8/100 42/42 [==============================] - 0s 1ms/step - loss: 3.6054 - accuracy: 0.7337 - val_loss: 3.9324 - val_accuracy: 0.7308 Epoch 9/100 42/42 [==============================] - 0s 1ms/step - loss: 3.4001 - accuracy: 0.7215 - val_loss: 3.4334 - val_accuracy: 0.7308 Epoch 10/100 42/42 [==============================] - 0s 1ms/step - loss: 2.8922 - accuracy: 0.7288 - val_loss: 2.8330 - val_accuracy: 0.7308 Epoch 11/100 42/42 [==============================] - 0s 1ms/step - loss: 2.4768 - accuracy: 0.7433 - val_loss: 1.2727 - val_accuracy: 0.7308 Epoch 12/100 42/42 [==============================] - 0s 1ms/step - loss: 1.6051 - accuracy: 0.7240 - val_loss: 0.6996 - val_accuracy: 0.7404 Epoch 13/100 42/42 [==============================] - 0s 1ms/step - loss: 1.3042 - accuracy: 0.7409 - val_loss: 0.6729 - val_accuracy: 0.7500 Epoch 14/100 42/42 [==============================] - 0s 1ms/step - loss: 1.2543 - accuracy: 0.7361 - val_loss: 0.6578 - val_accuracy: 0.7500 Epoch 15/100 42/42 [==============================] - 0s 1ms/step - loss: 1.2196 - accuracy: 0.7094 - val_loss: 0.5239 - val_accuracy: 0.7788 Epoch 16/100 42/42 [==============================] - 0s 1ms/step - loss: 0.9230 - accuracy: 0.7458 - val_loss: 0.5067 - val_accuracy: 0.7885 Epoch 17/100 42/42 [==============================] - 0s 1ms/step - loss: 0.9433 - accuracy: 0.7627 - val_loss: 0.4927 - val_accuracy: 0.8269 Epoch 18/100 42/42 [==============================] - 0s 1ms/step - loss: 0.8862 - accuracy: 0.7240 - val_loss: 0.3726 - val_accuracy: 0.8462 Epoch 19/100 42/42 [==============================] - 0s 1ms/step - loss: 0.7844 - accuracy: 0.7554 - val_loss: 0.3550 - val_accuracy: 0.8462 Epoch 20/100 42/42 [==============================] - 0s 1ms/step - loss: 0.8198 - accuracy: 0.7240 - val_loss: 0.3450 - val_accuracy: 0.8558 Epoch 21/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5507 - accuracy: 0.7506 - val_loss: 0.3382 - val_accuracy: 0.8558 Epoch 22/100 42/42 [==============================] - 0s 1ms/step - loss: 0.8161 - accuracy: 0.7458 - val_loss: 0.3326 - val_accuracy: 0.8462 Epoch 23/100 42/42 [==============================] - 0s 1ms/step - loss: 0.7026 - accuracy: 0.7603 - val_loss: 0.3232 - val_accuracy: 0.8654 Epoch 24/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5395 - accuracy: 0.7651 - val_loss: 0.3169 - val_accuracy: 0.8654 Epoch 25/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5820 - accuracy: 0.7942 - val_loss: 0.3098 - val_accuracy: 0.8558 Epoch 26/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5944 - accuracy: 0.7651 - val_loss: 0.3020 - val_accuracy: 0.8654 Epoch 27/100 42/42 [==============================] - 0s 1ms/step - loss: 0.6394 - accuracy: 0.7651 - val_loss: 0.2969 - val_accuracy: 0.8654 Epoch 28/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5938 - accuracy: 0.7651 - val_loss: 0.2896 - val_accuracy: 0.8750 Epoch 29/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5811 - accuracy: 0.7797 - val_loss: 0.2835 - val_accuracy: 0.8750 Epoch 30/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3789 - accuracy: 0.8329 - val_loss: 0.2797 - val_accuracy: 0.8846 Epoch 31/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5529 - accuracy: 0.8063 - val_loss: 0.2734 - val_accuracy: 0.8846 Epoch 32/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5569 - accuracy: 0.8039 - val_loss: 0.2654 - val_accuracy: 0.8846 Epoch 33/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5183 - accuracy: 0.8039 - val_loss: 0.2590 - val_accuracy: 0.8942 Epoch 34/100 42/42 [==============================] - 0s 1ms/step - loss: 0.5420 - accuracy: 0.7845 - val_loss: 0.2572 - val_accuracy: 0.9038 Epoch 35/100 42/42 [==============================] - 0s 1ms/step - loss: 0.4040 - accuracy: 0.8015 - val_loss: 0.2511 - val_accuracy: 0.9135 Epoch 36/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3933 - accuracy: 0.8257 - val_loss: 0.2446 - val_accuracy: 0.9038 Epoch 37/100 42/42 [==============================] - 0s 1ms/step - loss: 0.4350 - accuracy: 0.8208 - val_loss: 0.2378 - val_accuracy: 0.9135 Epoch 38/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3923 - accuracy: 0.7748 - val_loss: 0.2341 - val_accuracy: 0.9135 Epoch 39/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3426 - accuracy: 0.8354 - val_loss: 0.2309 - val_accuracy: 0.9135 Epoch 40/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3220 - accuracy: 0.8354 - val_loss: 0.2272 - val_accuracy: 0.9135 Epoch 41/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2793 - accuracy: 0.8426 - val_loss: 0.2248 - val_accuracy: 0.9231 Epoch 42/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3744 - accuracy: 0.8208 - val_loss: 0.2247 - val_accuracy: 0.9135 Epoch 43/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2990 - accuracy: 0.8281 - val_loss: 0.2243 - val_accuracy: 0.9135 Epoch 44/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3358 - accuracy: 0.8136 - val_loss: 0.2193 - val_accuracy: 0.9231 Epoch 45/100 42/42 [==============================] - 0s 1ms/step - loss: 0.3040 - accuracy: 0.8378 - val_loss: 0.2149 - val_accuracy: 0.9231 Epoch 46/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2857 - accuracy: 0.8426 - val_loss: 0.2123 - val_accuracy: 0.9231 Epoch 47/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2844 - accuracy: 0.8475 - val_loss: 0.2135 - val_accuracy: 0.9135 Epoch 48/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2593 - accuracy: 0.8668 - val_loss: 0.2134 - val_accuracy: 0.9135 Epoch 49/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2941 - accuracy: 0.8668 - val_loss: 0.2089 - val_accuracy: 0.9135 Epoch 50/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2833 - accuracy: 0.8426 - val_loss: 0.2054 - val_accuracy: 0.9231 Epoch 51/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2422 - accuracy: 0.8547 - val_loss: 0.2017 - val_accuracy: 0.9231 Epoch 52/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2671 - accuracy: 0.8862 - val_loss: 0.1998 - val_accuracy: 0.9231 Epoch 53/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2401 - accuracy: 0.8765 - val_loss: 0.2019 - val_accuracy: 0.9231 Epoch 54/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2222 - accuracy: 0.8959 - val_loss: 0.1992 - val_accuracy: 0.9231 Epoch 55/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2345 - accuracy: 0.8717 - val_loss: 0.1948 - val_accuracy: 0.9231 Epoch 56/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2173 - accuracy: 0.8983 - val_loss: 0.1922 - val_accuracy: 0.9038 Epoch 57/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2781 - accuracy: 0.8765 - val_loss: 0.1908 - val_accuracy: 0.9135 Epoch 58/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2143 - accuracy: 0.9031 - val_loss: 0.1867 - val_accuracy: 0.9135
Epoch 59/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2553 - accuracy: 0.8959 - val_loss: 0.1814 - val_accuracy: 0.9135 Epoch 60/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1861 - accuracy: 0.9104 - val_loss: 0.1793 - val_accuracy: 0.9135 Epoch 61/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1968 - accuracy: 0.9128 - val_loss: 0.1737 - val_accuracy: 0.9135 Epoch 62/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1867 - accuracy: 0.9177 - val_loss: 0.1697 - val_accuracy: 0.9135 Epoch 63/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2166 - accuracy: 0.9056 - val_loss: 0.1616 - val_accuracy: 0.9135 Epoch 64/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2100 - accuracy: 0.8910 - val_loss: 0.1572 - val_accuracy: 0.9231 Epoch 65/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2105 - accuracy: 0.9249 - val_loss: 0.1564 - val_accuracy: 0.9135 Epoch 66/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2176 - accuracy: 0.9201 - val_loss: 0.1505 - val_accuracy: 0.9327 Epoch 67/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1684 - accuracy: 0.9177 - val_loss: 0.1471 - val_accuracy: 0.9327 Epoch 68/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1757 - accuracy: 0.9177 - val_loss: 0.1545 - val_accuracy: 0.9135 Epoch 69/100 42/42 [==============================] - 0s 1ms/step - loss: 0.2016 - accuracy: 0.9177 - val_loss: 0.1478 - val_accuracy: 0.9135 Epoch 70/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1990 - accuracy: 0.9322 - val_loss: 0.1423 - val_accuracy: 0.9327 Epoch 71/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1628 - accuracy: 0.9370 - val_loss: 0.1374 - val_accuracy: 0.9423 Epoch 72/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1443 - accuracy: 0.9370 - val_loss: 0.1339 - val_accuracy: 0.9423 Epoch 73/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1568 - accuracy: 0.9298 - val_loss: 0.1317 - val_accuracy: 0.9423 Epoch 74/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1604 - accuracy: 0.9274 - val_loss: 0.1323 - val_accuracy: 0.9423 Epoch 75/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1412 - accuracy: 0.9492 - val_loss: 0.1252 - val_accuracy: 0.9423 Epoch 76/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1372 - accuracy: 0.9370 - val_loss: 0.1184 - val_accuracy: 0.9519 Epoch 77/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1375 - accuracy: 0.9370 - val_loss: 0.1173 - val_accuracy: 0.9519 Epoch 78/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1366 - accuracy: 0.9225 - val_loss: 0.1097 - val_accuracy: 0.9519 Epoch 79/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1279 - accuracy: 0.9395 - val_loss: 0.1112 - val_accuracy: 0.9519 Epoch 80/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1340 - accuracy: 0.9322 - val_loss: 0.1087 - val_accuracy: 0.9519 Epoch 81/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1659 - accuracy: 0.9419 - val_loss: 0.1091 - val_accuracy: 0.9615 Epoch 82/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1200 - accuracy: 0.9322 - val_loss: 0.1007 - val_accuracy: 0.9615 Epoch 83/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1404 - accuracy: 0.9443 - val_loss: 0.0999 - val_accuracy: 0.9615 Epoch 84/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1102 - accuracy: 0.9540 - val_loss: 0.0964 - val_accuracy: 0.9615 Epoch 85/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1389 - accuracy: 0.9564 - val_loss: 0.1013 - val_accuracy: 0.9615 Epoch 86/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1249 - accuracy: 0.9685 - val_loss: 0.1036 - val_accuracy: 0.9615 Epoch 87/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1446 - accuracy: 0.9467 - val_loss: 0.0920 - val_accuracy: 0.9615 Epoch 88/100 42/42 [==============================] - 0s 1ms/step - loss: 0.0985 - accuracy: 0.9588 - val_loss: 0.0894 - val_accuracy: 0.9615 Epoch 89/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1088 - accuracy: 0.9443 - val_loss: 0.1060 - val_accuracy: 0.9615 Epoch 90/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1456 - accuracy: 0.9443 - val_loss: 0.0969 - val_accuracy: 0.9615 Epoch 91/100 42/42 [==============================] - 0s 1ms/step - loss: 0.0906 - accuracy: 0.9661 - val_loss: 0.0977 - val_accuracy: 0.9615 Epoch 92/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1322 - accuracy: 0.9564 - val_loss: 0.2185 - val_accuracy: 0.9615 Epoch 93/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1119 - accuracy: 0.9395 - val_loss: 0.2174 - val_accuracy: 0.9615 Epoch 94/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1040 - accuracy: 0.9443 - val_loss: 0.1006 - val_accuracy: 0.9615 Epoch 95/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1316 - accuracy: 0.9613 - val_loss: 0.2223 - val_accuracy: 0.9615 Epoch 96/100 42/42 [==============================] - 0s 1ms/step - loss: 0.0881 - accuracy: 0.9637 - val_loss: 0.2296 - val_accuracy: 0.9712 Epoch 97/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1257 - accuracy: 0.9467 - val_loss: 0.1126 - val_accuracy: 0.9712 Epoch 98/100 42/42 [==============================] - 0s 1ms/step - loss: 0.1157 - accuracy: 0.9685 - val_loss: 0.3307 - val_accuracy: 0.9712 Epoch 99/100 42/42 [==============================] - 0s 1ms/step - loss: 0.0810 - accuracy: 0.9613 - val_loss: 0.3307 - val_accuracy: 0.9712 Epoch 100/100 42/42 [==============================] - 0s 1ms/step - loss: 0.0871 - accuracy: 0.9588 - val_loss: 0.3281 - val_accuracy: 0.9712
Let’s check the accuracy
_, train_acc = model.evaluate(X_train, y_train, verbose=0)
_, valid_acc = model.evaluate(X_test, y_test, verbose=0)
print(‘Train: %.3f, Valid: %.3f’ % (train_acc, valid_acc))
Train: 0.976, Valid: 0.971
Let’s plot the corresponding taining/validation accuracy curves
plt.figure(figsize=[8,5])
plt.plot(history.history[‘accuracy’], label=’Train’)
plt.plot(history.history[‘val_accuracy’], label=’Valid’)
plt.legend()
plt.xlabel(‘Epochs’, fontsize=16)
plt.ylabel(‘Accuracy’, fontsize=16)
plt.title(‘Accuracy Curves Epoch 100, Batch Size 10’, fontsize=16)
plt.savefig(‘accuracycurveepoch100.png’)
plt.show()

Hyper-Parameter Optimization (HPO)
Let’s define the following function
Fit a model and plot learning curve
def fit_model(X_train, y_train, X_test, y_test, n_batch):
Define Model
model = Sequential()
model.add(Dense(6, input_dim=13, activation='relu'))
model.add(Dense(6, activation='relu'))
model.add(Dense(6, activation='sigmoid'))
model.add(Dropout(0.2))
model.add(Dense(1, activation = 'relu'))
Compile Model
model.compile(optimizer = 'adam',
metrics=['accuracy'],
loss = 'binary_crossentropy')
Fit Model
history = model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=100, verbose=0, batch_size=n_batch)
Plot Learning Curves
plt.plot(history.history['accuracy'], label='train')
plt.plot(history.history['val_accuracy'], label='test')
plt.title('batch='+str(n_batch))
plt.legend()
Let’s create learning curves for different batch sizes
batch_sizes = [4, 6, 10, 16, 32, 64, 128, 260]
plt.figure(figsize=(10,15))
for i in range(len(batch_sizes)):
Determine the Plot Number
plot_no = 420 + (i+1)
plt.subplot(plot_no)
Fit model and plot learning curves for a batch size
fit_model(X_train, y_train, X_test, y_test, batch_sizes[i])
Show learning curves
plt.savefig(‘accuracycurvebatchvar.png’)
plt.show()
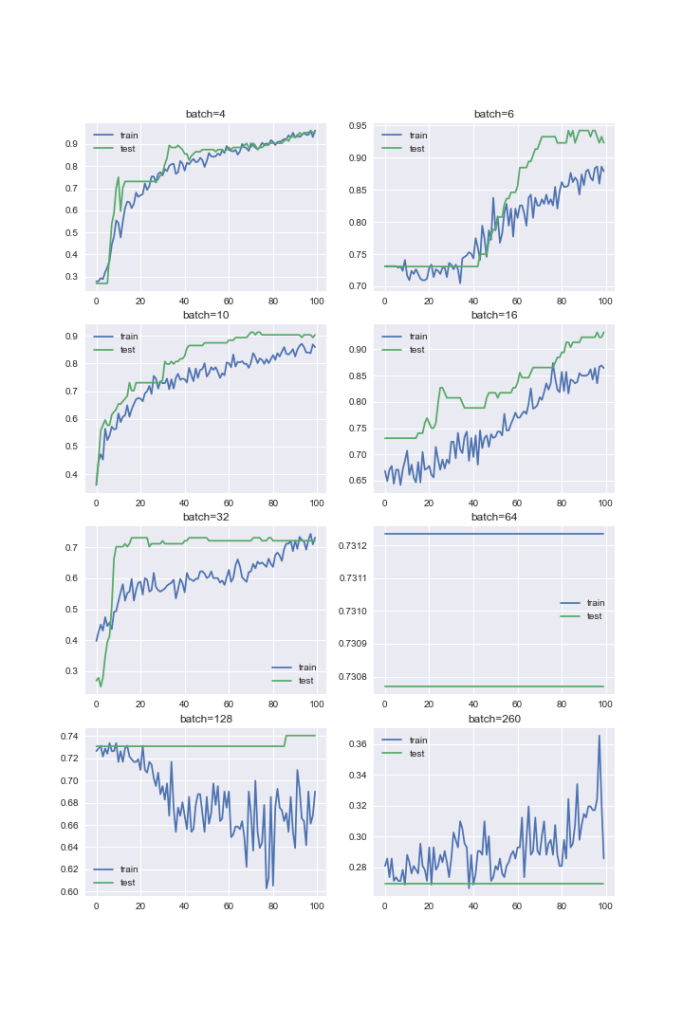
This accuracy graph shows that the value batch = 10 results in the optimal model performance.
Let’s define a function that fits a model and plot learning curve for different epoch given batch size of 10
def fit_model(trainX, trainy, validX, validy, n_epoch):
Define model
model = Sequential()
model.add(Dense(10, input_dim=13, activation='relu'))
model.add(Dense(10, activation='relu'))
model.add(Dense(10, activation='sigmoid'))
model.add(Dropout(0.2))
model.add(Dense(1, activation = 'relu'))
Compile model
model.compile(optimizer ='adam', metrics=['accuracy'], loss = 'binary_crossentropy')
Fit model
history = model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=n_epoch, verbose=0, batch_size=10)
# plot learning curves
plt.plot(history.history['accuracy'], label='train')
plt.plot(history.history['val_accuracy'], label='test')
plt.title('epoch='+str(n_epoch))
plt.legend()
Let’s create learning curves for different
epochs = [20, 50, 100, 120, 150, 200, 300, 400]
plt.figure(figsize=(10,15))
for i in range(len(batch_sizes)):
Determine the Plot Number
plot_no = 420 + (i+1)
plt.subplot(plot_no)
Fit model and plot learning curves for a specific value of epoch
fit_model(X_train, y_train, X_test, y_test, epochs[i])
Show learning curves
plt.savefig(‘accuracycurveepochvar.png’)
plt.show()

This accuracy graph shows that the values epoch>100 result in the optimal model performance.
Let’s define the final model
def init_model():
Define model
model = Sequential()
model.add(Dense(10, input_dim=13, activation='relu'))
model.add(Dense(10, activation='relu'))
model.add(Dense(10, activation='sigmoid'))
model.add(Dropout(0.2))
model.add(Dense(1, activation = 'relu'))
model.compile(optimizer ='adam',
metrics=['accuracy'],
loss = 'binary_crossentropy')
return model
Let’s apply our function init_model
model = init_model()
Simple early stopping
es = EarlyStopping(monitor=’val_loss’, mode=’min’, verbose=1, patience=150)
Model checkpoint
mc = ModelCheckpoint(‘best_model.h5′, monitor=’val_accuracy’, mode=’max’, verbose=1, save_best_only=True)
Fitting model
history = model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=250, verbose=0, batch_size=10, callbacks=[es, mc])
Epoch 1: val_accuracy improved from -inf to 0.73077, saving model to best_model.h5 Epoch 2: val_accuracy did not improve from 0.73077 Epoch 3: val_accuracy did not improve from 0.73077 Epoch 4: val_accuracy did not improve from 0.73077 Epoch 5: val_accuracy did not improve from 0.73077 Epoch 6: val_accuracy did not improve from 0.73077 Epoch 7: val_accuracy did not improve from 0.73077 Epoch 8: val_accuracy did not improve from 0.73077 Epoch 9: val_accuracy did not improve from 0.73077 Epoch 10: val_accuracy did not improve from 0.73077 Epoch 11: val_accuracy did not improve from 0.73077 Epoch 12: val_accuracy did not improve from 0.73077 Epoch 13: val_accuracy did not improve from 0.73077 Epoch 14: val_accuracy did not improve from 0.73077 Epoch 15: val_accuracy did not improve from 0.73077 Epoch 16: val_accuracy did not improve from 0.73077 Epoch 17: val_accuracy did not improve from 0.73077 Epoch 18: val_accuracy did not improve from 0.73077 Epoch 19: val_accuracy did not improve from 0.73077 Epoch 20: val_accuracy did not improve from 0.73077 Epoch 21: val_accuracy did not improve from 0.73077 Epoch 22: val_accuracy did not improve from 0.73077 Epoch 23: val_accuracy did not improve from 0.73077 Epoch 24: val_accuracy did not improve from 0.73077 Epoch 25: val_accuracy did not improve from 0.73077 Epoch 26: val_accuracy did not improve from 0.73077 Epoch 27: val_accuracy did not improve from 0.73077 Epoch 28: val_accuracy did not improve from 0.73077 Epoch 29: val_accuracy did not improve from 0.73077 Epoch 30: val_accuracy did not improve from 0.73077 Epoch 31: val_accuracy did not improve from 0.73077 Epoch 32: val_accuracy did not improve from 0.73077 Epoch 33: val_accuracy did not improve from 0.73077 Epoch 34: val_accuracy did not improve from 0.73077 Epoch 35: val_accuracy did not improve from 0.73077 Epoch 36: val_accuracy did not improve from 0.73077 Epoch 37: val_accuracy did not improve from 0.73077 Epoch 38: val_accuracy did not improve from 0.73077 Epoch 39: val_accuracy did not improve from 0.73077 Epoch 40: val_accuracy did not improve from 0.73077 Epoch 41: val_accuracy did not improve from 0.73077 Epoch 42: val_accuracy did not improve from 0.73077 Epoch 43: val_accuracy did not improve from 0.73077 Epoch 44: val_accuracy did not improve from 0.73077 Epoch 45: val_accuracy did not improve from 0.73077 Epoch 46: val_accuracy did not improve from 0.73077 Epoch 47: val_accuracy did not improve from 0.73077 Epoch 48: val_accuracy did not improve from 0.73077 Epoch 49: val_accuracy did not improve from 0.73077 Epoch 50: val_accuracy did not improve from 0.73077 Epoch 51: val_accuracy did not improve from 0.73077 Epoch 52: val_accuracy did not improve from 0.73077 Epoch 53: val_accuracy did not improve from 0.73077 Epoch 54: val_accuracy did not improve from 0.73077 Epoch 55: val_accuracy did not improve from 0.73077 Epoch 56: val_accuracy did not improve from 0.73077 Epoch 57: val_accuracy did not improve from 0.73077 Epoch 58: val_accuracy did not improve from 0.73077 Epoch 59: val_accuracy did not improve from 0.73077 Epoch 60: val_accuracy did not improve from 0.73077 Epoch 61: val_accuracy did not improve from 0.73077 Epoch 62: val_accuracy did not improve from 0.73077 Epoch 63: val_accuracy did not improve from 0.73077 Epoch 64: val_accuracy did not improve from 0.73077 Epoch 65: val_accuracy improved from 0.73077 to 0.74038, saving model to best_model.h5 Epoch 66: val_accuracy improved from 0.74038 to 0.77885, saving model to best_model.h5 Epoch 67: val_accuracy did not improve from 0.77885 Epoch 68: val_accuracy did not improve from 0.77885 Epoch 69: val_accuracy did not improve from 0.77885 Epoch 70: val_accuracy did not improve from 0.77885 Epoch 71: val_accuracy improved from 0.77885 to 0.79808, saving model to best_model.h5 Epoch 72: val_accuracy did not improve from 0.79808 Epoch 73: val_accuracy did not improve from 0.79808 Epoch 74: val_accuracy improved from 0.79808 to 0.80769, saving model to best_model.h5 Epoch 75: val_accuracy did not improve from 0.80769 Epoch 76: val_accuracy did not improve from 0.80769 Epoch 77: val_accuracy improved from 0.80769 to 0.81731, saving model to best_model.h5 Epoch 78: val_accuracy did not improve from 0.81731 Epoch 79: val_accuracy improved from 0.81731 to 0.83654, saving model to best_model.h5 Epoch 80: val_accuracy did not improve from 0.83654 Epoch 81: val_accuracy did not improve from 0.83654 Epoch 82: val_accuracy did not improve from 0.83654 Epoch 83: val_accuracy did not improve from 0.83654 Epoch 84: val_accuracy improved from 0.83654 to 0.84615, saving model to best_model.h5 Epoch 85: val_accuracy did not improve from 0.84615 Epoch 86: val_accuracy did not improve from 0.84615 Epoch 87: val_accuracy did not improve from 0.84615 Epoch 88: val_accuracy did not improve from 0.84615 Epoch 89: val_accuracy improved from 0.84615 to 0.85577, saving model to best_model.h5 Epoch 90: val_accuracy did not improve from 0.85577 Epoch 91: val_accuracy did not improve from 0.85577 Epoch 92: val_accuracy did not improve from 0.85577 Epoch 93: val_accuracy did not improve from 0.85577 Epoch 94: val_accuracy did not improve from 0.85577 Epoch 95: val_accuracy did not improve from 0.85577 Epoch 96: val_accuracy did not improve from 0.85577 Epoch 97: val_accuracy improved from 0.85577 to 0.86538, saving model to best_model.h5 Epoch 98: val_accuracy did not improve from 0.86538 Epoch 99: val_accuracy did not improve from 0.86538 Epoch 100: val_accuracy did not improve from 0.86538 Epoch 101: val_accuracy did not improve from 0.86538 Epoch 102: val_accuracy did not improve from 0.86538 Epoch 103: val_accuracy did not improve from 0.86538 Epoch 104: val_accuracy did not improve from 0.86538 Epoch 105: val_accuracy did not improve from 0.86538 Epoch 106: val_accuracy did not improve from 0.86538 Epoch 107: val_accuracy did not improve from 0.86538 Epoch 108: val_accuracy did not improve from 0.86538 Epoch 109: val_accuracy did not improve from 0.86538 Epoch 110: val_accuracy did not improve from 0.86538 Epoch 111: val_accuracy did not improve from 0.86538 Epoch 112: val_accuracy improved from 0.86538 to 0.87500, saving model to best_model.h5 Epoch 113: val_accuracy improved from 0.87500 to 0.88462, saving model to best_model.h5 Epoch 114: val_accuracy did not improve from 0.88462 Epoch 115: val_accuracy did not improve from 0.88462 Epoch 116: val_accuracy did not improve from 0.88462 Epoch 117: val_accuracy did not improve from 0.88462 Epoch 118: val_accuracy did not improve from 0.88462 Epoch 119: val_accuracy did not improve from 0.88462 Epoch 120: val_accuracy did not improve from 0.88462 Epoch 121: val_accuracy did not improve from 0.88462 Epoch 122: val_accuracy did not improve from 0.88462 Epoch 123: val_accuracy did not improve from 0.88462 Epoch 124: val_accuracy did not improve from 0.88462 Epoch 125: val_accuracy did not improve from 0.88462 Epoch 126: val_accuracy did not improve from 0.88462 Epoch 127: val_accuracy did not improve from 0.88462 Epoch 128: val_accuracy did not improve from 0.88462 Epoch 129: val_accuracy did not improve from 0.88462 Epoch 130: val_accuracy did not improve from 0.88462 Epoch 131: val_accuracy did not improve from 0.88462 Epoch 132: val_accuracy did not improve from 0.88462 Epoch 133: val_accuracy did not improve from 0.88462 Epoch 134: val_accuracy did not improve from 0.88462 Epoch 135: val_accuracy did not improve from 0.88462 Epoch 136: val_accuracy did not improve from 0.88462 Epoch 137: val_accuracy did not improve from 0.88462 Epoch 138: val_accuracy did not improve from 0.88462 Epoch 139: val_accuracy did not improve from 0.88462 Epoch 140: val_accuracy did not improve from 0.88462 Epoch 141: val_accuracy did not improve from 0.88462 Epoch 142: val_accuracy did not improve from 0.88462 Epoch 143: val_accuracy did not improve from 0.88462 Epoch 144: val_accuracy did not improve from 0.88462 Epoch 145: val_accuracy did not improve from 0.88462 Epoch 146: val_accuracy did not improve from 0.88462
Epoch 147: val_accuracy did not improve from 0.88462 Epoch 148: val_accuracy did not improve from 0.88462 Epoch 149: val_accuracy did not improve from 0.88462 Epoch 150: val_accuracy did not improve from 0.88462 Epoch 151: val_accuracy did not improve from 0.88462 Epoch 152: val_accuracy did not improve from 0.88462 Epoch 153: val_accuracy did not improve from 0.88462 Epoch 154: val_accuracy did not improve from 0.88462 Epoch 155: val_accuracy did not improve from 0.88462 Epoch 156: val_accuracy did not improve from 0.88462 Epoch 157: val_accuracy did not improve from 0.88462 Epoch 158: val_accuracy did not improve from 0.88462 Epoch 159: val_accuracy did not improve from 0.88462 Epoch 160: val_accuracy did not improve from 0.88462 Epoch 161: val_accuracy did not improve from 0.88462 Epoch 162: val_accuracy did not improve from 0.88462 Epoch 163: val_accuracy improved from 0.88462 to 0.89423, saving model to best_model.h5 Epoch 164: val_accuracy did not improve from 0.89423 Epoch 165: val_accuracy did not improve from 0.89423 Epoch 166: val_accuracy did not improve from 0.89423 Epoch 167: val_accuracy did not improve from 0.89423 Epoch 168: val_accuracy did not improve from 0.89423 Epoch 169: val_accuracy did not improve from 0.89423 Epoch 170: val_accuracy did not improve from 0.89423 Epoch 171: val_accuracy did not improve from 0.89423 Epoch 172: val_accuracy did not improve from 0.89423 Epoch 173: val_accuracy did not improve from 0.89423 Epoch 174: val_accuracy did not improve from 0.89423 Epoch 175: val_accuracy did not improve from 0.89423 Epoch 176: val_accuracy did not improve from 0.89423 Epoch 177: val_accuracy did not improve from 0.89423 Epoch 178: val_accuracy did not improve from 0.89423 Epoch 179: val_accuracy did not improve from 0.89423 Epoch 180: val_accuracy did not improve from 0.89423 Epoch 181: val_accuracy did not improve from 0.89423 Epoch 182: val_accuracy improved from 0.89423 to 0.91346, saving model to best_model.h5 Epoch 183: val_accuracy did not improve from 0.91346 Epoch 184: val_accuracy did not improve from 0.91346 Epoch 185: val_accuracy did not improve from 0.91346 Epoch 186: val_accuracy did not improve from 0.91346 Epoch 187: val_accuracy did not improve from 0.91346 Epoch 188: val_accuracy did not improve from 0.91346 Epoch 189: val_accuracy did not improve from 0.91346 Epoch 190: val_accuracy did not improve from 0.91346 Epoch 191: val_accuracy did not improve from 0.91346 Epoch 192: val_accuracy did not improve from 0.91346 Epoch 193: val_accuracy did not improve from 0.91346 Epoch 194: val_accuracy did not improve from 0.91346 Epoch 195: val_accuracy did not improve from 0.91346 Epoch 196: val_accuracy did not improve from 0.91346 Epoch 197: val_accuracy did not improve from 0.91346 Epoch 198: val_accuracy improved from 0.91346 to 0.92308, saving model to best_model.h5 Epoch 199: val_accuracy did not improve from 0.92308 Epoch 200: val_accuracy did not improve from 0.92308 Epoch 201: val_accuracy did not improve from 0.92308 Epoch 202: val_accuracy did not improve from 0.92308 Epoch 203: val_accuracy improved from 0.92308 to 0.93269, saving model to best_model.h5 Epoch 204: val_accuracy improved from 0.93269 to 0.94231, saving model to best_model.h5 Epoch 205: val_accuracy improved from 0.94231 to 0.95192, saving model to best_model.h5 Epoch 206: val_accuracy did not improve from 0.95192 Epoch 207: val_accuracy did not improve from 0.95192 Epoch 208: val_accuracy did not improve from 0.95192 Epoch 209: val_accuracy did not improve from 0.95192 Epoch 210: val_accuracy did not improve from 0.95192 Epoch 211: val_accuracy improved from 0.95192 to 0.96154, saving model to best_model.h5 Epoch 212: val_accuracy improved from 0.96154 to 0.97115, saving model to best_model.h5 Epoch 213: val_accuracy did not improve from 0.97115 Epoch 214: val_accuracy did not improve from 0.97115 Epoch 215: val_accuracy did not improve from 0.97115 Epoch 216: val_accuracy did not improve from 0.97115 Epoch 217: val_accuracy did not improve from 0.97115 Epoch 218: val_accuracy did not improve from 0.97115 Epoch 219: val_accuracy did not improve from 0.97115 Epoch 220: val_accuracy did not improve from 0.97115 Epoch 221: val_accuracy did not improve from 0.97115 Epoch 222: val_accuracy did not improve from 0.97115 Epoch 223: val_accuracy did not improve from 0.97115 Epoch 224: val_accuracy did not improve from 0.97115 Epoch 225: val_accuracy did not improve from 0.97115 Epoch 226: val_accuracy did not improve from 0.97115 Epoch 227: val_accuracy did not improve from 0.97115 Epoch 228: val_accuracy did not improve from 0.97115 Epoch 229: val_accuracy did not improve from 0.97115 Epoch 230: val_accuracy did not improve from 0.97115 Epoch 231: val_accuracy did not improve from 0.97115 Epoch 232: val_accuracy did not improve from 0.97115 Epoch 233: val_accuracy did not improve from 0.97115 Epoch 234: val_accuracy did not improve from 0.97115 Epoch 235: val_accuracy did not improve from 0.97115 Epoch 236: val_accuracy did not improve from 0.97115 Epoch 237: val_accuracy did not improve from 0.97115 Epoch 238: val_accuracy did not improve from 0.97115 Epoch 239: val_accuracy did not improve from 0.97115 Epoch 240: val_accuracy did not improve from 0.97115 Epoch 241: val_accuracy did not improve from 0.97115 Epoch 242: val_accuracy did not improve from 0.97115 Epoch 243: val_accuracy did not improve from 0.97115 Epoch 244: val_accuracy did not improve from 0.97115 Epoch 245: val_accuracy did not improve from 0.97115 Epoch 246: val_accuracy did not improve from 0.97115 Epoch 247: val_accuracy did not improve from 0.97115 Epoch 248: val_accuracy did not improve from 0.97115 Epoch 249: val_accuracy did not improve from 0.97115 Epoch 250: val_accuracy did not improve from 0.97115
Let’s plot training history
plt.plot(history.history[‘loss’], label=’train’)
plt.plot(history.history[‘val_loss’], label=’valid’)
plt.legend()
plt.xlabel(‘Epochs’, fontsize=14)
plt.ylabel(‘Loss’, fontsize=14)
plt.title(‘Loss Curves’, fontsize=16)
plt.savefig(‘losscurvesfinal.png’)
plt.show()

Let’s plot final accuracy plot
plt.figure(figsize=[8,5])
plt.plot(history.history[‘accuracy’], label=’Train’)
plt.plot(history.history[‘val_accuracy’], label=’Valid’)
plt.legend()
plt.xlabel(‘Epochs’, fontsize=16)
plt.ylabel(‘Accuracy’, fontsize=16)
plt.title(‘Accuracy Curves’, fontsize=16)
plt.savefig(‘accuracycurvesfinal.png’)
plt.show()
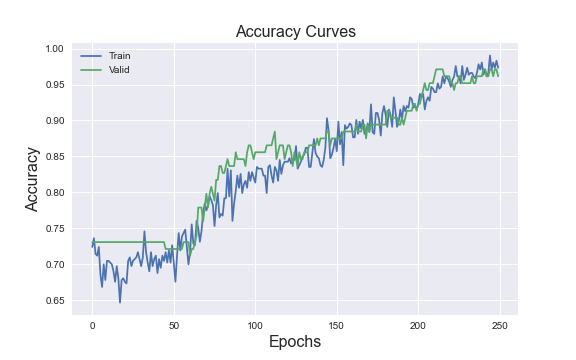
The final accuracy values are as follows
_, train_acc = model.evaluate(X_train, y_train, verbose=0)
_, valid_acc = model.evaluate(X_test, y_test, verbose=0)
print(‘Train: %.3f, Valid: %.3f’ % (train_acc, valid_acc))
Train: 0.985, Valid: 0.962
The final model is optimal in that it resolves a trade-off between training/test/validation data leakage/overfitting and overall precision. Thus, the ANN-based optimized early stopping with patience and model checkpoint method avoids data overfitting and yields the accuracy scores of the training and validation data of 98.5% and 96.2%, respectively.
Conclusions
The key success for controlling wildfires is the early detection and accurate prediction of these catastrophic phenomena. The present use-case study offers opportunities for the wildfire community to apply ANN-based deep learning techniques in early wildfire warning systems, hazard assessment and related risk management. This includes the EDA sequence that focuses on understanding the effects of the physical and environmental conditions such as temperature, wind, humidity and relative humidity on predicting the occurrence of fire in a given area. It was found that the final three-fold EDA+ANN+HPO workflow with an accuracy of prediction of 98% performed the best. The data for this project is taken from the NE Portugal Fire Dataset (courtsey of University of Minho, Portugal).


Leave a comment