The use of data analytics (DA) and artificial intelligence (AI) have been increasing in the pharmaceutical industry, including drug R&D, drug repurposing, improving pharmaceutical productivity, and clinical trials, among others. DA uses algorithms that can recognize patterns within a set of data to be further classified. Examples include drug recommendation, discovery and validation using knowledge graphs and Natural Language Processing (NLP) algorithms.
Leading biopharmaceutical companies’ belief in AI is due to growing awareness related to AI in the pharmaceutical sector and rising investment in drug development.
Contents:
- E2E Workflow
- Import Libraries
- Input Data
- Raw Data Cleaning
- Exploratory Data Analysis (EDA)
- Review Sentiments
- WordCloud Images
- Review Sentiment Editing
- Text Data Pre-Processing
- Sentiment Correlations
- Review Data Editing
- Feature Engineering
- NER Analysis
- Topic Modelling
- Train LDA
- Topic Correlations
- NLP Modelling
- BoW Vectorizer
- TF-IDF Vectorizer
- Word2Vec Vectorizer
- NLP Modelling
- Conclusion
- Appendix:
- Multi-Cloud NLP Drug Recommendation
Following the earlier DA-based study (cf. research paper and the source code), the objective of this project is to to build an AI-guided drug review system that recommends the most effective drug for a certain condition based on available reviews of various drugs used to treat this condition.
E2E Workflow
- Setup Jupyter notebook within the Anaconda IDE
- Import/install relevant Python libraries
- Download the Kaggle UCI ML Drug Review dataset
- Overview of the train/test dataset
- Reset the index after data concatenation
- Exploratory Data Analysis (EDA) using SNS plots
- Plot the WordCount images for positive/negative reviews
- Loading stop words from NLTK
- Text data NLP Pre-Processing (removing digits, extra spaces, lower case, etc.)
- nltk.sentiment.vader sentiment analysis using SentimentIntensityAnalyzer
- Adding the sentiment scores for reviews, preprocessed reviews as new features
- Feature Engineering – check the Pearson correlation matrix of various features
- Adding the word count, stopword count,char length, unique words count, mean word length, and puncation count
- Named entity recognition (NER) using spacy
- LDA topic modelling – prepare cleaned reviews
- Splitting the data into train, test and cross-validation (CV) datasets
- Encoding categorical, text and numerical features by applying LabelEncoder
- Vectorizing the cleaned reviews using BoW, TF-IDF (1 gram)
- Word2Vec Vectorization for reviews using pretrained glove model
- Compute the LDA confusion, precision, and recall matrices for the test data
Import Libraries
Let’s set the working directory YOURPATH
import os
os. getcwd()
os.chdir(‘YOURPATH’)
and import/install relevant Python libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings(‘ignore’)
from wordcloud import WordCloud
from wordcloud import STOPWORDS
import nltk
import regex as re
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
import re
from tqdm import tqdm
from nltk.corpus import stopwords
from nltk.sentiment.vader import SentimentIntensityAnalyzer
import string
import spacy
from tqdm import tqdm
import gensim
Input Data
Let’s read the Kaggle train and test data as csv files
data_train = pd.read_csv(‘drugsComTrain_raw.csv’)
data_test = pd.read_csv(‘drugsComTest_raw.csv’)
and check the dataset structure
print(‘Size of Train dataset is:’,data_train.shape)
print(‘Size of Test dataset is:’,data_test.shape)
Size of Train dataset is: (161297, 7) Size of Test dataset is: (53766, 7)
print(‘Columns of the dataset are:\n’,data_train.columns)
Columns of the dataset are:
Index(['uniqueID', 'drugName', 'condition', 'review', 'rating', 'date',
'usefulCount'],
dtype='object')
print(‘Overview of Train dataset:\n’)
data_train.head(10)
Overview of Train dataset:
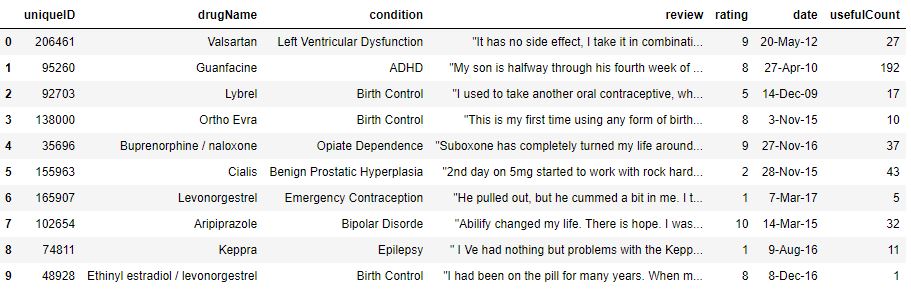
Similarly, let’s print the test Kaggle dataset
print(‘Overview of Test dataset:\n’)
data_test.head(10)
Overview of Test dataset:

Let’s concatenate the two datasets
data = pd.concat([data_train,data_test])
print(‘The size of the combined data is:’,data.shape)
The size of the combined data is: (215063, 7)
while resetting the index after concatenation
data.reset_index(inplace=True,drop=True)
data.tail()
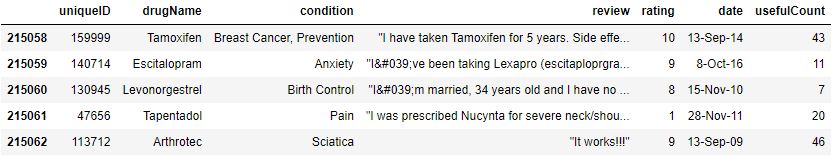
Let’s verify data types
data.dtypes
uniqueID int64 drugName object condition object review object rating int64 date object usefulCount int64 dtype: object
The descriptive statistics of numerical features is given by
data.describe()
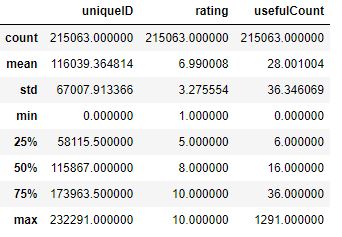
and the data info is
data.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 215063 entries, 0 to 215062 Data columns (total 7 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 uniqueID 215063 non-null int64 1 drugName 215063 non-null object 2 condition 213869 non-null object 3 review 215063 non-null object 4 rating 215063 non-null int64 5 date 215063 non-null object 6 usefulCount 215063 non-null int64 dtypes: int64(3), object(4) memory usage: 11.5+ MB
Raw Data Cleaning
Let’s check the above data for null values
data.isnull().any()
uniqueID False drugName False condition True review False rating False date False usefulCount False dtype: bool
and the percentage of null values is
null_size = data.isnull().sum()[‘condition’]
print(‘Total null values are:’,null_size)
data_size = data.shape[0]
print(‘Percentage of null values are:’,(null_size/data_size)*100)
Total null values are: 1194 Percentage of null values are: 0.5551861547546533
Let’ drop the entire rows with null values
data = data.dropna(axis=0)
print(‘Size of the dataset after dropping null values:’,data.shape)
Size of the dataset after dropping null values: (213869, 7)
and check for number of unique conditions
print(‘Number of unique conditions are:’,data[‘condition’].unique().shape[0])
Number of unique conditions are: 916
Exploratory Data Analysis (EDA)
Referring to the previous DA analysis, let’s apply EDA to our data after dropping null values.
Let’s plot the top 10 conditions
conditions = dict(data[‘condition’].value_counts())
top_conditions = list(conditions.keys())[0:10]
values = list(conditions.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=top_conditions,y=values,palette=’spring’)
plt.title(‘Top 10 Conditions’)
plt.xlabel(‘Conditions’)
plt.ylabel(‘Count’)
plt.xticks(fontsize=11)
plt.yticks(fontsize=11)
b.set_xlabel(“Conditions”,fontsize=20)
b.set_ylabel(“Count”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Top 10 Conditions”,fontsize=20)
plt.show()
plt.savefig(‘drugbartop10conditions.png’)

It is clear that Birth Control is the most frequently occurring condition in our dataset, followed by Depression, Pain, Anxiety, etc.
Let’s count the number of drugs provided or prescribed as a treatment for top 10 conditions
val=[]
for c in list(conditions.keys()):
val.append(data[data[‘condition’]==c][‘drugName’].nunique())
drug_cond = dict(zip(list(conditions.keys()),val))
Let’s plot the number of drugs provided or prescribed as a treatment for top 10 conditions
top_conditions = list(drug_cond.keys())[0:10]
values = list(drug_cond.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=top_conditions,y=values,palette=’spring’)
plt.title(‘Number of Drugs for each Top 10 Conditions’)
plt.xlabel(‘Conditions’)
plt.ylabel(‘Count of Drugs’)
b.set_xlabel(“Count of Patients used”,fontsize=20)
b.set_ylabel(“Drug Names”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Number of Drugs for each Top 10 Conditions”,fontsize=20)
plt.show()
plt.savefig(‘drugbarnumberofdrugs.png’)
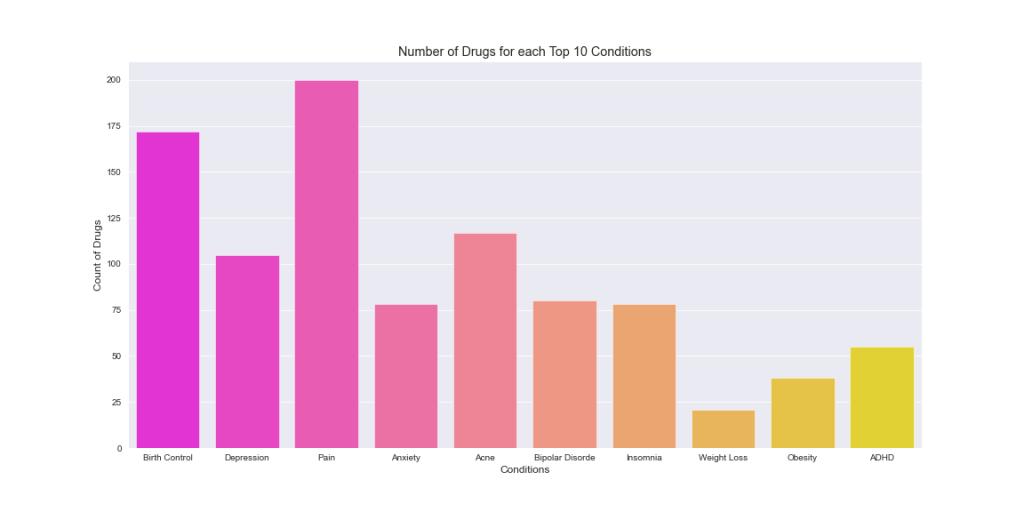
This plot shows that multiple drugs are normally used as a treatment for top 10 conditions. It seems that there will likely be no “silver bullet” medication for Pain and Birth Control conditions.
Let’s see what is the most frequently used Birth Control drug by plotting top 10 drugs used as a treatment for this particular condition
drugs_birth = dict(data[data[‘condition’]==’Birth Control’][‘drugName’].value_counts())
top_drugs = list(drugs_birth.keys())[0:10]
values = list(drugs_birth.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=values,y=top_drugs,palette=’spring’)
plt.title(‘Top 10 Drugs used for Birth Control’)
plt.ylabel(‘Drug Names’)
plt.xlabel(‘Count of Patients used’)
b.set_xlabel(“Count of Patients used”,fontsize=20)
b.set_ylabel(“Drug Names”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Top 10 Drugs used for Birth Control”,fontsize=20)
plt.show()
plt.savefig(‘drugbarnumbermostuseddrugs.png’)

It is clear that Etonogestrel is the most frequently used Birth Control drug, followed by Ethinyl estradiol, Levonorgestrel and Nexplanon.
Similarly, we can plot top 10 drugs used as a treatment for Pain
drugs_pain = dict(data[data[‘condition’]==’Pain’][‘drugName’].value_counts())
top_drugs = list(drugs_pain.keys())[0:10]
values = list(drugs_pain.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=values,y=top_drugs,palette=’spring’)
plt.title(‘Top 10 Drugs used for Pain’)
plt.ylabel(‘Drug Names’)
plt.xlabel(‘Count of Patients used’)
b.set_xlabel(“Count of Patients used”,fontsize=20)
b.set_ylabel(“Drug Names”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Top 10 Drugs used for Pain”,fontsize=20)
plt.show()
plt.savefig(‘drugbarnumbermostusedpain.png’)
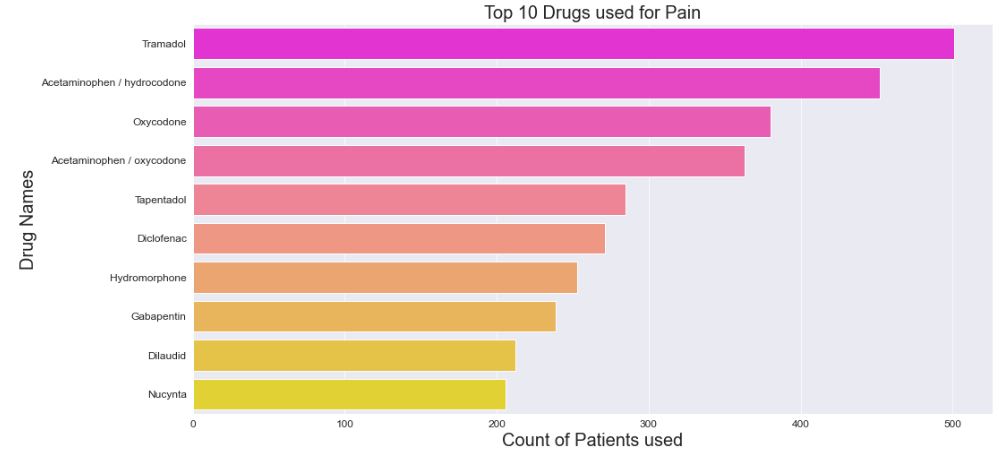
We can see that Tramadol is the most frequently used Pain drug.
Let’s plot the top 10 drugs rated as 10
drugs_rating = dict(data[data[‘rating’]==10][‘drugName’].value_counts())
top_drugs = list(drugs_rating.keys())[0:10]
values = list(drugs_rating.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=values,y=top_drugs,palette=’spring’)
plt.title(‘Top 10 Drugs rated as 10’)
plt.ylabel(‘Drug Names’)
plt.xlabel(‘Count of Ratings’)
b.set_xlabel(“Count of Ratings”,fontsize=20)
b.set_ylabel(“Drug Names”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Top 10 Drugs rated as 10”,fontsize=20)
plt.show()
plt.savefig(‘drugbartop10rate10.png’)

It appears that Birth Control drugs are top rated, so Etonogestrel and Levonorgestrel should be the top recommended drugs as they are (a) most frequently used and (b) rated as 10 by patients.
Likewise, let’s plot the top 10 drugs rated as 1
drugs_rating = dict(data[data[‘rating’]==1][‘drugName’].value_counts())
top_drugs = list(drugs_rating.keys())[0:10]
values = list(drugs_rating.values())[0:10]
plt.figure(figsize=(16,8))
sns.set_style(style=’darkgrid’)
b=sns.barplot(x=values,y=top_drugs,palette=’spring’)
plt.title(‘Top 10 Drugs rated as 1’)
plt.ylabel(‘Drug Names’)
plt.xlabel(‘Count of Ratings’)
b.set_xlabel(“Count of Ratings”,fontsize=20)
b.set_ylabel(“Drug Names”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Top 10 Drugs rated as 1”,fontsize=20)
plt.show()
plt.savefig(‘drugbartop10rate1.png’)
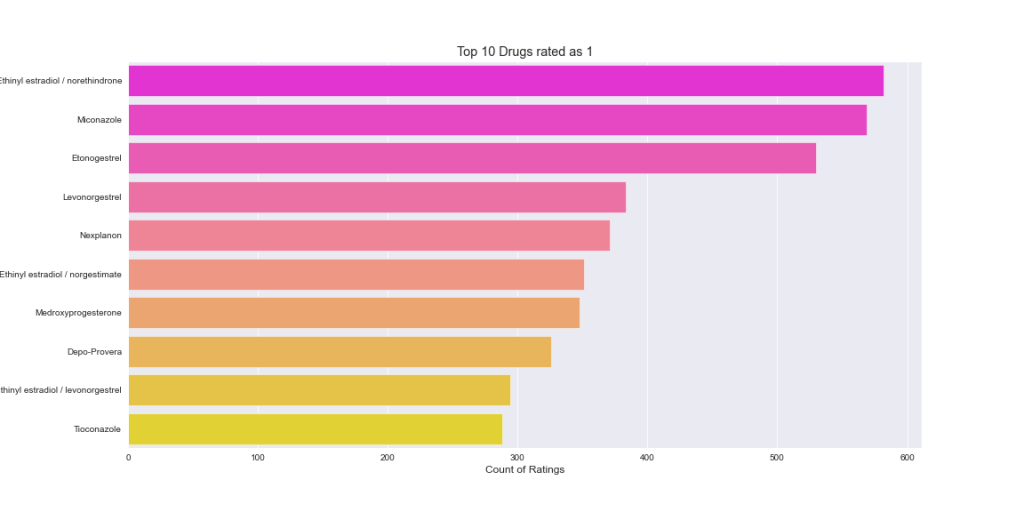
One can see that Levonorgestrel and Etonogestrel are among top 10 most frequently used drugs that have both ratings ‘10’ and ‘1’. The lowest rating may imply that these two drugs had side effects and/or not effective for certain patients.
Let’s plot the density distributions of ratings 1-10
f,ax = plt.subplots(1,2,figsize=(16,8))
ax1= sns.histplot(data[‘rating’],ax=ax[0])
ax1.set_title(‘Count of Ratings’)
ax2= sns.distplot(data[‘rating’],ax=ax[1],color=”r”)
ax2.set_title(‘Distribution of Ratings density’)
ax2.axes.set_title(“Distribution of Ratings Density”,fontsize=20)
ax2.set_xlabel(“Rating”,fontsize=20)
ax2.set_ylabel(“Count”, fontsize=20)
ax2.tick_params(labelsize=12)
ax1.set_xlabel(“Rating”,fontsize=20)
ax1.set_ylabel(“Count”, fontsize=20)
ax1.tick_params(labelsize=12)
ax1.axes.set_title(“Count of Ratings”,fontsize=20)
plt.show()
plt.savefig(‘drugdistplotratingsdens.png’)

We can see that rating 10 has the maximum density or frequency count.
Let’s look at the percentage distribution of ratings using pie chart
ratings_count = dict(data[‘rating’].value_counts())
count = list(ratings_count.values())
labels = list(ratings_count.keys())
plt.figure(figsize=(18,9))
plt.pie(count,labels=labels, autopct=’%1.1f%%’,textprops={‘fontsize’: 14})
plt.title(‘Pie Chart Representation of Ratings’,fontsize=20)
plt.legend(title=’Ratings’,fontsize=12)
plt.show()
plt.savefig(‘drugspiechartratings.png’)
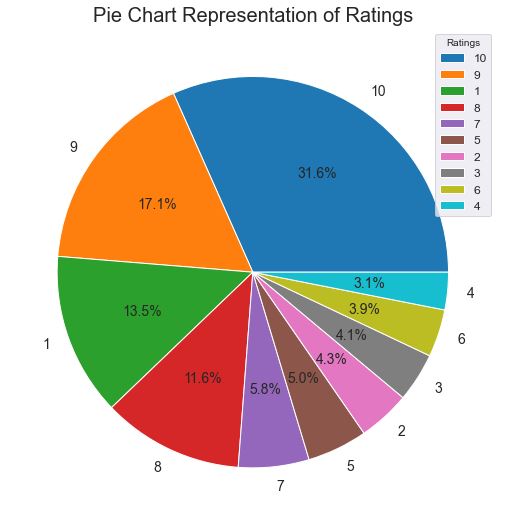
We can see that 31.6% and 13.5% drugs have ratings 10 and 1, respectively.
Let’s change the date format using to_datetime
data[‘date’]= pd.to_datetime(data[‘date’])
and count ratings per year
year_ratings = dict(data[‘date’].dt.year.value_counts())
years = list(year_ratings.keys())
values = list(year_ratings.values())
plt.figure(figsize=(18,9))
b=sns.barplot(x=years,y=values,palette=’spring’)
plt.xlabel(‘Years’)
plt.ylabel(‘Count of Ratings’)
plt.title(‘Count of Ratings in each Year’)
b.set_xlabel(“Year”,fontsize=20)
b.set_ylabel(“Count of Ratings”, fontsize=20)
b.tick_params(labelsize=12)
b.axes.set_title(“Count of Ratings in each Year”,fontsize=20)
plt.show()
plt.savefig(‘drugsbarratingsyear.png’)
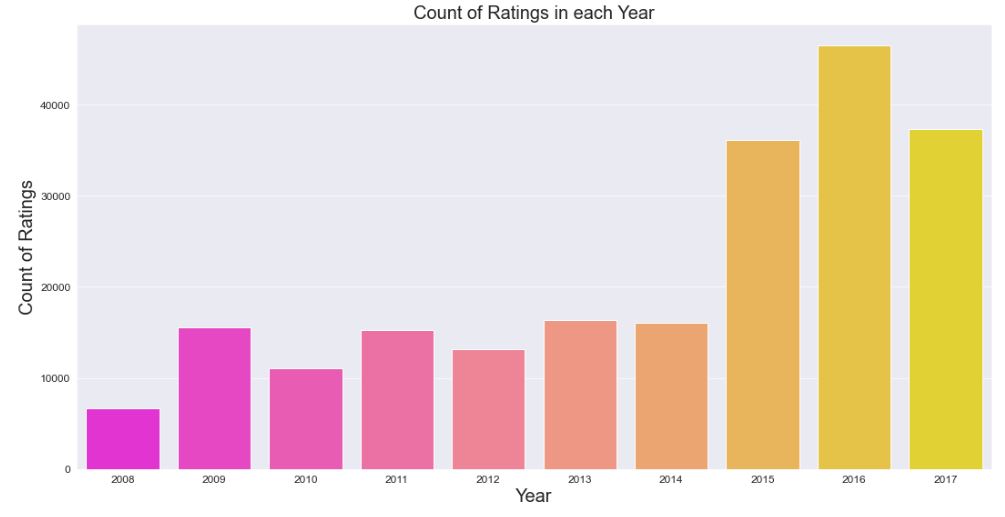
Notably drug ratings achieved their over-4000 peak in 2016.
Let’s check the distribution of usefulCount
plt.figure(figsize=(16,8))
plt.xlim(0, 200)
ax1 =sns.distplot(data[‘usefulCount’],color=’r’,bins=256)
ax1.set_xlabel(“usefulCount”,fontsize=20)
ax1.set_ylabel(“Density”, fontsize=20)
ax1.tick_params(labelsize=12)
plt.title(‘Distribution of usefulCount’,fontsize=20)
plt.show()
plt.savefig(‘drugdistusefulcount.png’)
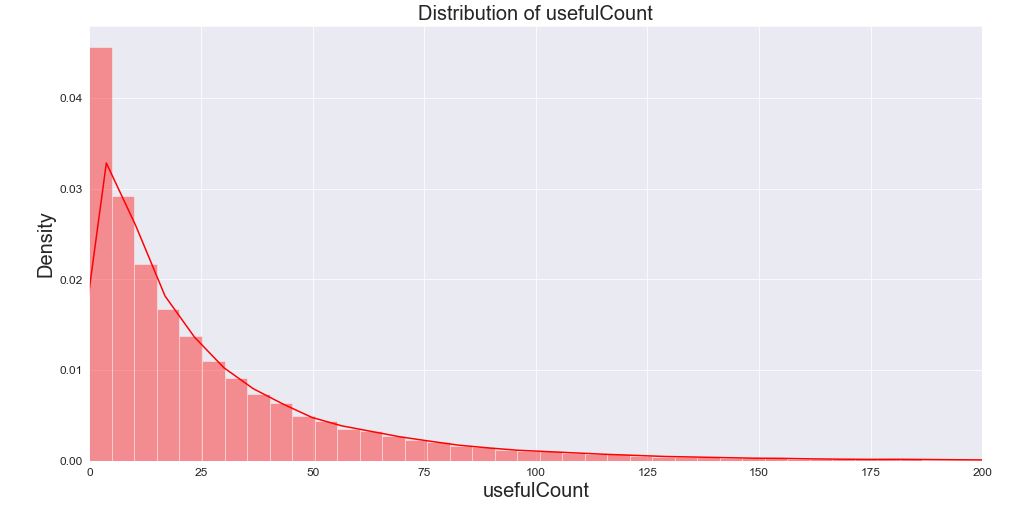
It is clear that there that the feature usefulCount has the limit of 200.
Review Sentiments
Let’s create the target feature review_sediment using ratings
data[‘review_sentiment’] = data[‘rating’].apply(lambda x: 1 if x > 5 else 0)
Here 1 and 0 represent positive and negative reviews, respectively.
data.head(10)
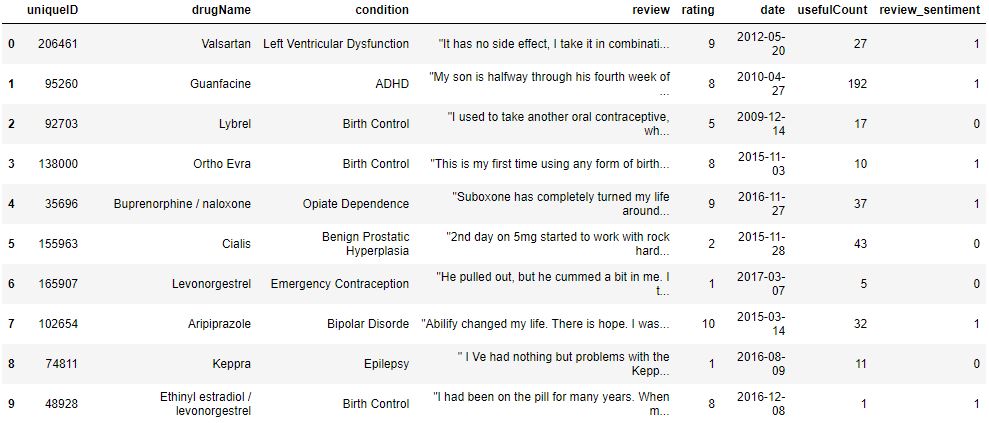
Let’s plot the pie chart for review sentiments
plt.figure(figsize=(14,7))
plt.pie(data[‘review_sentiment’].value_counts(),labels=[‘Positive’,’Negative’],autopct=’%1.1f%%’,textprops={‘fontsize’: 16})
plt.title(‘Pie Chart Representation of Review Sentiment’,fontsize=20)
plt.show()
plt.savefig(‘drugspiechartratings.png’)
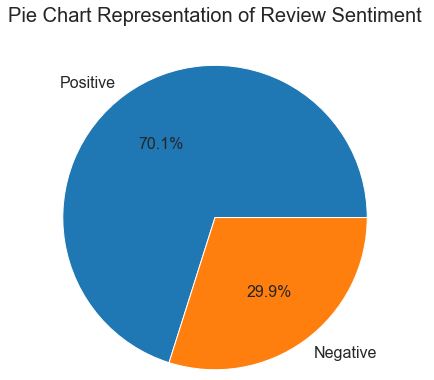
According to this chart, 70.1% of patients are likely to give positive reviews. Hence, this is an imbalanced dataset.
WordCloud Images
Let’s turn our attention to WordCloud – a technique for visualising frequent words in a text where the size of the words represents their frequency.
Let’s look at WordCloud for positive reviews
positive_reviews = ” “.join([review for review in data[‘review’][data[‘review_sentiment’] == 1]])
stop_words = set(STOPWORDS)
wordcloud = WordCloud(width = 1200, height = 800,background_color =’white’,stopwords = stop_words,min_font_size = 10).generate(positive_reviews)
while plotting the WordCloud image
plt.figure(figsize = (12, 8), facecolor = None)
plt.imshow(wordcloud)
plt.title(‘WordCloud for Positive Reviews’)
plt.axis(“off”)
plt.tight_layout(pad = 0)
plt.show()
plt.savefig(‘drugswordcloudpos.png’)

Similarly, we can apply WordCloud to negative reviews
negative_reviews = ” “.join([review for review in data[‘review’][data[‘review_sentiment’] == 0]])
wordcloud = WordCloud(width = 1200, height = 800,background_color =’white’,stopwords = stop_words,min_font_size = 10).generate(negative_reviews)
and plot the resulting image
plt.figure(figsize = (12, 8), facecolor = None)
plt.imshow(wordcloud)
plt.title(‘WordCloud for Negative Reviews’)
plt.axis(“off”)
plt.tight_layout(pad = 0)
plt.show()
plt.savefig(‘drugswordcloudneg.png’)
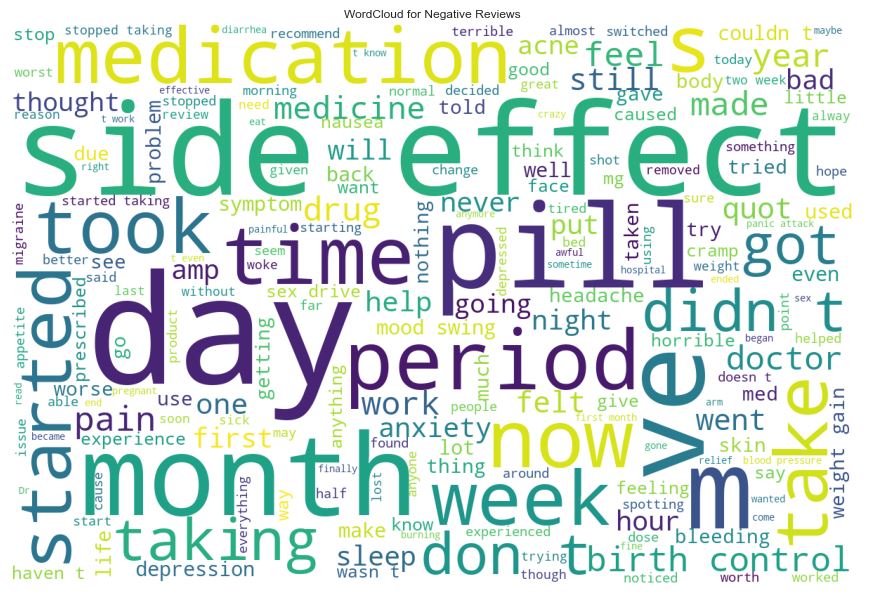
We can see that both positive and negative reviews contain most frequent words “side effect”, “day”, and “period”.
Review Sentiment Editing
Let’s remove some unwanted conditions in the review sentiment data
del_index = []
conds =[]
for c in data[‘condition’]:
if (‘helpful’ in c) or (‘Listed’ in c):
f= list(data[data[‘condition’]==c].index)
del_index.extend(f)
conds.append(c)
print(‘Size of the data before removing the conditions:’,data.shape)
Size of the data before removing the conditions: (213869, 8)
print(‘The removable conditions count is:’,len(conds))
The removable conditions count is: 1763
data.drop(del_index,inplace=True)
print(‘Size of the data after dropping the condtions:’,data.shape)
Size of the data after dropping the condtions: (212106, 8)
data.reset_index(inplace=True,drop=True)
data.tail()
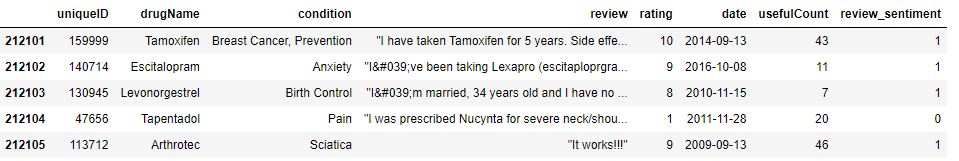
Text Data Pre-Processing
We need to invoke several functions to clean and preprocess the review text data by removing html tags, punctuations, special characters, stopwords, tabs, new lines, spaces and also stemming of words:
def decontracted(phrase):
# specific
phrase = re.sub(r”won’t”, “will not”, phrase)
phrase = re.sub(r”can\’t”, “can not”, phrase)
# general
phrase = re.sub(r"n\'t", " not", phrase)
phrase = re.sub(r"\'re", " are", phrase)
phrase = re.sub(r"\'s", " is", phrase)
phrase = re.sub(r"\'d", " would", phrase)
phrase = re.sub(r"\'ll", " will", phrase)
phrase = re.sub(r"\'t", " not", phrase)
phrase = re.sub(r"\'ve", " have", phrase)
phrase = re.sub(r"\'m", " am", phrase)
return phrase
def preprocess_text(text_data):
text_data = decontracted(text_data)
text_data = text_data.replace('\n',' ')
text_data = text_data.replace('\r',' ')
text_data = text_data.replace('\t',' ')
text_data = text_data.replace('-',' ')
text_data = text_data.replace("/",' ')
text_data = text_data.replace(">",' ')
text_data = text_data.replace('"',' ')
text_data = text_data.replace('?',' ')
return text_data
Let’s load stop words from the nltk library
stop_words = set(stopwords.words(‘english’))
stemmer = SnowballStemmer(‘english’)
Let’s remove ‘no’ from the stop words list due to the importance of ‘side effects’ and ‘no side effects’ in reviews
stop_words.remove(‘no’)
Let’s define the functional block that process the text data by removing digits, extra spaces, stop words, while converting words to lower case and stemming words
def nlp_preprocessing(review):
if type(review) is not int:
string = “”
review = preprocess_text(review)
review = re.sub(‘[^a-zA-Z]’, ‘ ‘, review)
review = re.sub('\s+',' ', review)
review = review.lower()
for word in review.split():
if not word in stop_words:
word = stemmer.stem(word)
string += word + " "
return string
Let’s apply the NLP processing
data[‘cleaned_review’] = data[‘review’].apply(nlp_preprocessing)
and convert the content of drugName and condition to lower case
data[‘drugName’] = data[‘drugName’].apply(lambda x:x.lower())
data[‘condition’] = data[‘condition’].apply(lambda x:x.lower())
data.head()
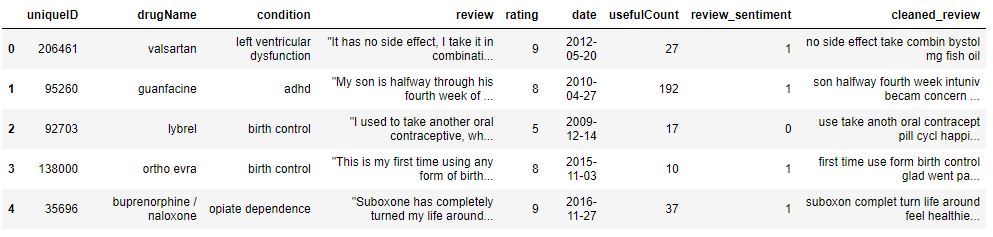
data.shape
(212106, 9)
Let’s add the sentiment scores for reviews and preprocessed reviews as new features using SentimentIntensityAnalyzer
sid = SentimentIntensityAnalyzer()
data[‘sentiment_score’] = [sid.polarity_scores(v)[‘compound’] for v in data[‘review’]]
data[‘sentiment_score_clean’] = [sid.polarity_scores(v)[‘compound’] for v in data[‘cleaned_review’]]
data.head()

These text features and sentiment scores are the basic feature extractions for the text data classification problem.
Sentiment Correlations
Let’s check the crrelation of features present in the sentiment data
data.corr()
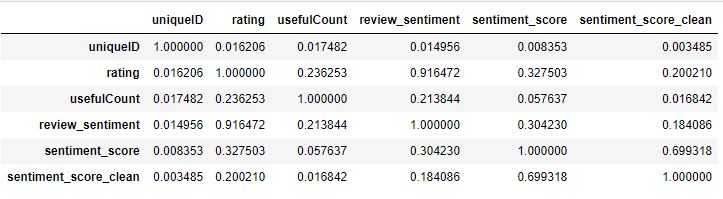
Let’s plot the Pearson’s correlation heatmap lower triangle with Seaborn
corr_data = data.corr(method=’pearson’)
mask_ut=np.triu(np.ones(corr_data.shape)).astype(np.bool)
f, ax = plt.subplots(figsize=(11, 9))
sns.set(font_scale=1)
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(corr_data, mask=mask_ut, cmap=cmap,annot=True, annot_kws={“size”:16})
hmap.figure.savefig(“Correlation_Heatmap_Lower_Triangle_with_Seaborn.png”,
format=’png’,
dpi=150)

Let’s save the data for future reference
data.to_csv(‘new_data_processed.csv’,index=False)
Review Data Editing
Let’s check the content of data
data.head(10)
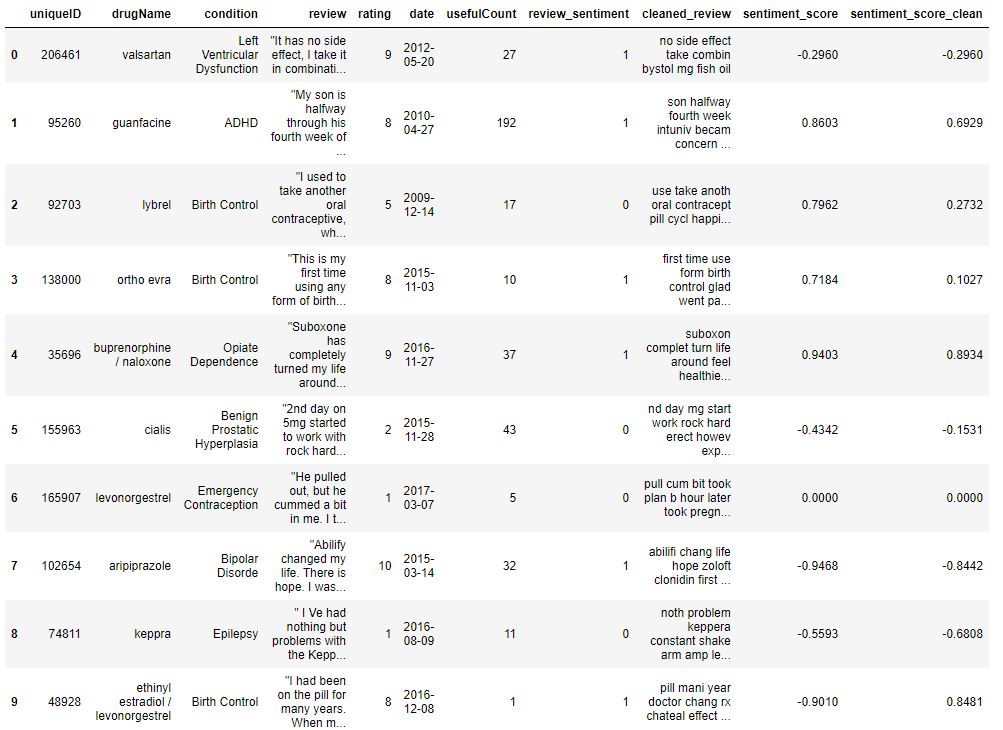
checking for any nan values in cleaned_review feature
data[‘cleaned_review’].isna().sum()
0
and droping all rows containing nan values
print(‘The data size before:’,data.shape)
data = data.dropna(axis=0)
data.reset_index(inplace=True,drop=True)
print(‘The data size after dropping:’,data.shape)
The data size before: (212106, 11) The data size after dropping: (212106, 11)
We can see that there are no nan values in our processed dataset
Let’s add the year as feature and check the data structure
data[‘date’] = pd.to_datetime(data[‘date’])
data[‘year’] = data[‘date’].dt.year
data.head(10)
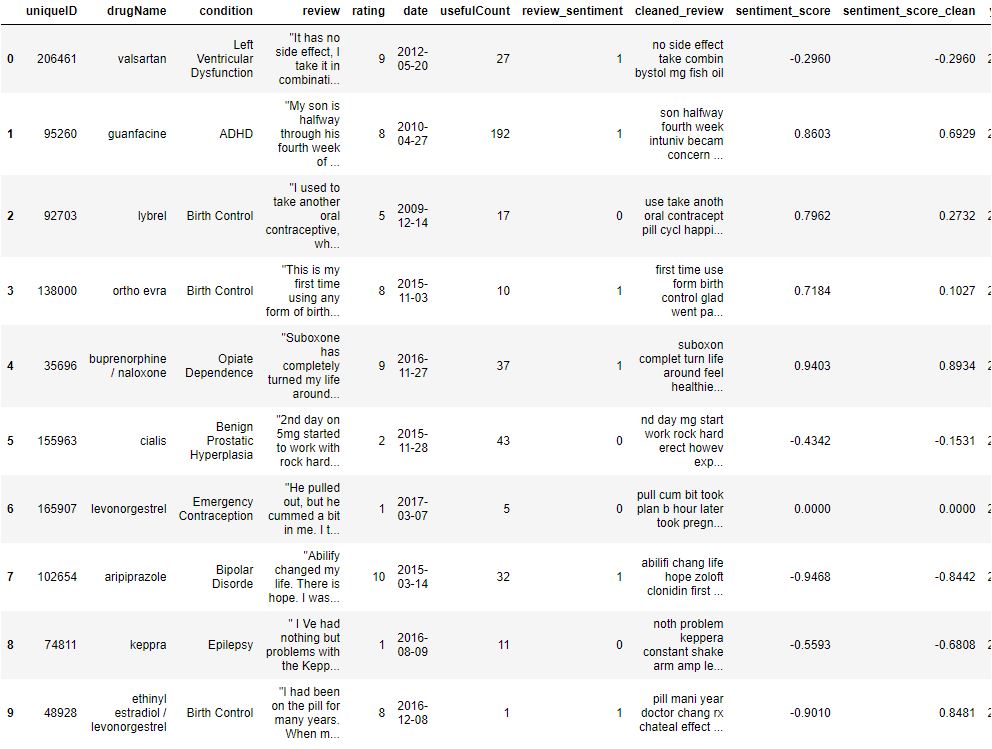
Let’s add the word count, stopword count, char length, unique words count, mean word length, and puncation count
stop_words = set(stopwords.words(‘english’))
data[‘word_count’]=data[“cleaned_review”].apply(lambda x: len(str(x).split()))
data[‘unique_word_count’]=data[“cleaned_review”].apply(lambda x: len(set(str(x).split())))
data[‘char_length’]=data[“cleaned_review”].apply(lambda x: len(str(x)))
data[“count_punctuations”] = data[“review”].apply(lambda x: len([c for c in str(x) if c in string.punctuation]))
data[“stopword_count”] = data[“review”].apply(lambda x: len([w for w in str(x).lower().split() if w in stop_words]))
data[“mean_word_len”] = data[“cleaned_review”].apply(lambda x: np.mean([len(w) for w in str(x).split()]))
Let’s check the updated content
data.head(10)
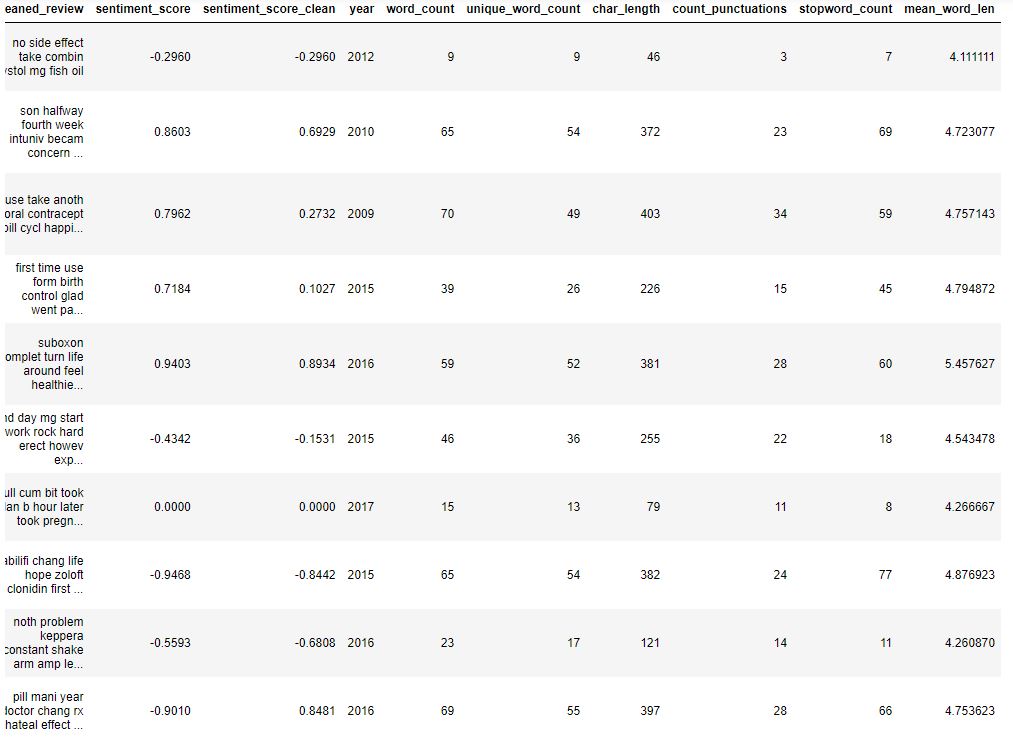
Feature Engineering
Let’s compute data correlations
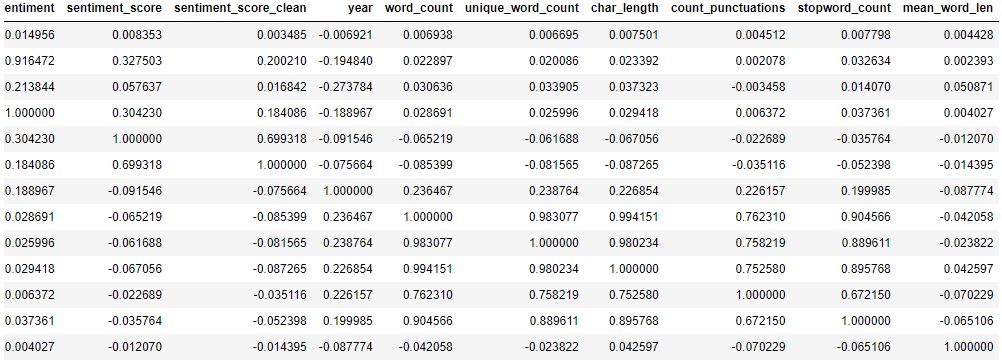
Let’s compute the correlation heatmap lower triangle with seaborn
corr1_data = data.corr(method=’pearson’)
mask_ut=np.triu(np.ones(corr1_data.shape)).astype(np.bool)
f, ax = plt.subplots(figsize=(11, 9))
sns.set(font_scale=1)
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(corr1_data, mask=mask_ut, cmap=”Spectral”,annot=True, annot_kws={“size”:12})
hmap.figure.savefig(“Correlation_Heatmap_feature_Lower_Triangle_with_Seaborn.png”,
format=’png’,
dpi=150)
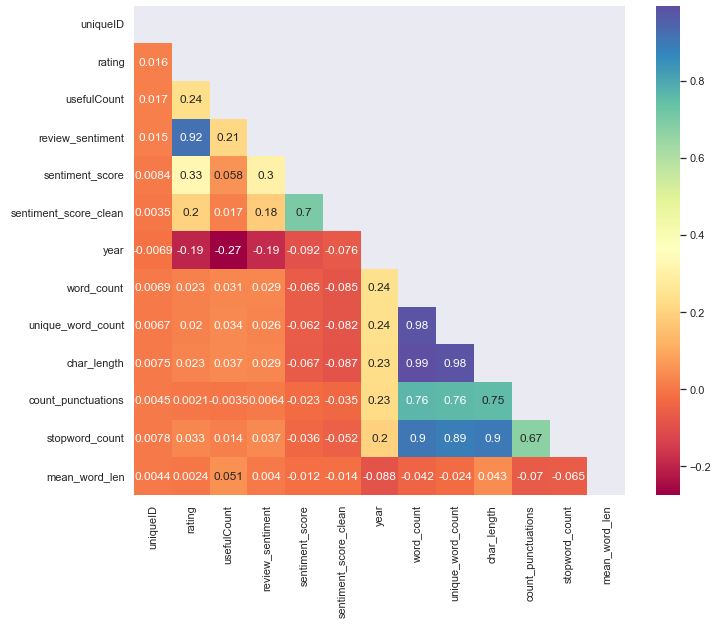
NER Analysis
Let’s perform Named Entity Recognition (NER) using spacy
nlp = spacy.load(“en_core_web_sm”)
def subj_obj_count(review):
sent = review
doc=nlp(sent)
sub_words = set([str(word) for word in doc if (word.dep_ == "nsubj")])
obj_words = set([str(word) for word in doc if (word.dep_ == "dobj")])
return len(sub_words),len(obj_words)
count = []
for r in tqdm(data[‘review’]):
count.append(subj_obj_count(r))
100%|██████████| 212106/212106 [41:58<00:00, 84.22it/s]
sub_obj = pd.DataFrame(count,columns=[‘subj_count’,’obj_count’])
sub_obj.head(10)

sub_obj.to_csv(‘sub_obj.csv’,index=False)
sub_obj = pd.read_csv(‘sub_obj.csv’)
sub_obj.shape
(212106, 2)
ner_lst = nlp.pipe_labels[‘ner’]
def ner(review):
sent = review
doc=nlp(sent)
dic = {}.fromkeys(ner_lst,0)
for word in doc.ents:
dic[word.label_]+=1
return dic
entity = pd.DataFrame([ner(r) for r in tqdm(data[‘cleaned_review’])])
100%|██████████| 212106/212106 [33:50<00:00, 104.47it/s]
entity.to_csv(‘entities.csv’,index=False)
entity = pd.read_csv(‘entities.csv’)
print(entity.shape)
entity.head(10)
(212106, 18)
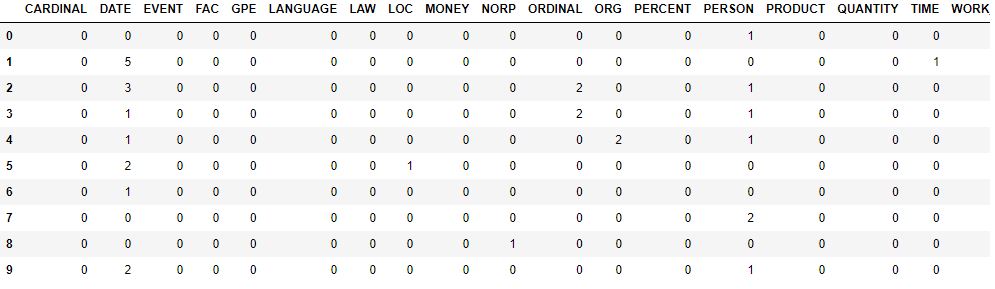
Topic Modelling
Topic Modelling in NLP analyzes the text data and finds the topics that best describes the set of documents. It is an Unsupervised approach as it recognizes and extracts the topics by detecting hidden patterns like clustering algorithms. Also it doesn’t require any predefined tags or training data.
Let’s pre-process the feature
corpus = data[‘cleaned_review’]
lst_corpus = []
for string in tqdm(corpus):
lst_words = string.split()
lst_grams = [” “.join(lst_words[i:i + 1]) for i in range(0, len(lst_words), 1)]
lst_corpus.append(lst_grams)
and map words to an id
id2word = gensim.corpora.Dictionary(lst_corpus)
while creating the dictionary word:freq
dic_corpus = [id2word.doc2bow(word) for word in lst_corpus]
Train LDA
Let’s train the LDA model
lda_model = gensim.models.ldamodel.LdaModel(corpus=dic_corpus, id2word=id2word, num_topics=20, chunksize=100, passes=10, alpha=’auto’, per_word_topics=True)
100%|██████████| 212106/212106 [00:02<00:00, 74286.58it/s]
storing the topic vectors for each review in a list
train_vecs = []
for i in range(len(corpus)):
top_topics = (
lda_model.get_document_topics(dic_corpus[i],
minimum_probability=0.0)
)
topic_vec = [top_topics[i][1] for i in range(20)]
train_vecs.append(topic_vec)
topics = pd.DataFrame(train_vecs)
print(topics.shape)
topics.head(10)
(212106, 20)
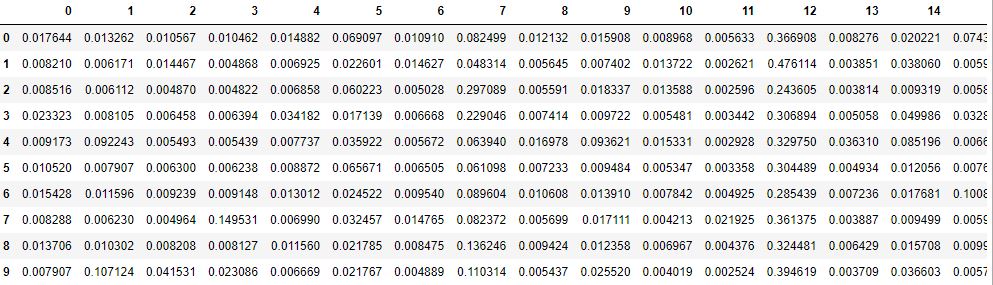
Let’s export this table to csv
topics.to_csv(‘topics.csv’,index=False)
Topic Correlations
Let’s read the topics data
topics = pd.read_csv(‘topics.csv’)
topics.shape
(212106, 20)
while concatenating sub_obj, entity and topics
data = pd.concat([data,sub_obj,entity,topics],axis=1)
print(data.shape)
data.tail(10)
(212106, 58)
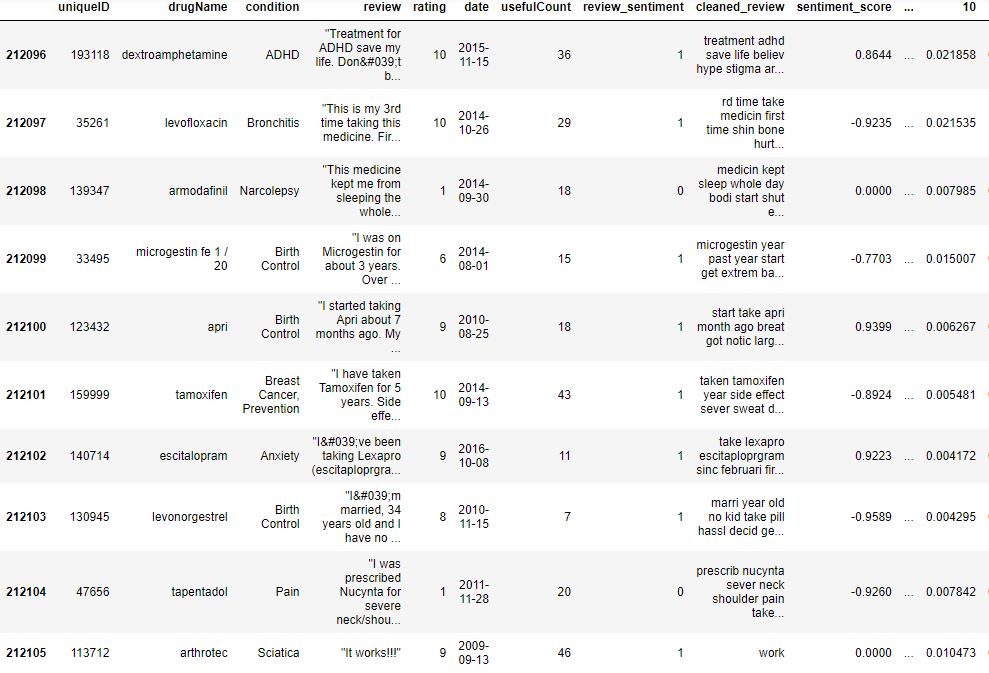
Let’s calculate correlations as 53 rows × 53 columns
data.corr()

Let’s plot the corresponding correlation heatmap lower triangle with seaborn
corr2_data = data.corr(method=’pearson’)
mask_ut=np.triu(np.ones(corr2_data.shape)).astype(np.bool)
f, ax = plt.subplots(figsize=(22, 18))
sns.set(font_scale=1)
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(corr2_data, mask=mask_ut, cmap=”Spectral”,annot=True, annot_kws={“size”:6})
hmap.figure.savefig(“Correlation_Heatmap_topics_Lower_Triangle_with_Seaborn.png”,
format=’png’,
dpi=150)
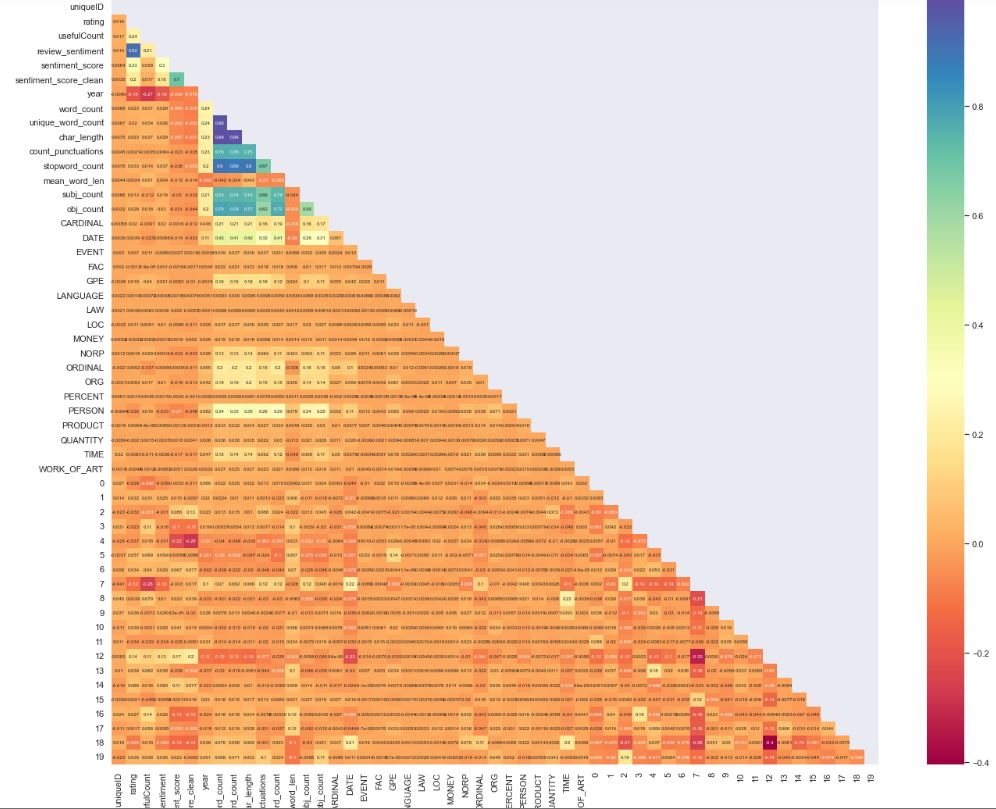
Let’s export the newly processed data
data.to_csv(‘final_new_data_processed.csv’,index=False)
NLP Modelling
Finally, we are ready to build training models for classifying reviews. This would require the following libraries:
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings(‘ignore’)
from wordcloud import WordCloud
from wordcloud import STOPWORDS
import nltk
import regex as re
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
Let’s read the data
data = pd.read_csv(‘final_new_data_processed.csv’)
print(data.shape)
data.head(10)
(212106, 58)
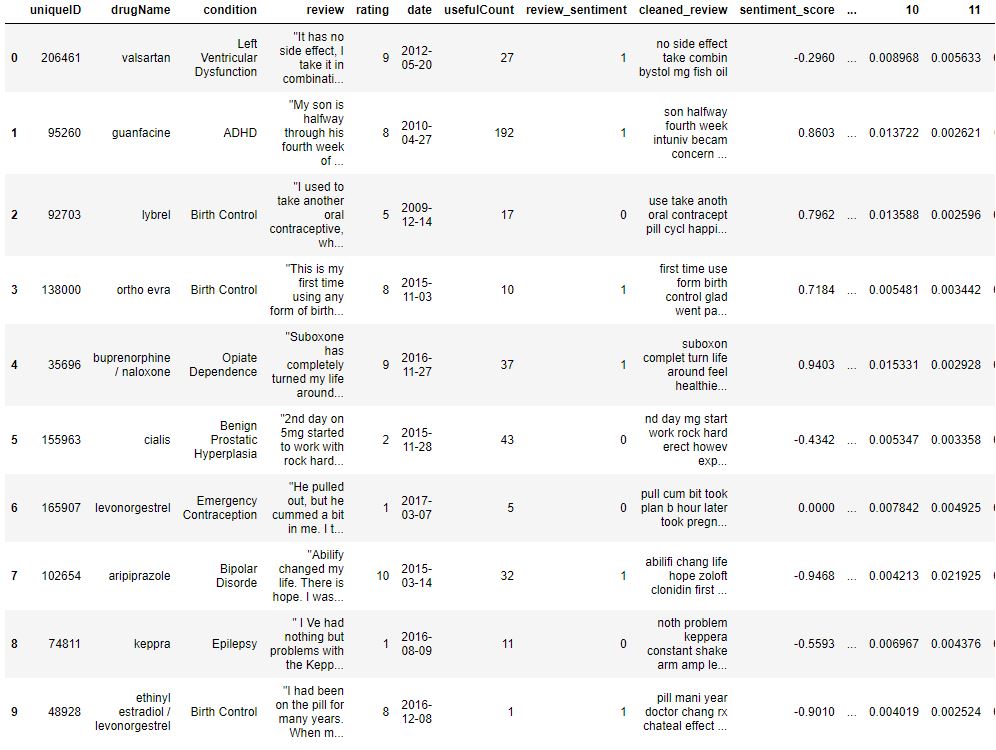
Let’s drop 5 unwanted columns
X = data.drop([‘uniqueID’,’review’,’rating’,’date’,’review_sentiment’],axis=1)
y = data[‘review_sentiment’].values
print(X.shape)
(212106, 53)
Let’s split our data into the 70% training and 30% test datasets while creating 21% cross-validation (CV) data using train_test_split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,stratify=y,test_size=0.30,random_state=42)
X_train, X_cv, y_train, y_cv = train_test_split(X_train, y_train,stratify=y_train,test_size=0.30,random_state=42)
print(‘Train data size is:’,X_train.shape)
print(‘Cross Validation data size is:’,X_cv.shape)
print(‘Test data size is:’,X_test.shape)
Train data size is: (103931, 53) Cross Validation data size is: (44543, 53) Test data size is: (63632, 53)
Let’s perform encoding Categorical, text and numerical features using LabelEncoder
from sklearn.preprocessing import LabelEncoder
lab_enc_cond = LabelEncoder()
lab_enc_cond.fit(X[‘condition’].values)
X_train_condition = lab_enc_cond.transform(X_train[‘condition’].values).reshape(-1,1)
X_test_condition = lab_enc_cond.transform(X_test[‘condition’].values).reshape(-1,1)
X_cv_condition = lab_enc_cond.transform(X_cv[‘condition’].values).reshape(-1,1)
print(‘After Encoding’)
print(‘Train data shape’,X_train_condition.shape)
print(‘Test data shape’,X_test_condition.shape)
print(‘CV data shape’,X_cv_condition.shape)
After Encoding Train data shape (103931, 1) Test data shape (63632, 1) CV data shape (44543, 1)
Let’s export the condition encoder
import joblib
print(‘Saving condition encoder..’)
joblib.dump(lab_enc_cond,’condition_encoder.pkl’)
Saving condition encoder..
Out[95]:
['condition_encoder.pkl']
Let’s apply label encoder
lab_enc_year = LabelEncoder()
lab_enc_year.fit(X[‘year’].values)
X_train_year = lab_enc_year.transform(X_train[‘year’].values).reshape(-1,1)
X_test_year = lab_enc_year.transform(X_test[‘year’].values).reshape(-1,1)
X_cv_year = lab_enc_year.transform(X_cv[‘year’].values).reshape(-1,1)
print(‘After Encoding’)
print(‘Train data shape’,X_train_year.shape)
print(‘Test data shape’,X_test_year.shape)
print(‘CV data shape’,X_cv_year.shape)
After Encoding Train data shape (103931, 1) Test data shape (63632, 1) CV data shape (44543, 1)
Let’s export the year encoder
print(‘Saving year encoder..’)
joblib.dump(lab_enc_year,’year_encoder.pkl’)
Saving year encoder..
Out[97]:
['year_encoder.pkl']
BoW Vectorizer
The text data can be encoded in different forms.
The ‘cleaned review’ text data can be vectorized using Bag of Words (BoW) as 1 gram tokens
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
from sklearn.preprocessing import Normalizer,StandardScaler,MinMaxScaler
import joblib
vect_bow_1 = CountVectorizer(min_df=10,ngram_range=(1,1))
X_train1 = X_train.dropna()
vect_bow_1.fit(X_train1[‘cleaned_review’].values)
CountVectorizer(min_df=10)
X_train_review_bow_1 = vect_bow_1.transform(X_train1[‘cleaned_review’].values)
X_test1 = X_test.dropna()
X_test_review_bow_1 = vect_bow_1.transform(X_test1[‘cleaned_review’].values)
X_cv1=X_cv.dropna()
X_cv_review_bow_1 = vect_bow_1.transform(X_cv1[‘cleaned_review’].values)
print(‘After Vectorization’)
print(‘Train data shape:’,X_train_review_bow_1.shape)
print(‘Test data shape:’,X_test_review_bow_1.shape)
print(‘CV data shape:’,X_cv_review_bow_1.shape)
After Vectorization Train data shape: (103927, 7294) Test data shape: (63628, 7294) CV data shape: (44543, 7294)
print(‘Vectorizer for BoW is saved..’)
joblib.dump(vect_bow_1,’vectorizer_bow.pkl’)
Vectorizer for BoW is saved..
Out[108]:
['vectorizer_bow.pkl']
TF-IDF Vectorizer
Let’s vectorize our cleaned review data using the Term Frequency and Inverse Document Frequency (TF-IDF) as 1 gram tokens fitted on training data only
vect_tfidf_1 = TfidfVectorizer(min_df=10,ngram_range=(1,1))
vect_tfidf_1.fit(X_train1[‘cleaned_review’].values)
X_train_review_tfidf_1 = vect_tfidf_1.transform(X_train1[‘cleaned_review’].values)
X_test_review_tfidf_1 = vect_tfidf_1.transform(X_test1[‘cleaned_review’].values)
X_cv_review_tfidf_1 = vect_tfidf_1.transform(X_cv1[‘cleaned_review’].values)
print(‘After Vectorization’)
print(‘Train data shape:’,X_train_review_tfidf_1.shape)
print(‘Test data shape:’,X_test_review_tfidf_1.shape)
print(‘CV data shape:’,X_cv_review_tfidf_1.shape)
After Vectorization Train data shape: (103927, 7294) Test data shape: (63628, 7294) CV data shape: (44543, 7294)
print(‘Vectorizer for TF-IDF is saved..’)
joblib.dump(vect_tfidf_1,’vectorizer_tfidf.pkl’)
Word2Vec Vectorizer
The cleaned review text data is now vectorized using Word2Vec
from tqdm import tqdm
cleaned_reviews = data[‘cleaned_review’].values
In doing so, we need to import the pre-trained Glove model as glove.6B.300d.txt
def loadGloveModel(gloveFile):
print (“Loading Glove Model”)
f = open(gloveFile,’r’, encoding=”utf8″)
model = {}
for line in tqdm(f):
splitLine = line.split()
word = splitLine[0]
embedding = np.array([float(val) for val in splitLine[1:]])
model[word] = embedding
print (“Done.”,len(model),” words loaded!”)
return model
model = loadGloveModel(‘glove.6B.300d.txt’)
Loading Glove Model
400000it [00:34, 11432.30it/s]
Done. 400000 words loaded!
words = []
for i in cleaned_reviews:
words.extend(str(i).split(‘ ‘))
Let' check the number of words that are present in both glove vectors and our corpus
print(“all the words in the corpus”, len(words))
words = set(words)
print(“the unique words in the corpus”, len(words))
inter_words = set(model.keys()).intersection(words)
print(“The number of words that are present in both glove vectors and our corpus”, \
len(inter_words),”(“,np.round(len(inter_words)/len(words)*100,3),”%)”)
words_courpus = {}
words_glove = set(model.keys())
for i in words:
if i in words_glove:
words_courpus[i] = model[i]
print(“word 2 vec length”, len(words_courpus))
all the words in the corpus 8868702 the unique words in the corpus 34667 The number of words that are present in both glove vectors and our corpus 13620 ( 39.288 %) word 2 vec length 13620
Let’s invoke the pickle binary protocol for serializing/de-serializing our vectors
import pickle
with open(‘glove_vectors’, ‘wb’) as f:
pickle.dump(words_courpus, f)
with open(‘glove_vectors’, ‘rb’) as f:
model = pickle.load(f)
glove_words = set(model.keys())
print(len(glove_words))
13620
Let’s create the list of columns
columns = [‘usefulCount’,’word_count’,’unique_word_count’,’char_length’,’count_punctuations’,’stopword_count’,
‘mean_word_len’,’subj_count’,’obj_count’,’CARDINAL’,’DATE’,’EVENT’,’FAC’,’GPE’,’LANGUAGE’,’LAW’,
‘LOC’,’MONEY’,’NORP’,’ORDINAL’,’ORG’, ‘PERCENT’,’PERSON’, ‘PRODUCT’,’QUANTITY’,’TIME’,’WORK_OF_ART’,
‘0’,’1′,’2′,’3′,’4′,’5′,’6′,’7′,’8′,’9′,’10’,’11’,’12’,’13’,’14’,’15’,’16’,’17’,’18’,’19’]
and apply Normalizer() to the train, test and CV data
normalizer = Normalizer()
X_train_num_1 = normalizer.fit_transform(X_train1[columns])
X_test_num_1 = normalizer.fit_transform(X_test1[columns])
X_cv_num_1 = normalizer.fit_transform(X_cv1[columns])
print(“After vectorizations”)
print(X_train_num_1.shape, y_train.shape)
print(X_test_num_1.shape, y_test.shape)
print(X_cv_num_1.shape, y_cv.shape)
After vectorizations (103927, 47) (103931,) (63628, 47) (63632,) (44543, 47) (44543,)
Let’s create the subsets with sentiment_score and sentiment_score_clean values
X_train_sent_score = X_train1[[‘sentiment_score’,’sentiment_score_clean’]].values
X_test_sent_score = X_test1[[‘sentiment_score’,’sentiment_score_clean’]].values
X_cv_sent_score = X_cv1[[‘sentiment_score’,’sentiment_score_clean’]].values
print(“After vectorizations”)
print(X_train_sent_score.shape, y_train.shape)
print(X_test_sent_score.shape, y_test.shape)
print(X_cv_sent_score.shape, y_cv.shape)
After vectorizations (103927, 2) (103931,) (63628, 2) (63632,) (44543, 2) (44543,)
Let’s concatenate all encoded features (all extracted features + sentiment scores)
from scipy.sparse import hstack
X_tr_1 = np.concatenate((X_train_num_1,X_train_sent_score),axis=1)
X_te_1 = np.concatenate((X_test_num_1,X_test_sent_score),axis=1)
X_cv_1 = np.concatenate((X_cv_num_1,X_cv_sent_score),axis=1)
print(“Final Data matrix”)
print(X_tr_1.shape, y_train.shape)
print(X_te_1.shape, y_test.shape)
print(X_cv_1.shape, y_cv.shape)
Final Data matrix (103927, 49) (103931,) (63628, 49) (63632,) (44543, 49) (44543,)
NLP Modelling
We need a few performance metrics
from sklearn.metrics import log_loss, accuracy_score,confusion_matrix, f1_score,roc_auc_score,roc_curve
and the following functions
def plot_confusion_matrix(test_y, predict_y):
C = confusion_matrix(test_y, predict_y)
print(“Number of misclassified points “,(len(test_y)-np.trace(C))/len(test_y)*100)
A =(((C.T)/(C.sum(axis=1))).T)
B =(C/C.sum(axis=0))
plt.figure(figsize=(20,4))
labels = [0,1]
cmap=sns.light_palette("green")
plt.subplot(1, 3, 1)
sns.heatmap(C, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Confusion matrix")
plt.subplot(1, 3, 2)
sns.heatmap(B, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Precision matrix")
plt.subplot(1, 3, 3)
sns.heatmap(A, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.title("Recall matrix")
plt.show()
def model_metrics(clf,train_data,test_data,cv_data):
print('**LogLoss**')
predict_y = clf.predict_proba(train_data)
print ("The train log loss is:",log_loss(y_train, predict_y))
predict_y = clf.predict_proba(cv_data)
print( "The cross validation log loss is:",log_loss(y_cv, predict_y))
predict_y = clf.predict_proba(test_data)
print( "The test log loss is:",log_loss(y_test, predict_y))
print(50*'-')
print('**Accuracy**')
y_pred_tr = clf.predict(train_data)
print ("The train Accuracy is:",accuracy_score(y_train, y_pred_tr))
y_pred_cv = clf.predict(cv_data)
print( "The cross validation Accuracy is:",accuracy_score(y_cv, y_pred_cv))
y_pred_te = clf.predict(test_data)
print( "The test Accuracy is:",accuracy_score(y_test, y_pred_te))
print(50*'-')
print('**F1 Score**')
print ("The train F1 score is:",f1_score(y_train, y_pred_tr))
print( "The cross validation F1 score is:",f1_score(y_cv, y_pred_cv))
print( "The test F1 score is:",f1_score(y_test, y_pred_te))
print(50*'-')
print('**AUC**')
print ("The train AUC is:",roc_auc_score(y_train, y_pred_tr))
print( "The cross validation AUC is:",roc_auc_score(y_cv, y_pred_cv))
print( "The test AUC is:",roc_auc_score(y_test, y_pred_te))
print(50*'-')
As an example, let’s deploy the Random Model
test_len = len(y_test)
predicted_y = np.zeros((test_len,2))
for i in range(test_len):
rand_probs = np.random.rand(1,2)
predicted_y[i] = ((rand_probs/sum(sum(rand_probs)))[0])
print(“Log loss on Test Data using Random Model”,log_loss(y_test, predicted_y, eps=1e-15))
predicted_y =np.argmax(predicted_y, axis=1)
print(“Accuray on Test Data using Random Model”,accuracy_score(y_test, predicted_y))
print(“F1 score on Test Data using Random Model”,f1_score(y_test, predicted_y))
print(“AUC on Test Data using Random Model”,roc_auc_score(y_test, predicted_y))
Log loss on Test Data using Random Model 0.8817010788104614 Accuray on Test Data using Random Model 0.5019644204174001 F1 score on Test Data using Random Model 0.5859171860504618 AUC on Test Data using Random Model 0.5014991363225234
plot_confusion_matrix(y_test,predicted_y)
plt.show()
plt.savefig(‘drugsconfmatrix.png’)
Number of misclassified points 49.80355795826
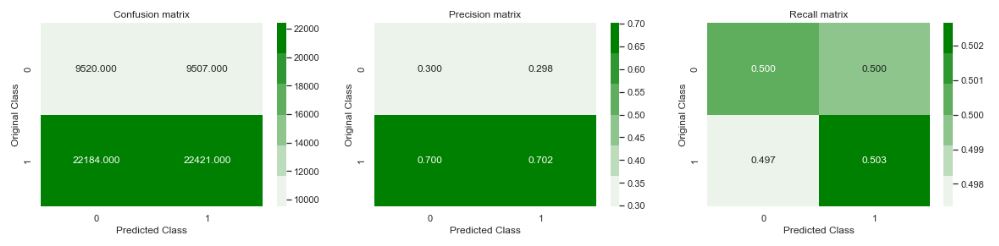
Conclusion
Following recent ML and DA studies, we have addressed the problem of building an NLP-based drug recommendation system in Python. It appears that the Sentiment Analysis, Topic Modelling and Word2Vec techniques play a major role in classifying the drug reviews thereby recommending the effective drugs.
The simplest way to deploy our trained LDA model is to create a web service using the Flask web framework. A final web app can be deployed within the multi-cloud environment discussed in Appendix.
Appendix:
Multi-Cloud NLP Drug Recommendation
One can use Amazon Comprehend Medical to extract medication names and medical conditions to monitor drug safety and adverse events. Amazon Comprehend Medical is a NLP service that uses AI to easily extract relevant medical information from unstructured text. We query the OpenFDA API (an open-source API published by the FDA) and Clinicaltrials.gov API (another open-source API published by the National Library of Medicine (NLM) at the National Institutes of Health (NIH)) to get information on past adverse events, recalls, and clinical trials for the drug or medical condition in question. One can then use this data in population scale studies to further analyze the drug’s safety and efficacy.
Several successful case studies can guide you through the process of creating an ERC20 token recommendation system built with TensorFlow, Cloud Machine Learning Engine, Cloud Endpoints, and App Engine created at Google Cloud. The key point is the collaborative filtering technique for generating user recommendations. Collaborative filtering relies only on observed user behavior to make recommendations — no profile data or content access is necessary. The entire workflow looks like this:
- Creating and training the model for token recommendation system
- Query token ratings from BigQuery
- Train the model locally and in Google ML Engine
- Tuning hyperparameters in Cloud ML Engine
- Deploying the recommendation system to Google App Engine
With the development of e-commerce, a growing number of people prefer to purchase medicine online. To provide online medication guidance, the novel cloud-assisted drug recommendation system CADRE can recommend users with top-N related medicines according to symptoms.
Healthcare organizations are using Azure products and services—including hybrid cloud, mixed reality, AI, and IoT—to drive better health outcomes, improve security, scale faster, and enhance data interoperability.
Text Analytics is now Azure Language Service. Specifically, Sentiment Analysis and Opinion Mining, Named Entity Recognition (NER), Entity Linking, Key Phrase Extraction, Language Detection, Text Analytics for health, and Text Summarization are all part of the Language Service as they exist today.
Using the Text Analytics NER, users can extract known types like person names, geographical locations, datetimes, and organizations. However, lots of information of interest is more specific than the standard types. The capability allows you to build your own custom entity extractors by providing labelled examples of text to train models.

Leave a comment