The goal of this post is a comparison of available binary classifiers in Scikit-Learn on the breast cancer (BC) dataset. The BC dataset comes with the Scikit-Learn package itself.
Contents:
- Data Analysis
- ML Preparation
- Learning Curves
- Feature Dominance
- Calibration Curves
- Confusion Matrix
- ROC Curve
- Precision-Recall Curve
- KS Statistic Plot
- Cumulative Gains Plots
- Lift Curves
- PCA
- Classification Report
- ROC-AUC Score
- Summary
- Explore More
- Infographics
Data Analysis
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
and load the BC dataset
from sklearn import datasets
data = datasets.load_breast_cancer()
with the following keys
print(data.keys())
dict_keys(['data', 'target', 'frame', 'target_names', 'DESCR', 'feature_names', 'filename', 'data_module'])
Let’s check the contents of this dataset
print(data.DESCR)
.. _breast_cancer_dataset:
Breast cancer wisconsin (diagnostic) dataset
--------------------------------------------
**Data Set Characteristics:**
:Number of Instances: 569
:Number of Attributes: 30 numeric, predictive attributes and the class
:Attribute Information:
- radius (mean of distances from center to points on the perimeter)
- texture (standard deviation of gray-scale values)
- perimeter
- area
- smoothness (local variation in radius lengths)
- compactness (perimeter^2 / area - 1.0)
- concavity (severity of concave portions of the contour)
- concave points (number of concave portions of the contour)
- symmetry
- fractal dimension ("coastline approximation" - 1)
The mean, standard error, and "worst" or largest (mean of the three
worst/largest values) of these features were computed for each image,
resulting in 30 features. For instance, field 0 is Mean Radius, field
10 is Radius SE, field 20 is Worst Radius.
- class:
- WDBC-Malignant
- WDBC-Benign
:Summary Statistics:
===================================== ====== ======
Min Max
===================================== ====== ======
radius (mean): 6.981 28.11
texture (mean): 9.71 39.28
perimeter (mean): 43.79 188.5
area (mean): 143.5 2501.0
smoothness (mean): 0.053 0.163
compactness (mean): 0.019 0.345
concavity (mean): 0.0 0.427
concave points (mean): 0.0 0.201
symmetry (mean): 0.106 0.304
fractal dimension (mean): 0.05 0.097
radius (standard error): 0.112 2.873
texture (standard error): 0.36 4.885
perimeter (standard error): 0.757 21.98
area (standard error): 6.802 542.2
smoothness (standard error): 0.002 0.031
compactness (standard error): 0.002 0.135
concavity (standard error): 0.0 0.396
concave points (standard error): 0.0 0.053
symmetry (standard error): 0.008 0.079
fractal dimension (standard error): 0.001 0.03
radius (worst): 7.93 36.04
texture (worst): 12.02 49.54
perimeter (worst): 50.41 251.2
area (worst): 185.2 4254.0
smoothness (worst): 0.071 0.223
compactness (worst): 0.027 1.058
concavity (worst): 0.0 1.252
concave points (worst): 0.0 0.291
symmetry (worst): 0.156 0.664
fractal dimension (worst): 0.055 0.208
===================================== ====== ======
:Missing Attribute Values: None
:Class Distribution: 212 - Malignant, 357 - Benign
:Creator: Dr. William H. Wolberg, W. Nick Street, Olvi L. Mangasarian
:Donor: Nick Street
:Date: November, 1995
This is a copy of UCI ML Breast Cancer Wisconsin (Diagnostic) datasets.
https://goo.gl/U2Uwz2
Features are computed from a digitized image of a fine needle
aspirate (FNA) of a breast mass. They describe
characteristics of the cell nuclei present in the image.
Separating plane described above was obtained using
Multisurface Method-Tree (MSM-T) [K. P. Bennett, "Decision Tree
Construction Via Linear Programming." Proceedings of the 4th
Midwest Artificial Intelligence and Cognitive Science Society,
pp. 97-101, 1992], a classification method which uses linear
programming to construct a decision tree. Relevant features
were selected using an exhaustive search in the space of 1-4
features and 1-3 separating planes.
The actual linear program used to obtain the separating plane
in the 3-dimensional space is that described in:
[K. P. Bennett and O. L. Mangasarian: "Robust Linear
Programming Discrimination of Two Linearly Inseparable Sets",
Optimization Methods and Software 1, 1992, 23-34].
This database is also available through the UW CS ftp server:
ftp ftp.cs.wisc.edu
cd math-prog/cpo-dataset/machine-learn/WDBC/
.. topic:: References
- W.N. Street, W.H. Wolberg and O.L. Mangasarian. Nuclear feature extraction
for breast tumor diagnosis. IS&T/SPIE 1993 International Symposium on
Electronic Imaging: Science and Technology, volume 1905, pages 861-870,
San Jose, CA, 1993.
- O.L. Mangasarian, W.N. Street and W.H. Wolberg. Breast cancer diagnosis and
prognosis via linear programming. Operations Research, 43(4), pages 570-577,
July-August 1995.
- W.H. Wolberg, W.N. Street, and O.L. Mangasarian. Machine learning techniques
to diagnose breast cancer from fine-needle aspirates. Cancer Letters 77 (1994)
163-171.
print(data.target_names)
['malignant' 'benign']
print(data.feature_names)
['mean radius' 'mean texture' 'mean perimeter' 'mean area' 'mean smoothness' 'mean compactness' 'mean concavity' 'mean concave points' 'mean symmetry' 'mean fractal dimension' 'radius error' 'texture error' 'perimeter error' 'area error' 'smoothness error' 'compactness error' 'concavity error' 'concave points error' 'symmetry error' 'fractal dimension error' 'worst radius' 'worst texture' 'worst perimeter' 'worst area' 'worst smoothness' 'worst compactness' 'worst concavity' 'worst concave points' 'worst symmetry' 'worst fractal dimension']
import pandas as pd
df = pd.DataFrame(data.data, columns=data.feature_names)
df[‘target’] = data.target
df.head()
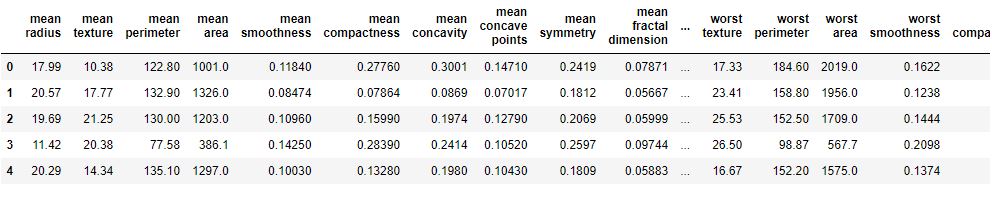
df.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 569 entries, 0 to 568 Data columns (total 31 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 mean radius 569 non-null float64 1 mean texture 569 non-null float64 2 mean perimeter 569 non-null float64 3 mean area 569 non-null float64 4 mean smoothness 569 non-null float64 5 mean compactness 569 non-null float64 6 mean concavity 569 non-null float64 7 mean concave points 569 non-null float64 8 mean symmetry 569 non-null float64 9 mean fractal dimension 569 non-null float64 10 radius error 569 non-null float64 11 texture error 569 non-null float64 12 perimeter error 569 non-null float64 13 area error 569 non-null float64 14 smoothness error 569 non-null float64 15 compactness error 569 non-null float64 16 concavity error 569 non-null float64 17 concave points error 569 non-null float64 18 symmetry error 569 non-null float64 19 fractal dimension error 569 non-null float64 20 worst radius 569 non-null float64 21 worst texture 569 non-null float64 22 worst perimeter 569 non-null float64 23 worst area 569 non-null float64 24 worst smoothness 569 non-null float64 25 worst compactness 569 non-null float64 26 worst concavity 569 non-null float64 27 worst concave points 569 non-null float64 28 worst symmetry 569 non-null float64 29 worst fractal dimension 569 non-null float64 30 target 569 non-null int32 dtypes: float64(30), int32(1) memory usage: 135.7 KB
ML Preparation
Let’s split the data
X = data.data
y = data.target
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y,train_size=0.8,random_state=1)
and apply RobustScaler
ss_train = RobustScaler()
X_train = ss_train.fit_transform(X_train)
ss_test = RobustScaler()
X_test = ss_test.fit_transform(X_test)
Learning Curves
Let’s import the key libraries
import scikitplot as skplt
import sklearn
from sklearn.datasets import load_digits, load_boston, load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor, GradientBoostingClassifier, ExtraTreesClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import sys
import warnings
warnings.filterwarnings(“ignore”)
print(“Scikit Plot Version : “, skplt.version)
print(“Scikit Learn Version : “, sklearn.version)
print(“Python Version : “, sys.version)
%matplotlib inline
Scikit Plot Version : 0.3.7 Scikit Learn Version : 1.1.3 Python Version : 3.9.13 (main, Aug 25 2022, 23:51:50) [MSC v.1916 64 bit (AMD64)]
Let’s look at the train/test data Scikit-Plot learning curves
from sklearn.neighbors import KNeighborsClassifier
logreg = KNeighborsClassifier(n_neighbors=6)
logreg.fit(X_train, y_train)
logreg.score(X_test, y_test)
0.9385964912280702
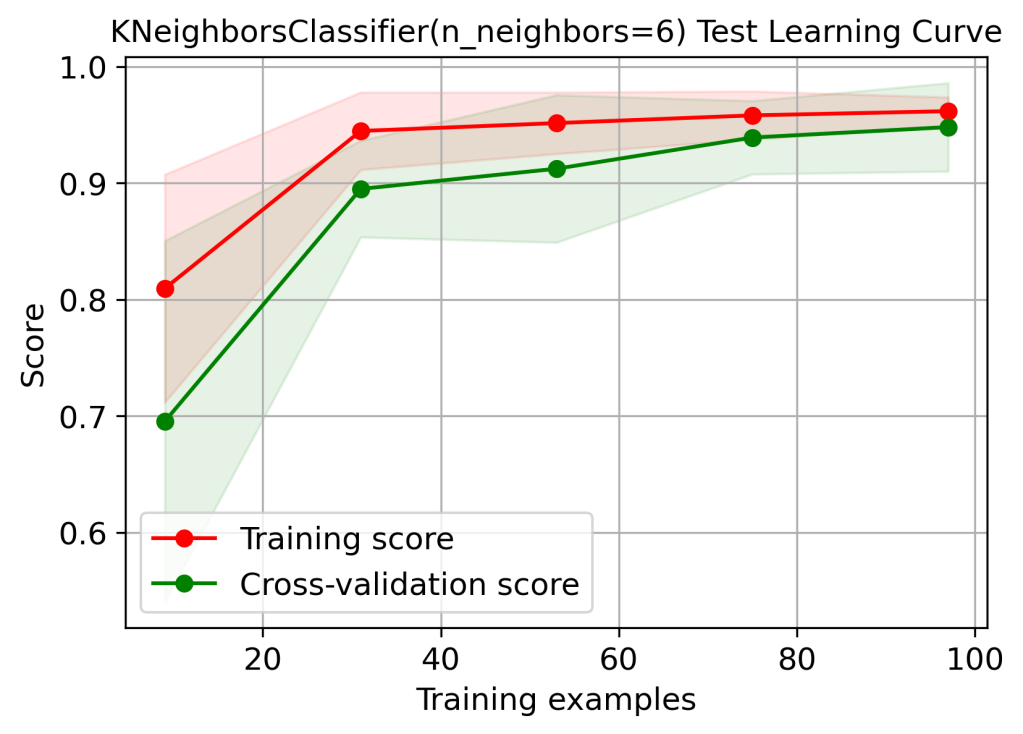
skplt.estimators.plot_learning_curve(logreg, X_test, y_test,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”KNeighborsClassifier(n_neighbors=6) Test Learning Curve”);
plt.savefig(‘learningcurveknn.png’, dpi=300, bbox_inches=’tight’)
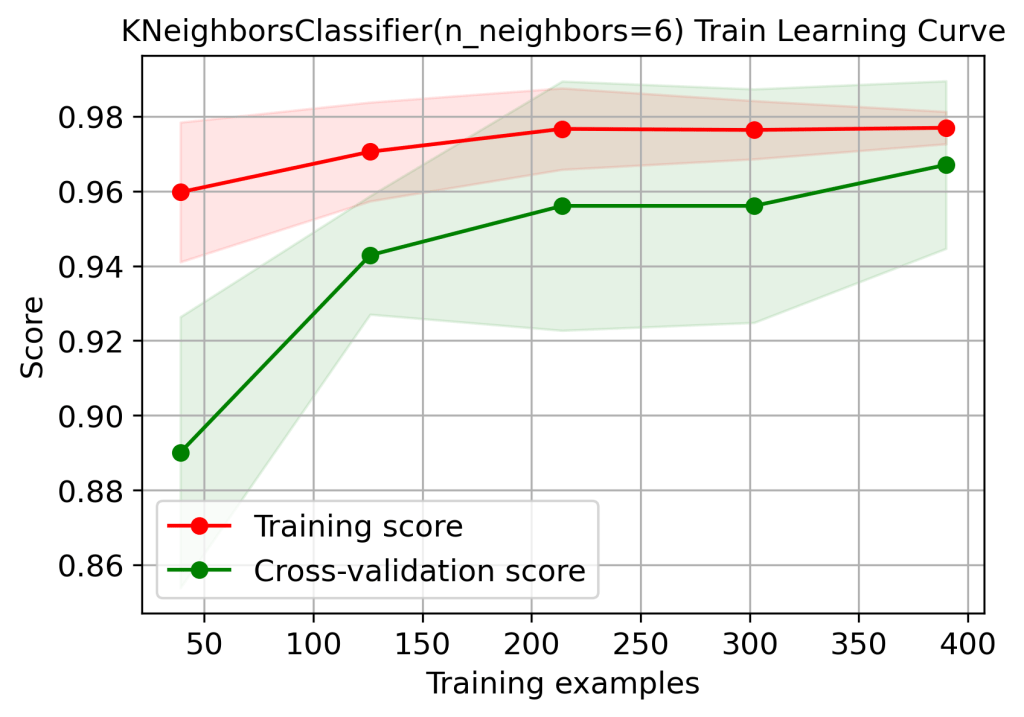
skplt.estimators.plot_learning_curve(logreg, X_train, y_train,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”KNeighborsClassifier(n_neighbors=6) Train Learning Curve”);
plt.savefig(‘learningcurveknntrain.png’, dpi=300, bbox_inches=’tight’)
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
rf.score(X_test, y_test)
0.9298245614035088
skplt.estimators.plot_learning_curve(rf, X_test, y_test,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”RandomForestClassifier() Test Learning Curve”);
plt.savefig(‘learningcurverfctest.png’, dpi=300, bbox_inches=’tight’)

skplt.estimators.plot_learning_curve(rf, X_train, y_train,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”RandomForestClassifier() Train Learning Curve”);
plt.savefig(‘learningcurverfctrain.png’, dpi=300, bbox_inches=’tight’)

gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
gb.score(X_test, y_test)
0.9210526315789473
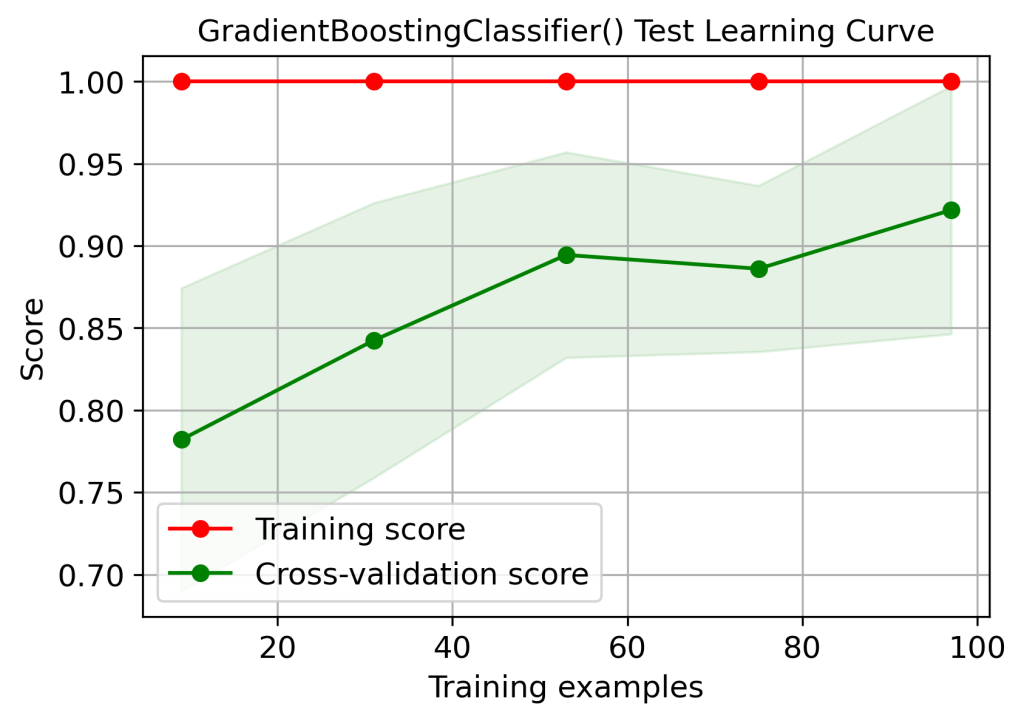
skplt.estimators.plot_learning_curve(gb, X_test, y_test,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”GradientBoostingClassifier() Test Learning Curve”);
plt.savefig(‘learningcurvegbctest.png’, dpi=300, bbox_inches=’tight’)
skplt.estimators.plot_learning_curve(gb, X_train, y_train,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”GradientBoostingClassifier() Train Learning Curve”);
plt.savefig(‘learningcurvegbctrain.png’, dpi=300, bbox_inches=’tight’)

xt = ExtraTreesClassifier()
xt.fit(X_train, y_train)
xt.score(X_test, y_test)
0.9473684210526315
skplt.estimators.plot_learning_curve(xt, X_test, y_test,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”ExtraTreesClassifier() Test Learning Curve”);
plt.savefig(‘learningcurveetctest.png’, dpi=300, bbox_inches=’tight’)
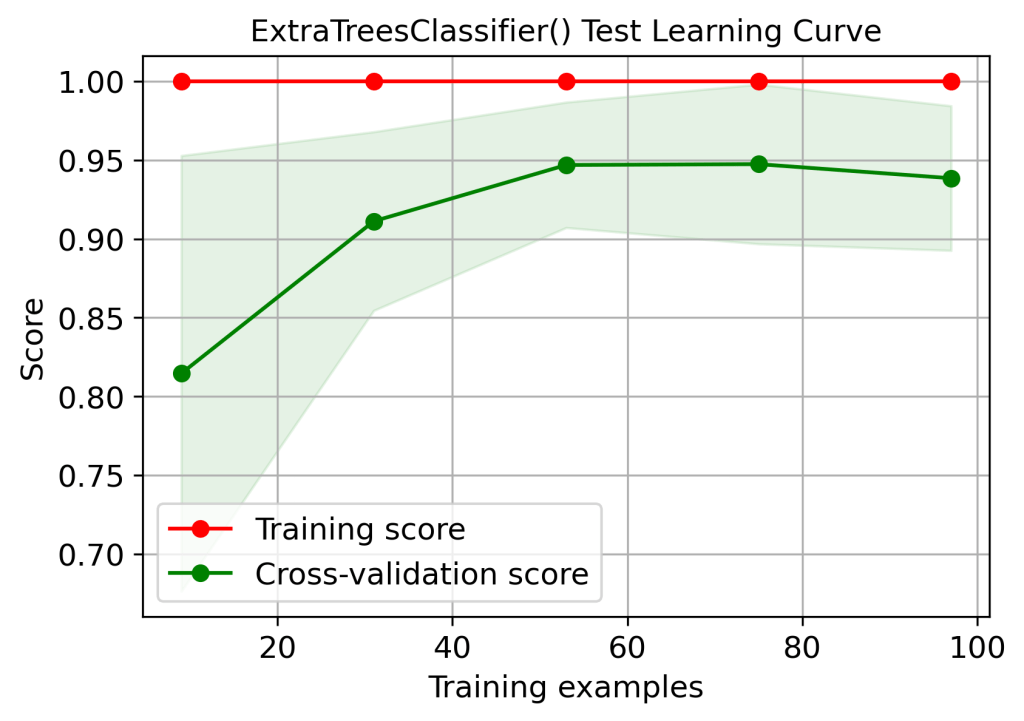
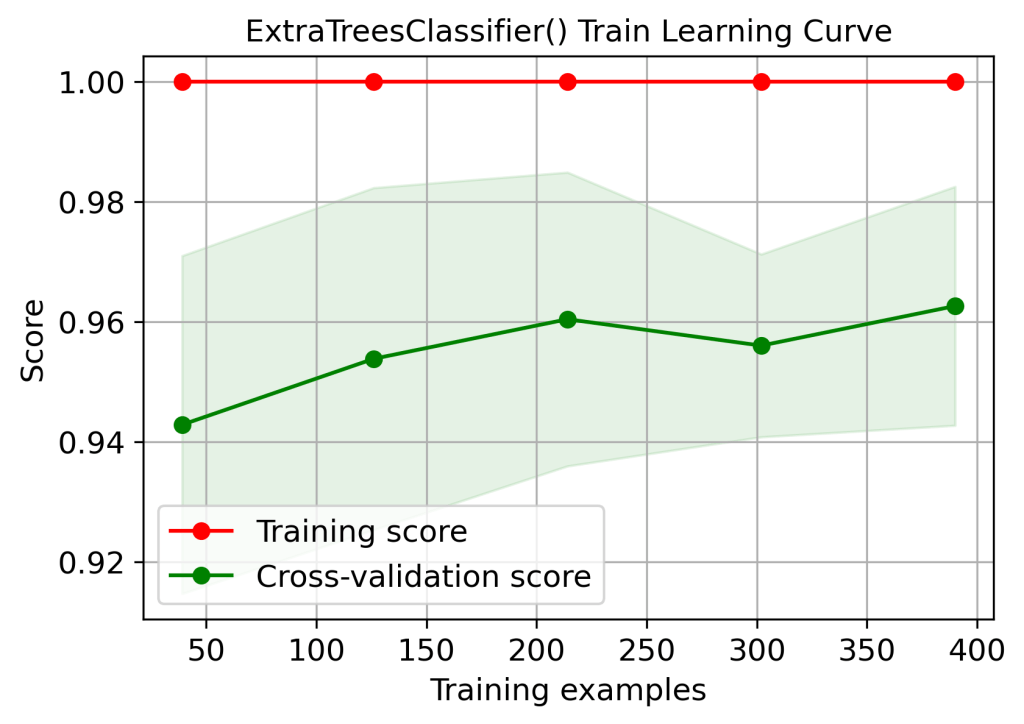
skplt.estimators.plot_learning_curve(xt, X_train, y_train,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”ExtraTreesClassifier() Train Learning Curve”);
plt.savefig(‘learningcurveetctrain.png’, dpi=300, bbox_inches=’tight’)
lr = LogisticRegression()
lr.fit(X_train, y_train)
lr.score(X_test, y_test)
0.9649122807017544
skplt.estimators.plot_learning_curve(lr, X_test, y_test,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”LogisticRegression() Test Learning Curve”);
plt.savefig(‘learningcurvelrtest.png’, dpi=300, bbox_inches=’tight’)
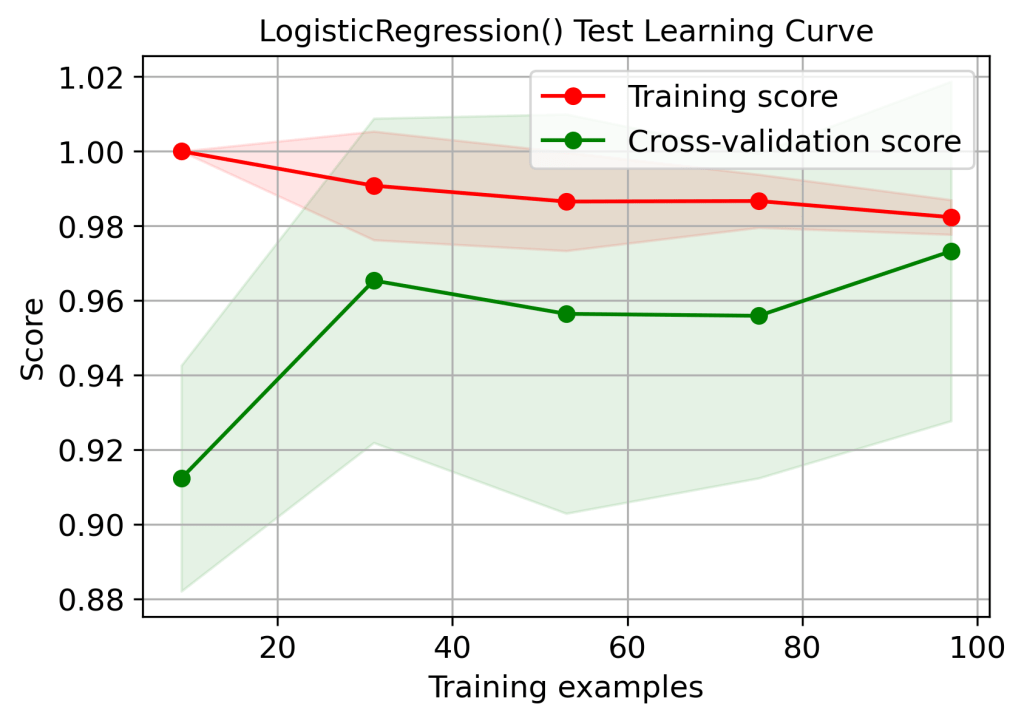
skplt.estimators.plot_learning_curve(lr, X_train, y_train,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”LogisticRegression() Train Learning Curve”);
plt.savefig(‘learningcurvelrtrain.png’, dpi=300, bbox_inches=’tight’)
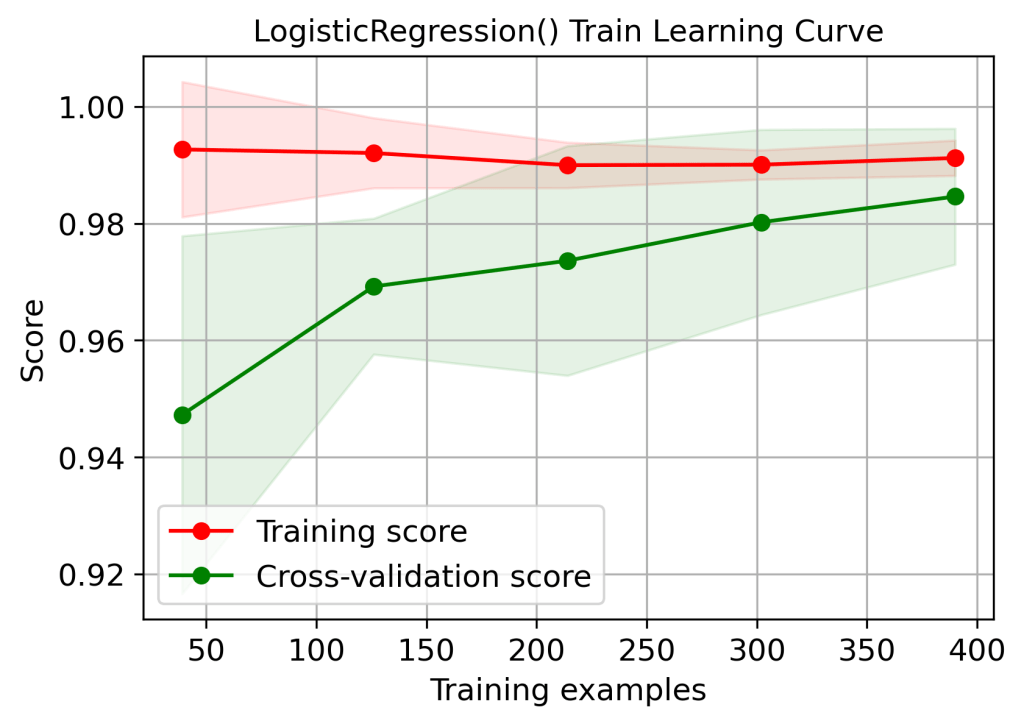
Feature Dominance
Let’s compare the feature dominance coefficients
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(121)
skplt.estimators.plot_feature_importances(rf, feature_names=data.feature_names,
title=”Random Forest Classifier Feature Importance”,
x_tick_rotation=90, order=”ascending”,
ax=ax1);
ax2 = fig.add_subplot(122)
skplt.estimators.plot_feature_importances(xt, feature_names=data.feature_names,
title=”Extra Trees Classifier Feature Importance”,
x_tick_rotation=90,
ax=ax2);
plt.tight_layout()
plt.savefig(‘featureimportancerfxt.png’, dpi=300, bbox_inches=’tight’)
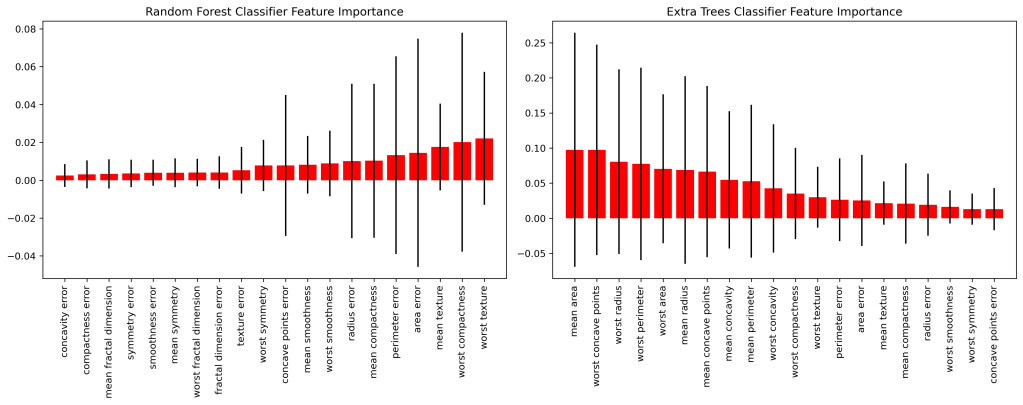
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(121)
skplt.estimators.plot_feature_importances(rf, feature_names=data.feature_names,
title=”RandomForest Feature Importance”,
x_tick_rotation=90, order=”ascending”,
ax=ax1);
ax2 = fig.add_subplot(122)
skplt.estimators.plot_feature_importances(gb, feature_names=data.feature_names,
title=”Gradient Boosting Classifier Feature Importance”,
x_tick_rotation=90,
ax=ax2);
plt.tight_layout()
plt.savefig(‘featureimportancerfgb.png’, dpi=300, bbox_inches=’tight’)

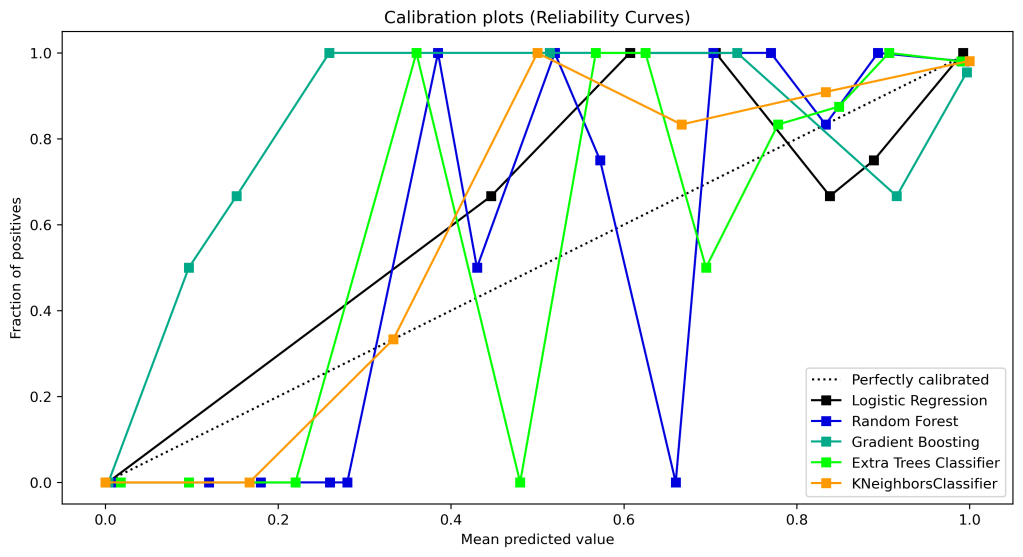
Calibration Curves
Let’s look at the calibration curves
lr_probas = LogisticRegression().fit(X_train, y_train).predict_proba(X_test)
rf_probas = RandomForestClassifier().fit(X_train, y_train).predict_proba(X_test)
gb_probas = GradientBoostingClassifier().fit(X_train, y_train).predict_proba(X_test)
et_scores = ExtraTreesClassifier().fit(X_train, y_train).predict_proba(X_test)
kn_scores = KNeighborsClassifier(n_neighbors=6).fit(X_train, y_train).predict_proba(X_test)
probas_list = [lr_probas, rf_probas, gb_probas, et_scores,kn_scores]
clf_names = [‘Logistic Regression’, ‘Random Forest’, ‘Gradient Boosting’, ‘Extra Trees Classifier’,’KNeighborsClassifier’]
skplt.metrics.plot_calibration_curve(y_test,
probas_list,
clf_names, n_bins=15,
figsize=(12,6)
);
plt.savefig(‘calibrationcurves.png’, dpi=300, bbox_inches=’tight’)
Confusion Matrix
Let’s plot the confusion matrix
lr = LogisticRegression()
lr.fit(X_train, y_train)
y_test_pred_lr = lr.predict(X_test)
gb=GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_test_pred_gb = gb.predict(X_test)
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(121)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_lr,
title=”LogisticRegression Confusion Matrix”,
cmap=”Oranges”,normalize=’all’,
ax=ax1)
ax2 = fig.add_subplot(122)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_gb,
normalize=’all’,
title=”GradientBoostingClassifier Confusion Matrix”,
cmap=”Purples”,
ax=ax2);
plt.savefig(‘confusionmatriceslrgb.png’, dpi=300, bbox_inches=’tight’)
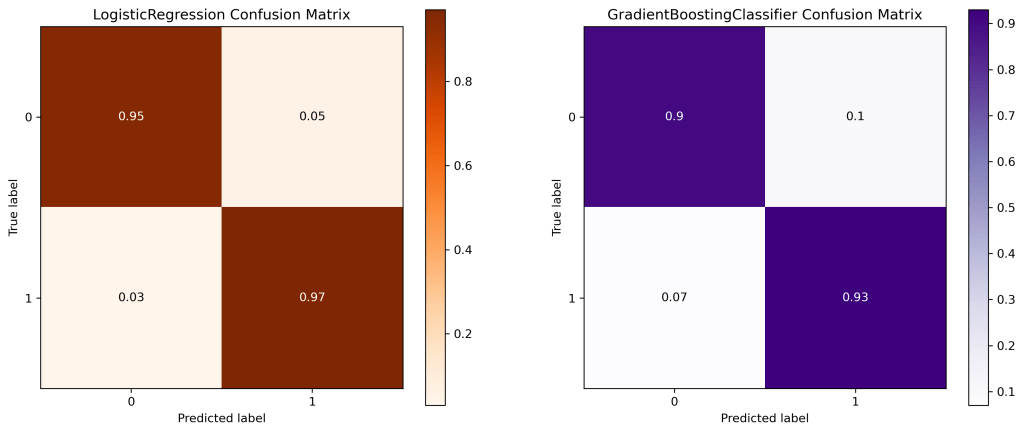
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(121)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_rf,
title=”RandomForest Confusion Matrix”,
cmap=”Oranges”,normalize=’all’,
ax=ax1)
ax2 = fig.add_subplot(122)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_knn,
normalize=’all’,
title=”KNeighbors Confusion Matrix”,
cmap=”Purples”,
ax=ax2);
plt.savefig(‘confusionmatricesrfknn.png’, dpi=300, bbox_inches=’tight’)

lr = LogisticRegression()
lr.fit(X_train, y_train)
y_test_pred_lr = lr.predict(X_test)
xt=ExtraTreesClassifier()
xt.fit(X_train, y_train)
y_test_pred_xt = xt.predict(X_test)
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(121)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_lr,
title=”LogisticRegression Confusion Matrix”,
cmap=”Oranges”,normalize=’all’,
ax=ax1)
ax2 = fig.add_subplot(122)
skplt.metrics.plot_confusion_matrix(y_test, y_test_pred_xt,
normalize=’all’,
title=”ExtraTreesClassifier Confusion Matrix”,
cmap=”Purples”,
ax=ax2);
plt.savefig(‘confusionmatriceslrxt.png’, dpi=300, bbox_inches=’tight’)

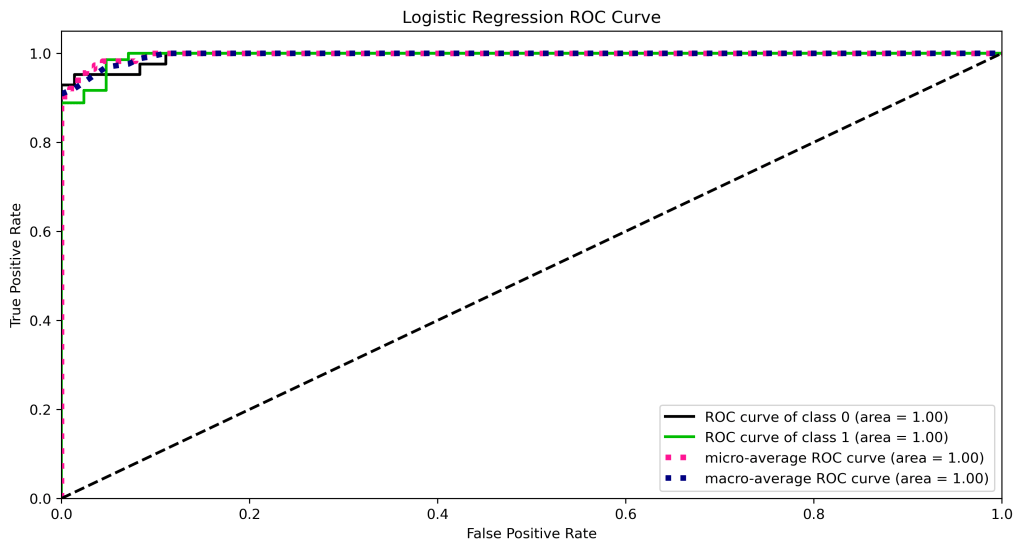
ROC Curve
Let’s compare the ROC curves
y_test_probs = lr.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, y_test_probs,
title=”Logistic Regression ROC Curve”, figsize=(12,6));
plt.savefig(‘roclr.png’, dpi=300, bbox_inches=’tight’)
y_test_probs = xt.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, y_test_probs,
title=”Extra Trees ROC Curve”, figsize=(12,6));
plt.savefig(‘rocxt.png’, dpi=300, bbox_inches=’tight’)

gb=GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_test_probs = gb.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, y_test_probs,
title=”Gradient Boosting ROC Curve”, figsize=(12,6));
plt.savefig(‘rocgb.png’, dpi=300, bbox_inches=’tight’)
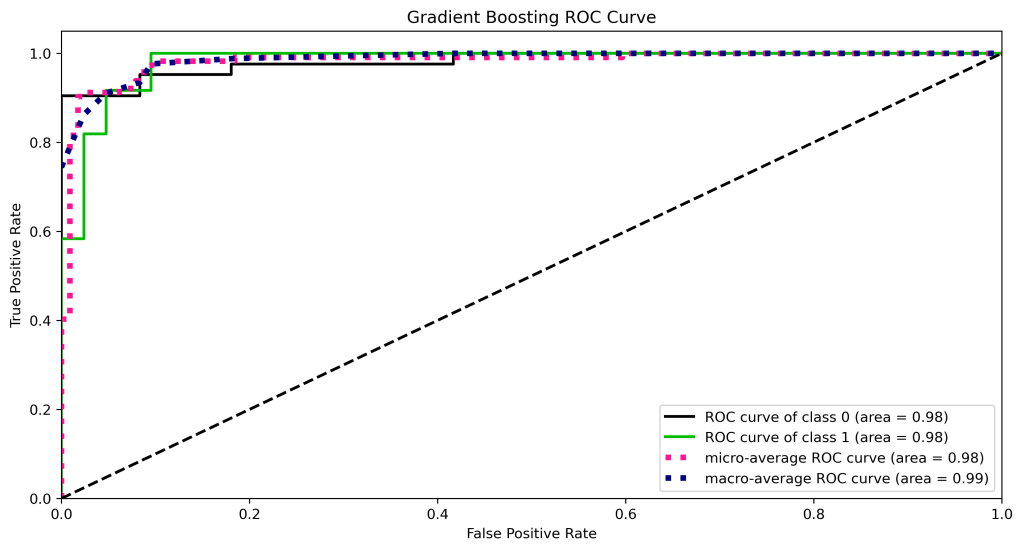
from sklearn.neighbors import KNeighborsClassifier
logreg = KNeighborsClassifier(n_neighbors=6)
logreg.fit(X_train, y_train)
logreg.score(X_test, y_test)
y_test_probs = logreg.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, y_test_probs,
title=”KNeighbors ROC Curve”, figsize=(12,6));
plt.savefig(‘rocknn.png’, dpi=300, bbox_inches=’tight’)
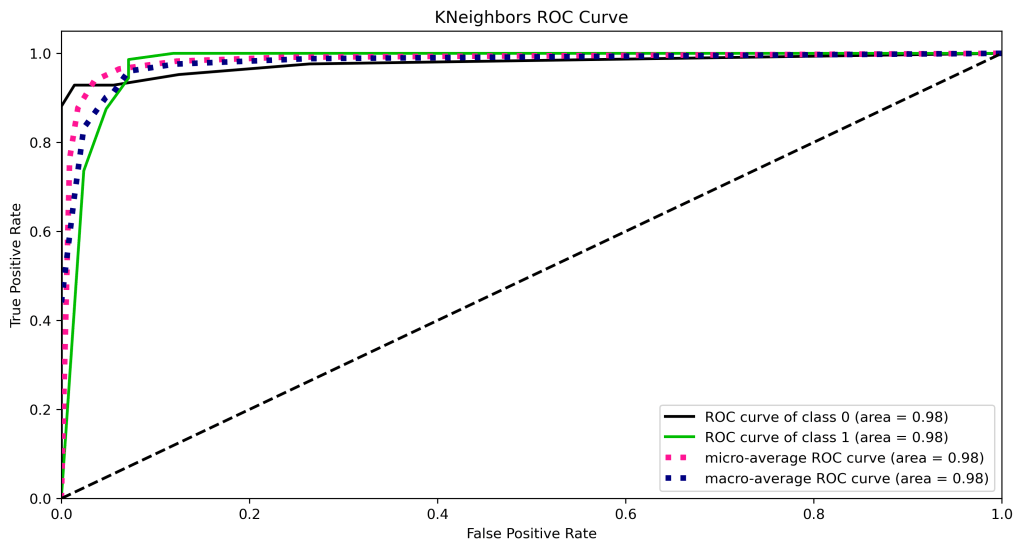
y_test_probs = rf.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, y_test_probs,
title=”RandomForest ROC Curve”, figsize=(12,6));
plt.savefig(‘rocrfc.png’, dpi=300, bbox_inches=’tight’)
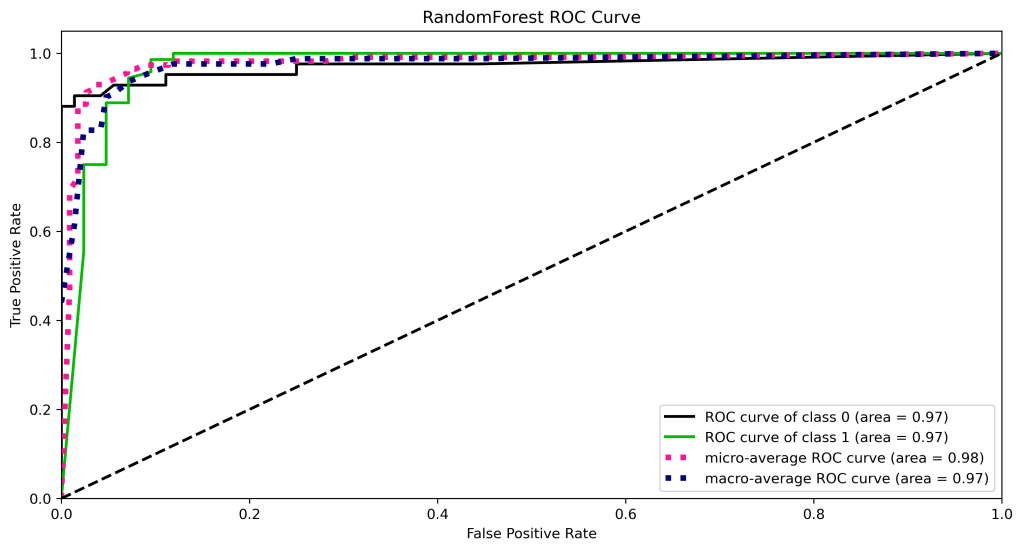
Precision-Recall Curve
Let’s compare the precision-recall curves
y_test_probs = lr.predict_proba(X_test)
skplt.metrics.plot_precision_recall_curve(y_test, y_test_probs,
title=”Logistic Regression Precision-Recall Curve”, figsize=(12,6));
plt.savefig(‘precision-recalllr.png’, dpi=300, bbox_inches=’tight’)
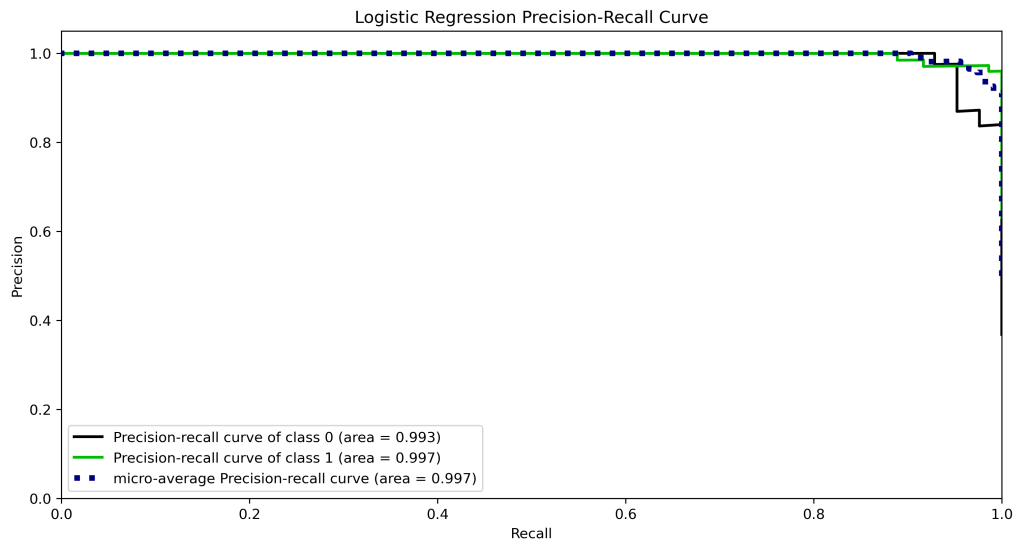
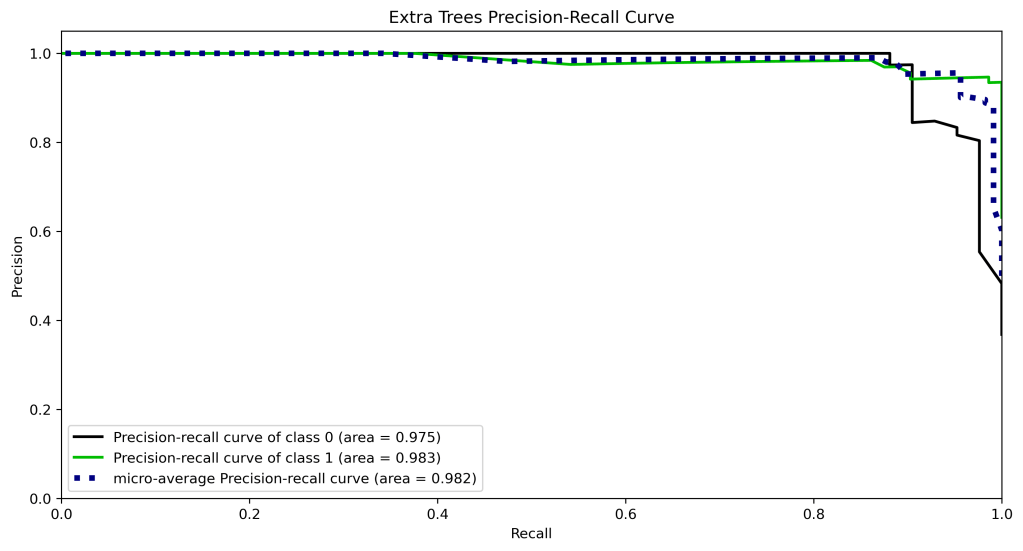
y_test_probs = xt.predict_proba(X_test)
skplt.metrics.plot_precision_recall_curve(y_test, y_test_probs,
title=”Extra Trees Precision-Recall Curve”, figsize=(12,6));
plt.savefig(‘precision-recallxt.png’, dpi=300, bbox_inches=’tight’)
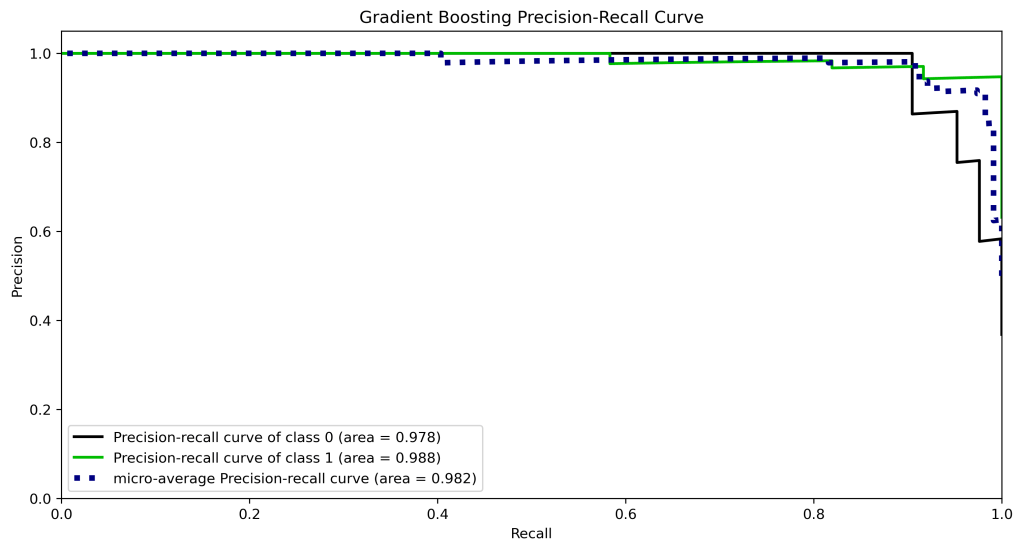
y_test_probs = gb.predict_proba(X_test)
skplt.metrics.plot_precision_recall_curve(y_test, y_test_probs,
title=”Gradient Boosting Precision-Recall Curve”, figsize=(12,6));
plt.savefig(‘precision-recallgb.png’, dpi=300, bbox_inches=’tight’)
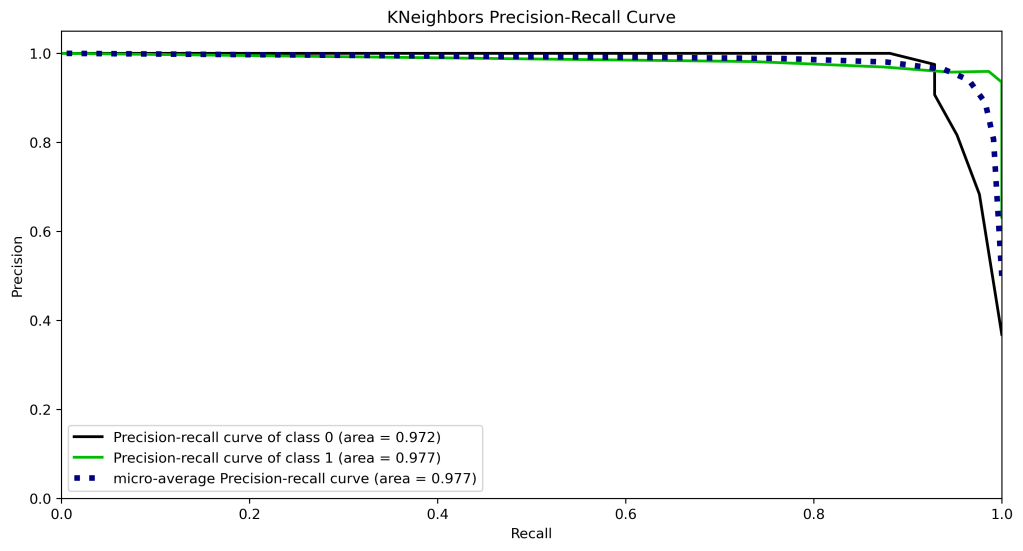
y_test_probs = logreg.predict_proba(X_test)
skplt.metrics.plot_precision_recall_curve(y_test, y_test_probs,
title=”KNeighbors Precision-Recall Curve”, figsize=(12,6));
plt.savefig(‘precision-recallknn.png’, dpi=300, bbox_inches=’tight’)
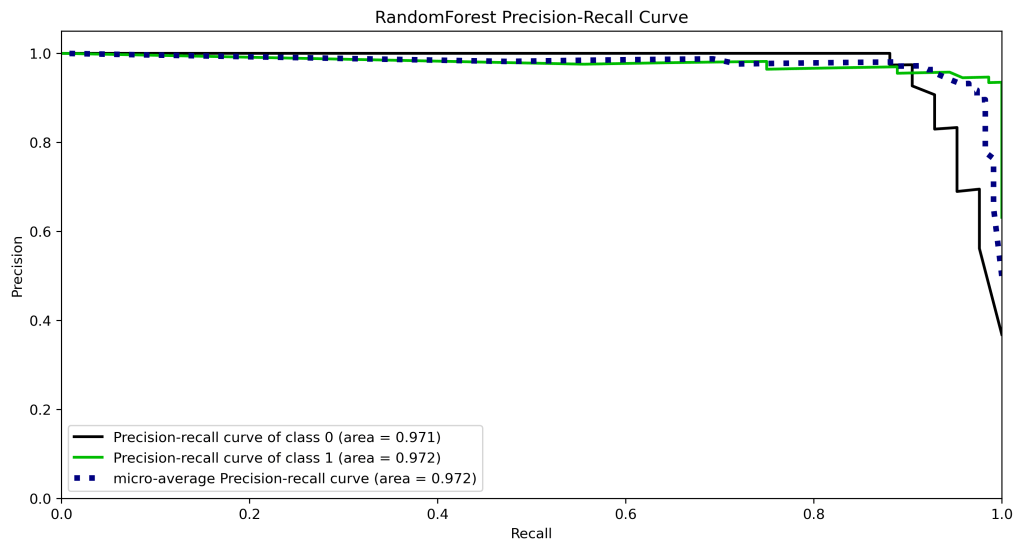
y_test_probs = rf.predict_proba(X_test)
skplt.metrics.plot_precision_recall_curve(y_test, y_test_probs,
title=”RandomForest Precision-Recall Curve”, figsize=(12,6));
plt.savefig(‘precision-recallrfc.png’, dpi=300, bbox_inches=’tight’)
KS Statistic Plot
Let’s compare the KS statistic plots
lr = LogisticRegression()
lr.fit(X_train, y_train)
y_test_pred_proba_lr = lr.predict_proba(X_test)
skplt.metrics.plot_ks_statistic(y_test, y_test_pred_proba_lr, figsize=(10,6),title=”Logistic Regression KS Statistic Plot”);
plt.savefig(‘kstatlr.png’, dpi=300, bbox_inches=’tight’)

xt = ExtraTreesClassifier()
xt.fit(X_train, y_train)
y_test_pred_proba_xt = xt.predict_proba(X_test)
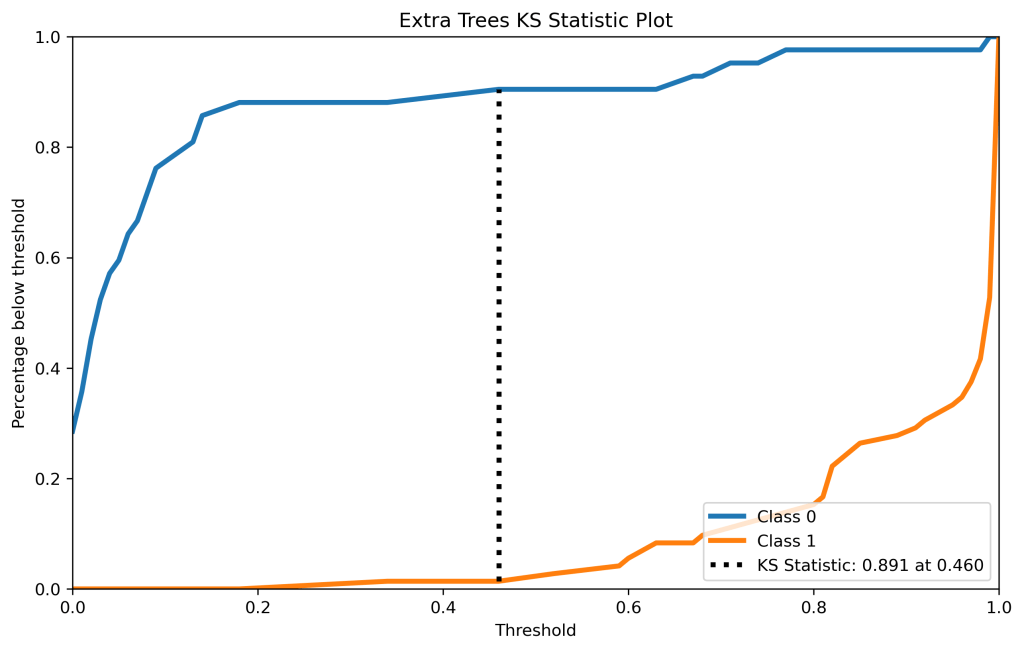
skplt.metrics.plot_ks_statistic(y_test, y_test_pred_proba_xt, figsize=(10,6),title=”Extra Trees KS Statistic Plot”);
plt.savefig(‘kstatxt.png’, dpi=300, bbox_inches=’tight’)
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_test_pred_proba_gb = gb.predict_proba(X_test)
skplt.metrics.plot_ks_statistic(y_test, y_test_pred_proba_gb, figsize=(10,6),title=”Gradient Boosting KS Statistic Plot”);
plt.savefig(‘kstatgb.png’, dpi=300, bbox_inches=’tight’)

rf = RandomForestClassifier()
rf.fit(X_train, y_train)
y_test_pred_proba_rf = rf.predict_proba(X_test)
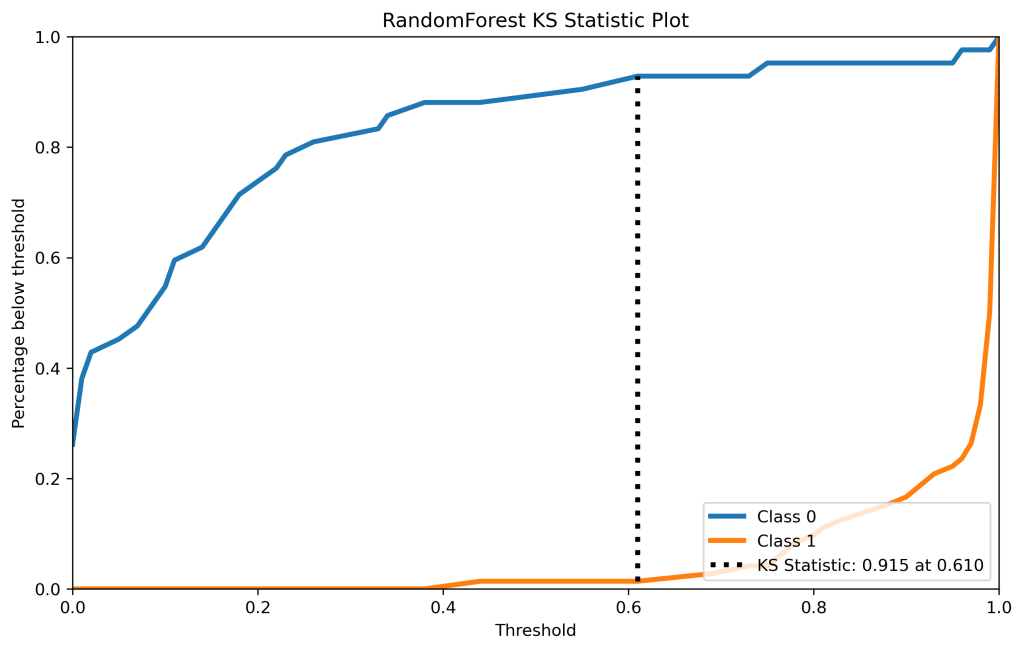
skplt.metrics.plot_ks_statistic(y_test, y_test_pred_proba_rf, figsize=(10,6),title=”Random Forest KS Statistic Plot”);
plt.savefig(‘kstatrf.png’, dpi=300, bbox_inches=’tight’)
logreg = KNeighborsClassifier(n_neighbors=6)
logreg.fit(X_train, y_train)
y_test_pred_proba_knn = logreg.predict_proba(X_test)
skplt.metrics.plot_ks_statistic(y_test, y_test_pred_proba_knn, figsize=(10,6),title=”KNeighbors KS Statistic Plot”);
plt.savefig(‘kstatknn.png’, dpi=300, bbox_inches=’tight’)

Cumulative Gains Plots
lr_probas = LogisticRegression().fit(X_train, y_train).predict_proba(X_test)
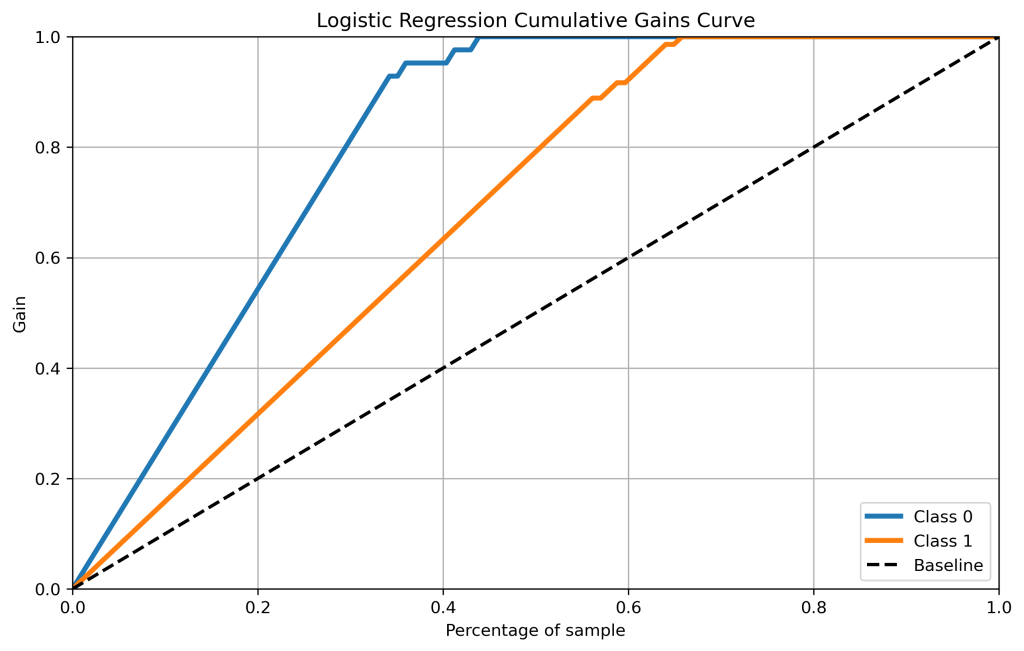
skplt.metrics.plot_cumulative_gain(y_test, lr_probas, figsize=(10,6),title=”Logistic Regression Cumulative Gains Curve”);
plt.savefig(‘cumgainlr.png’, dpi=300, bbox_inches=’tight’)
xt_probas = ExtraTreesClassifier().fit(X_train, y_train).predict_proba(X_test)
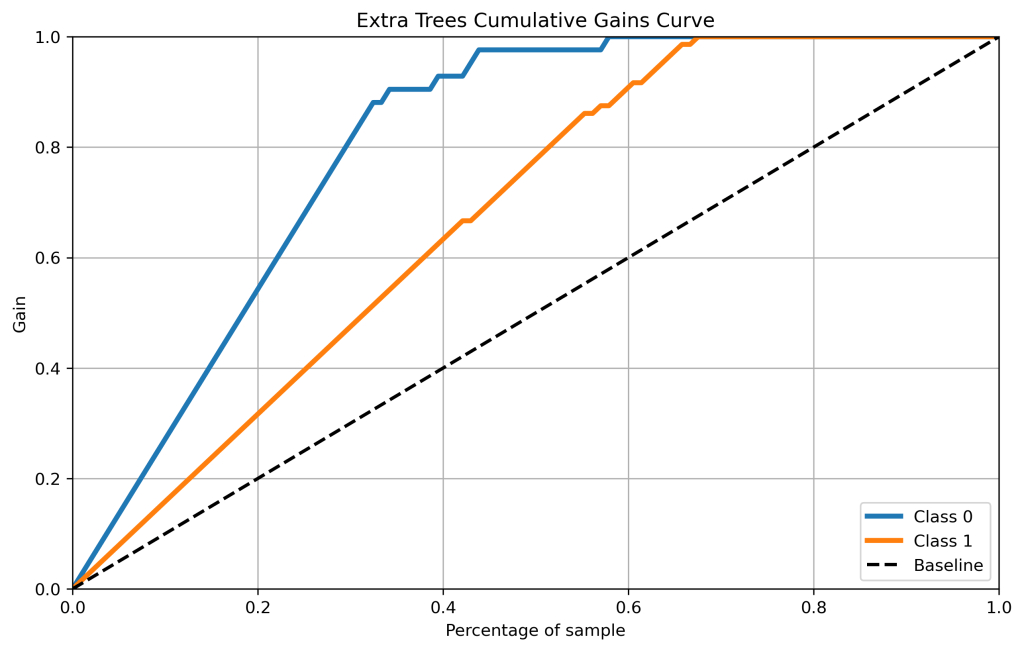
skplt.metrics.plot_cumulative_gain(y_test, xt_probas, figsize=(10,6),title=”Extra Trees Cumulative Gains Curve”);
plt.savefig(‘cumgainxt.png’, dpi=300, bbox_inches=’tight’)
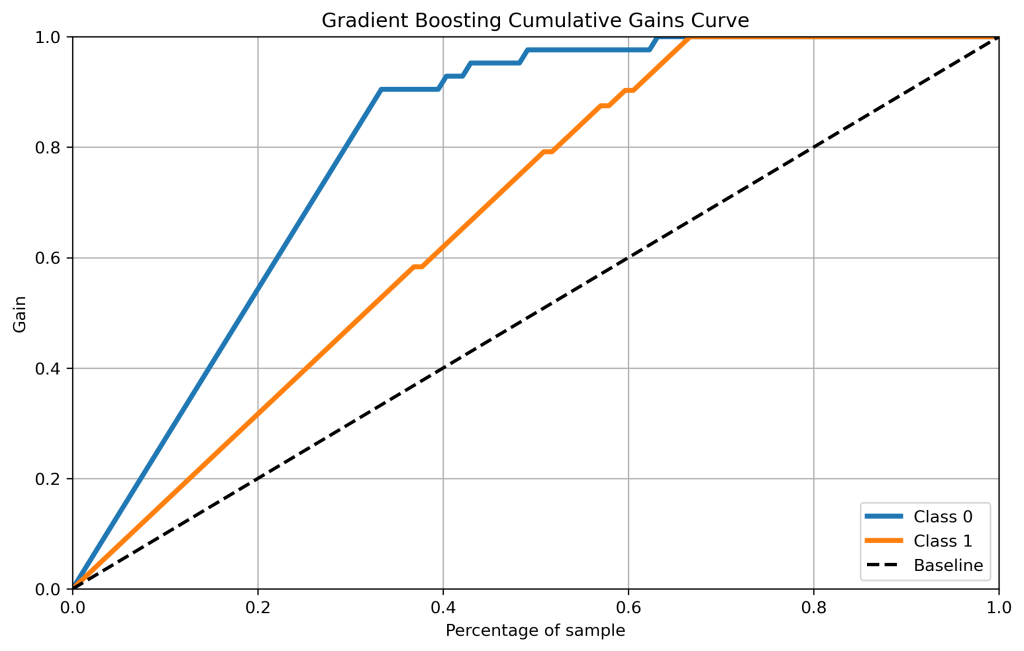
gb_probas = GradientBoostingClassifier().fit(X_train, y_train).predict_proba(X_test)
skplt.metrics.plot_cumulative_gain(y_test, gb_probas, figsize=(10,6),title=”Gradient Boosting Cumulative Gains Curve”);
plt.savefig(‘cumgaingb.png’, dpi=300, bbox_inches=’tight’)
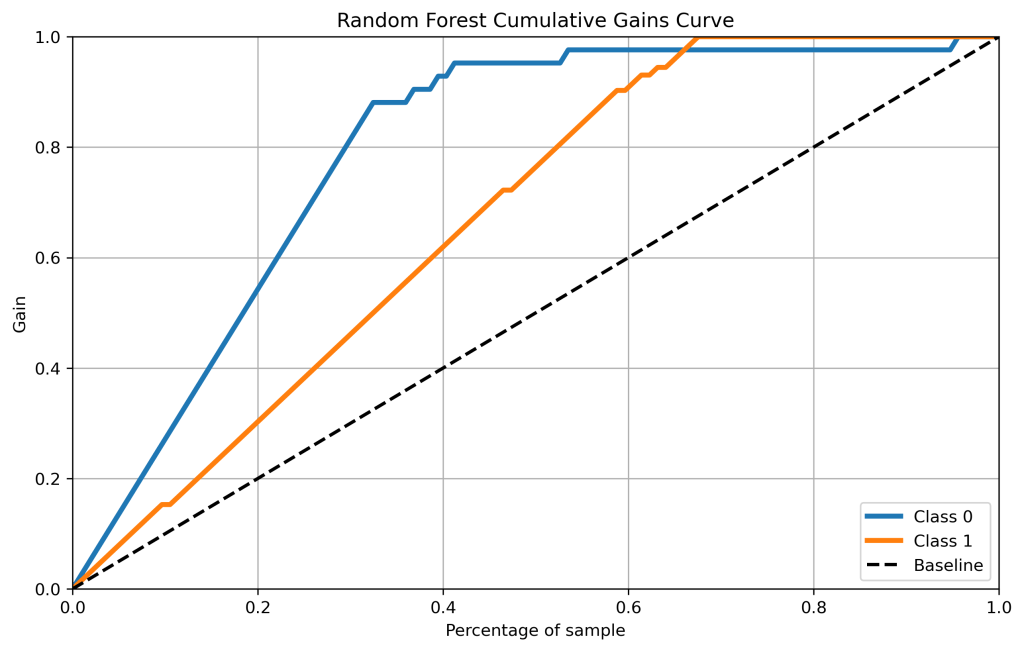
rf_probas = RandomForestClassifier().fit(X_train, y_train).predict_proba(X_test)
skplt.metrics.plot_cumulative_gain(y_test, rf_probas, figsize=(10,6),title=”Random Forest Cumulative Gains Curve”);
plt.savefig(‘cumgainrf.png’, dpi=300, bbox_inches=’tight’)
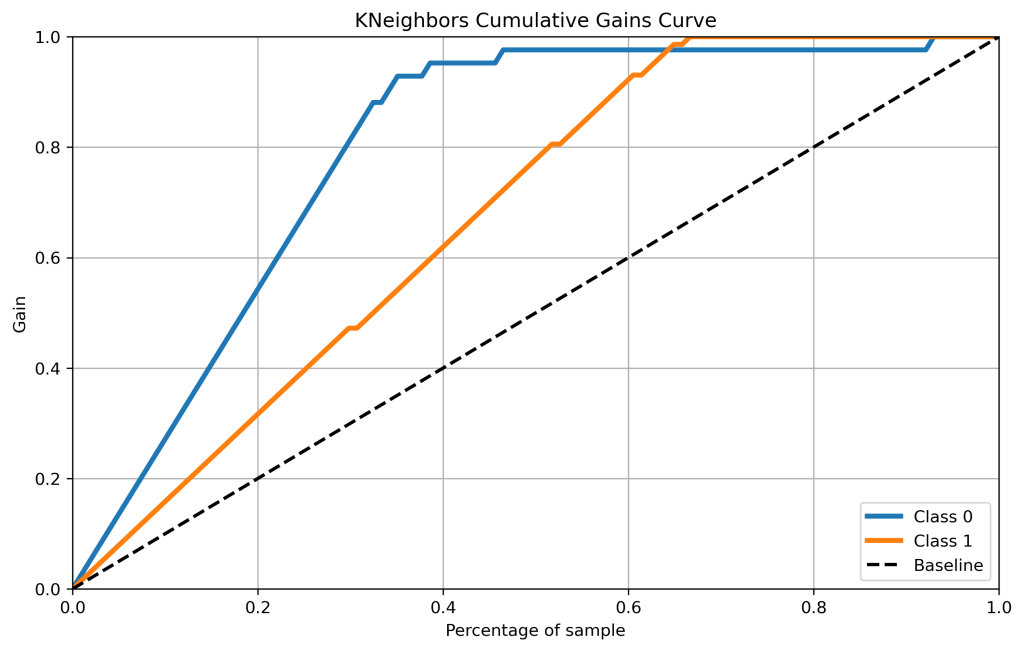
skplt.metrics.plot_cumulative_gain(y_test, kn_scores, figsize=(10,6),title=”KNeighbors Cumulative Gains Curve”);
plt.savefig(‘cumgainknn.png’, dpi=300, bbox_inches=’tight’)
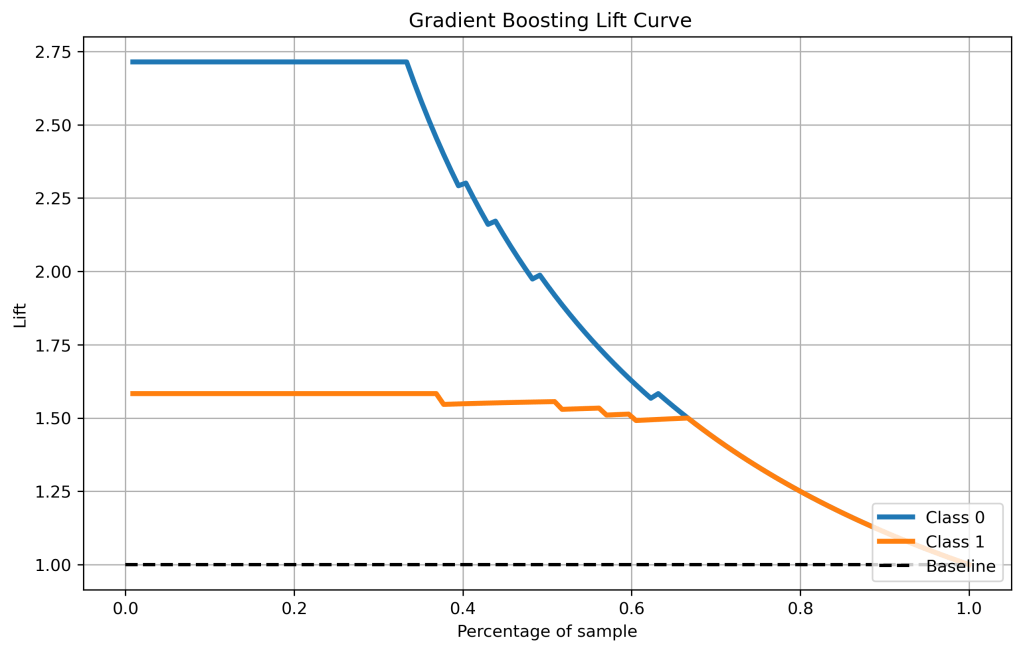
Lift Curves
Let’s compare the lift curves
skplt.metrics.plot_lift_curve(y_test, lr_probas, figsize=(10,6),title=” Logistic Regression Lift Curve”);
plt.savefig(‘liftlr.png’, dpi=300, bbox_inches=’tight’)

xt = ExtraTreesClassifier()
xt.fit(X_train, y_train)
y_test_pred_proba_xt = xt.predict_proba(X_test)
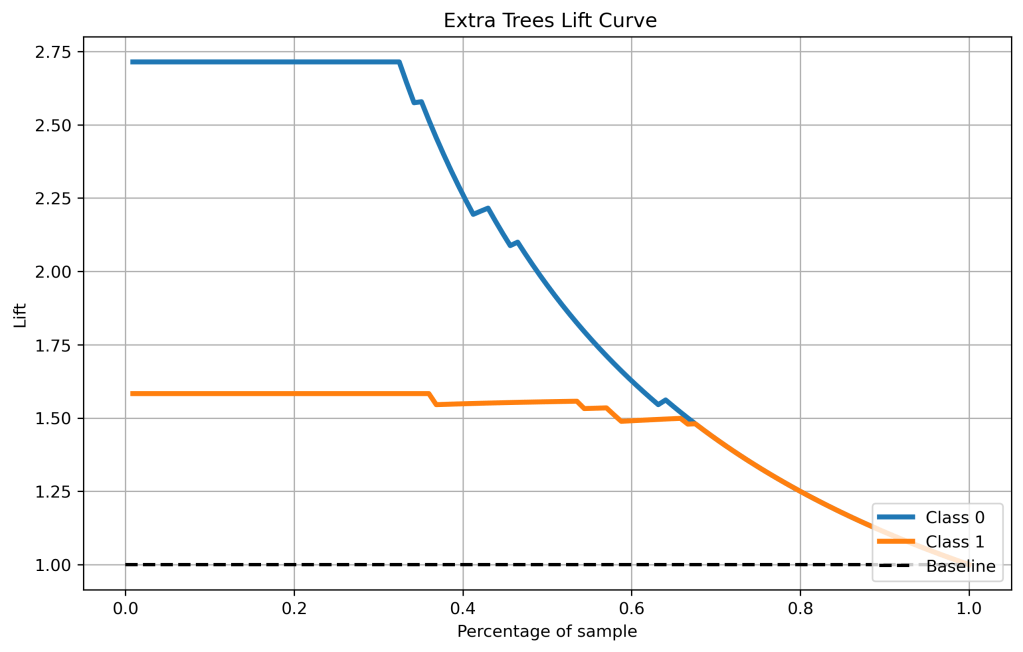
skplt.metrics.plot_lift_curve(y_test, y_test_pred_proba_xt, figsize=(10,6),title=” Extra Trees Lift Curve”);
plt.savefig(‘liftxt.png’, dpi=300, bbox_inches=’tight’)
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_test_pred_proba_gb = gb.predict_proba(X_test)
skplt.metrics.plot_lift_curve(y_test, y_test_pred_proba_gb, figsize=(10,6),title=” Gradient Boosting Lift Curve”);
plt.savefig(‘liftgb.png’, dpi=300, bbox_inches=’tight’)
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
y_test_pred_proba_rf = rf.predict_proba(X_test)
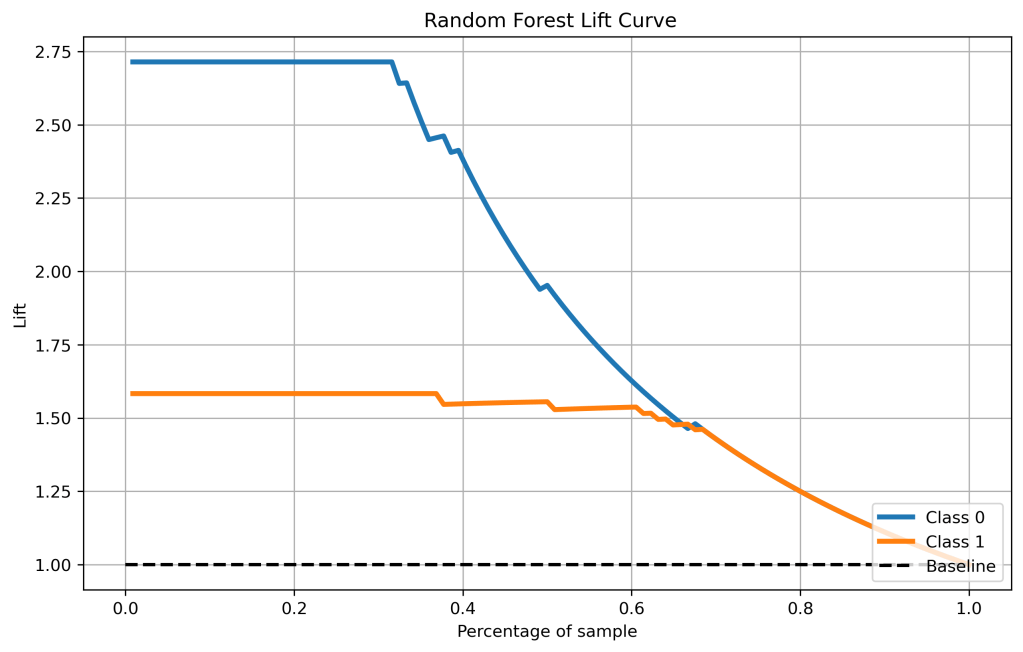
skplt.metrics.plot_lift_curve(y_test, y_test_pred_proba_rf, figsize=(10,6),title=” Random Forest Lift Curve”);
plt.savefig(‘liftrf.png’, dpi=300, bbox_inches=’tight’)
skplt.metrics.plot_lift_curve(y_test, kn_scores, figsize=(10,6),title=” KNeighbors Lift Curve”);
plt.savefig(‘liftknn.png’, dpi=300, bbox_inches=’tight’)

PCA
skplt.cluster.plot_elbow_curve(KMeans(random_state=1),
X,
cluster_ranges=range(2, 20),
figsize=(8,6));
plt.savefig(‘elbowplot.png’, dpi=300, bbox_inches=’tight’)

kmeans = KMeans(n_clusters=10, random_state=1)
kmeans.fit(X_train, y_train)
cluster_labels = kmeans.predict(X_test)
skplt.metrics.plot_silhouette(X_test, cluster_labels,
figsize=(8,6));
plt.savefig(‘silhouette.png’, dpi=300, bbox_inches=’tight’)

pca = PCA(random_state=1)
pca.fit(X)
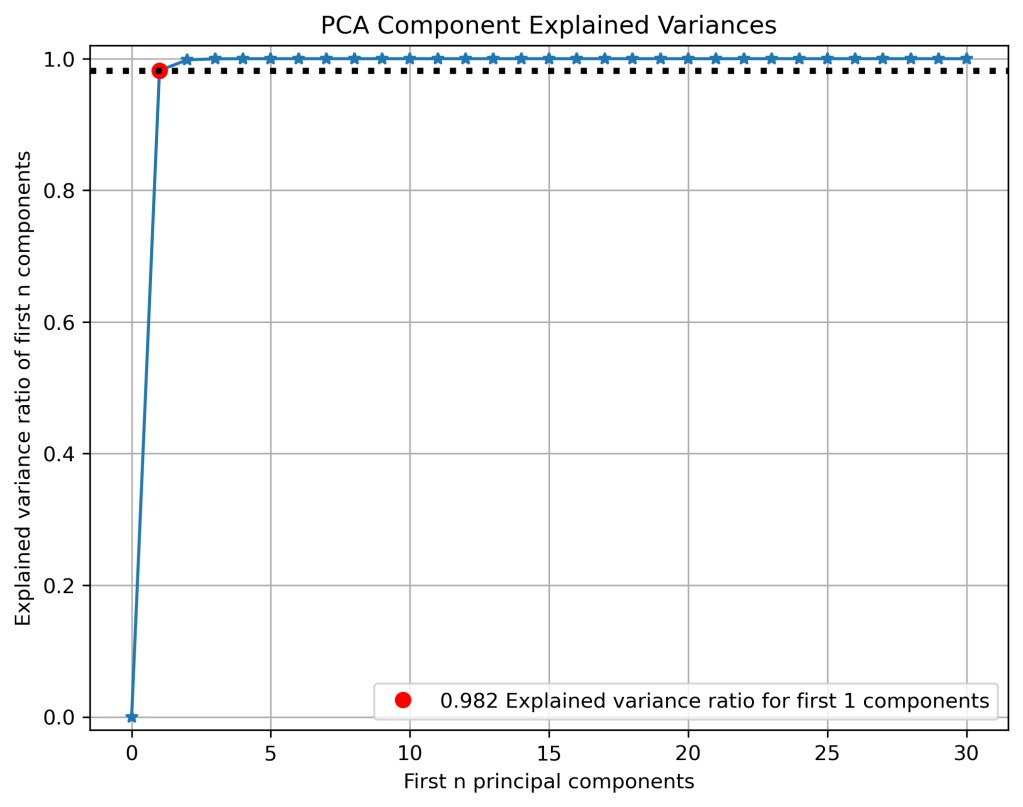
skplt.decomposition.plot_pca_component_variance(pca, figsize=(8,6));
plt.savefig(‘pcavariances.png’, dpi=300, bbox_inches=’tight’)
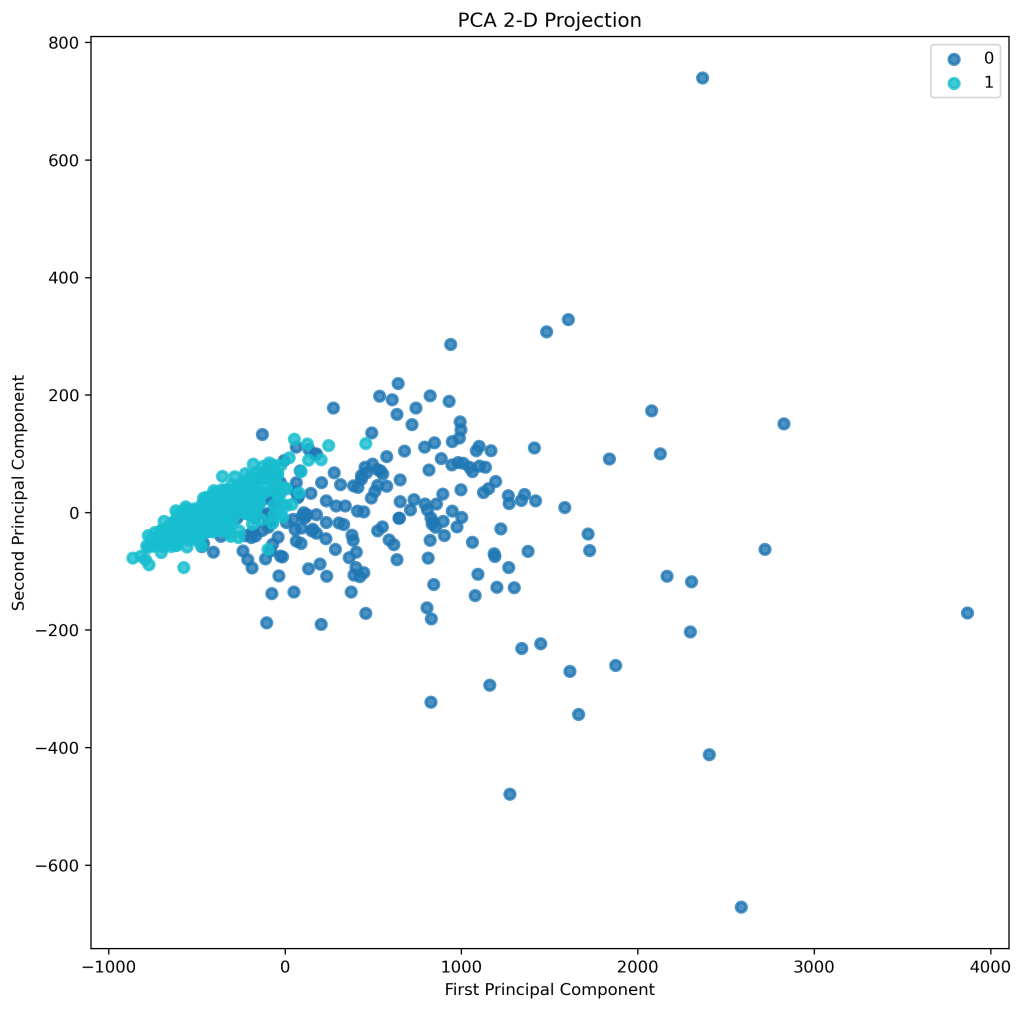
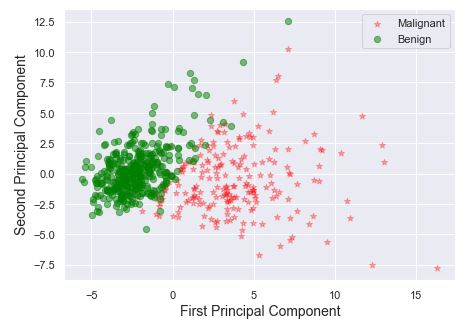
skplt.decomposition.plot_pca_2d_projection(pca, X, y,
figsize=(10,10),
cmap=”tab10″);
plt.savefig(‘pca2dprojection.png’, dpi=300, bbox_inches=’tight’)
Classification Report
Let’s compare the classification reports
lr = LogisticRegression()
lr.fit(X_train, y_train)
y_test_pred_lr = lr.predict(X_test)
print(accuracy_score(y_test_pred_lr, y_test))
print(confusion_matrix(y_test_pred_lr, y_test))
print(classification_report(y_test_pred_lr, y_test))
0.9649122807017544
[[40 2]
[ 2 70]]
precision recall f1-score support
0 0.95 0.95 0.95 42
1 0.97 0.97 0.97 72
accuracy 0.96 114
macro avg 0.96 0.96 0.96 114
weighted avg 0.96 0.96 0.96 114
from sklearn.metrics import ConfusionMatrixDisplay
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
Roc curve and auc score:
from sklearn.datasets import make_classification
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
class_names = [‘0’, ‘1’]
Run classifier, using a model that is too regularized (C too low) to see
the impact on the results
classifier = lr.fit(X_train, y_train)
np.set_printoptions(precision=2)
Plot non-normalized confusion matrix
titles_options = [
(“LR Confusion matrix, without normalization”, None),
(“LR Normalized confusion matrix”, “true”),
]
for title, normalize in titles_options:
disp = ConfusionMatrixDisplay.from_estimator(
classifier,
X_test,
y_test,
display_labels=class_names,
cmap=plt.cm.Blues,
normalize=normalize,
)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.savefig(‘confmatrixlr.png’, dpi=300, bbox_inches=’tight’)
LR Confusion matrix, without normalization [[40 2] [ 2 70]] LR Normalized confusion matrix [[0.95 0.05] [0.03 0.97]]
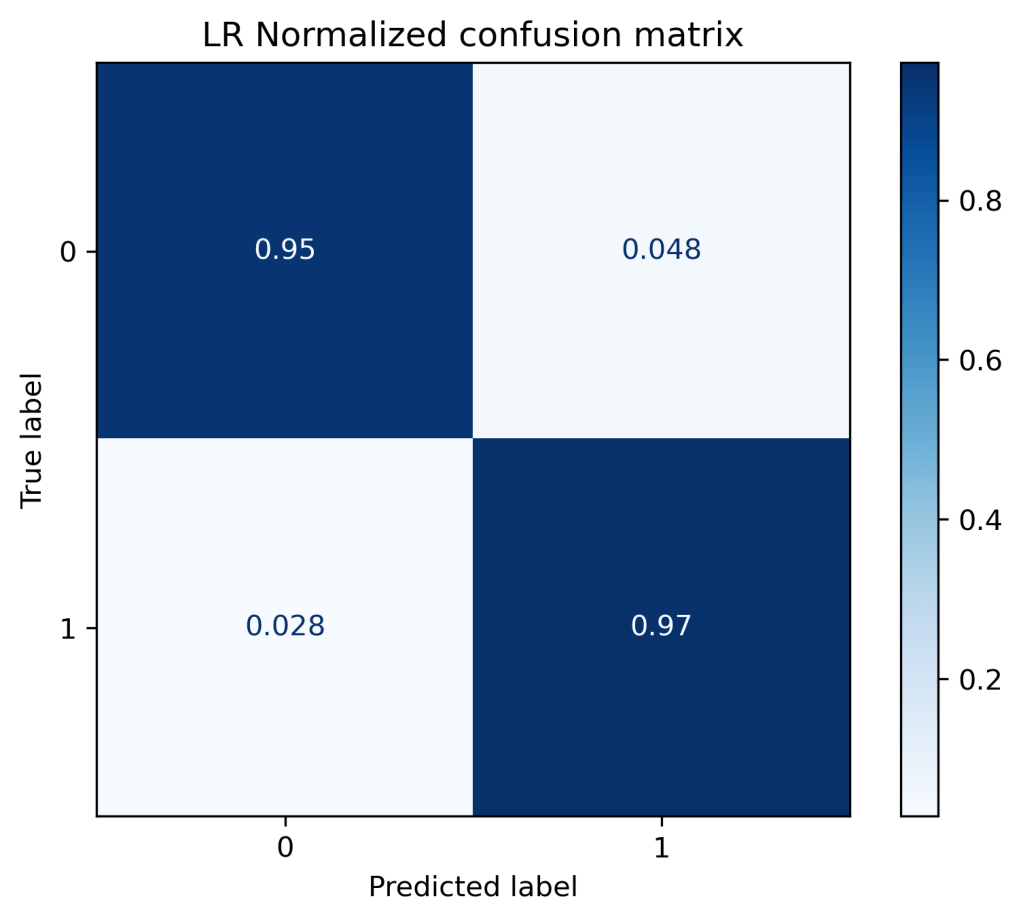
xt = ExtraTreesClassifier()
xt.fit(X_train, y_train)
y_test_pred_xt = xt.predict(X_test)
print(accuracy_score(y_test_pred_xt, y_test))
print(confusion_matrix(y_test_pred_xt, y_test))
print(classification_report(y_test_pred_xt, y_test))
0.9385964912280702
[[37 2]
[ 5 70]]
precision recall f1-score support
0 0.88 0.95 0.91 39
1 0.97 0.93 0.95 75
accuracy 0.94 114
macro avg 0.93 0.94 0.93 114
weighted avg 0.94 0.94 0.94 114
titles_options = [
(“ExtraTrees Confusion matrix, without normalization”, None),
(“ExtraTrees Normalized confusion matrix”, “true”),
]
for title, normalize in titles_options:
disp = ConfusionMatrixDisplay.from_estimator(
classifier,
X_test,
y_test,
display_labels=class_names,
cmap=plt.cm.Blues,
normalize=normalize,
)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.savefig(‘confmatrixxt.png’, dpi=300, bbox_inches=’tight’)
ExtraTrees Confusion matrix, without normalization [[38 4] [ 2 70]] ExtraTrees Normalized confusion matrix [[0.9 0.1 ] [0.03 0.97]]
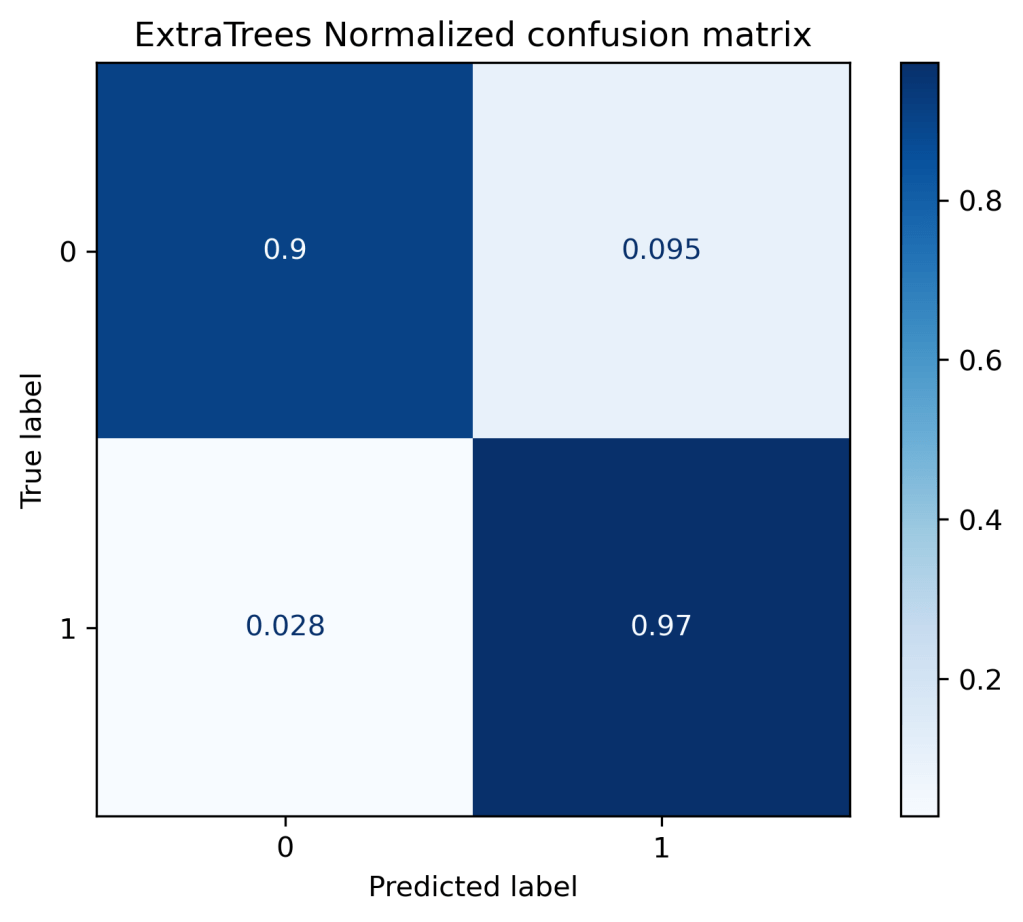
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_test_pred_gb = gb.predict(X_test)
print(accuracy_score(y_test_pred_gb, y_test))
print(confusion_matrix(y_test_pred_gb, y_test))
print(classification_report(y_test_pred_gb, y_test))
0.9210526315789473
[[38 5]
[ 4 67]]
precision recall f1-score support
0 0.90 0.88 0.89 43
1 0.93 0.94 0.94 71
accuracy 0.92 114
macro avg 0.92 0.91 0.92 114
weighted avg 0.92 0.92 0.92 114
classifier = gb.fit(X_train, y_train)
titles_options = [
(“Gradient Boosting Confusion Matrix, without normalization”, None),
(“Gradient Boosting Confusion Matrix”, “true”),
]
for title, normalize in titles_options:
disp = ConfusionMatrixDisplay.from_estimator(
classifier,
X_test,
y_test,
display_labels=class_names,
cmap=plt.cm.Blues,
normalize=normalize,
)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.savefig(‘confmatrixgb.png’, dpi=300, bbox_inches=’tight’)
Gradient Boosting Confusion Matrix, without normalization [[38 4] [ 4 68]] Gradient Boosting Confusion Matrix [[0.9 0.1 ] [0.06 0.94]]
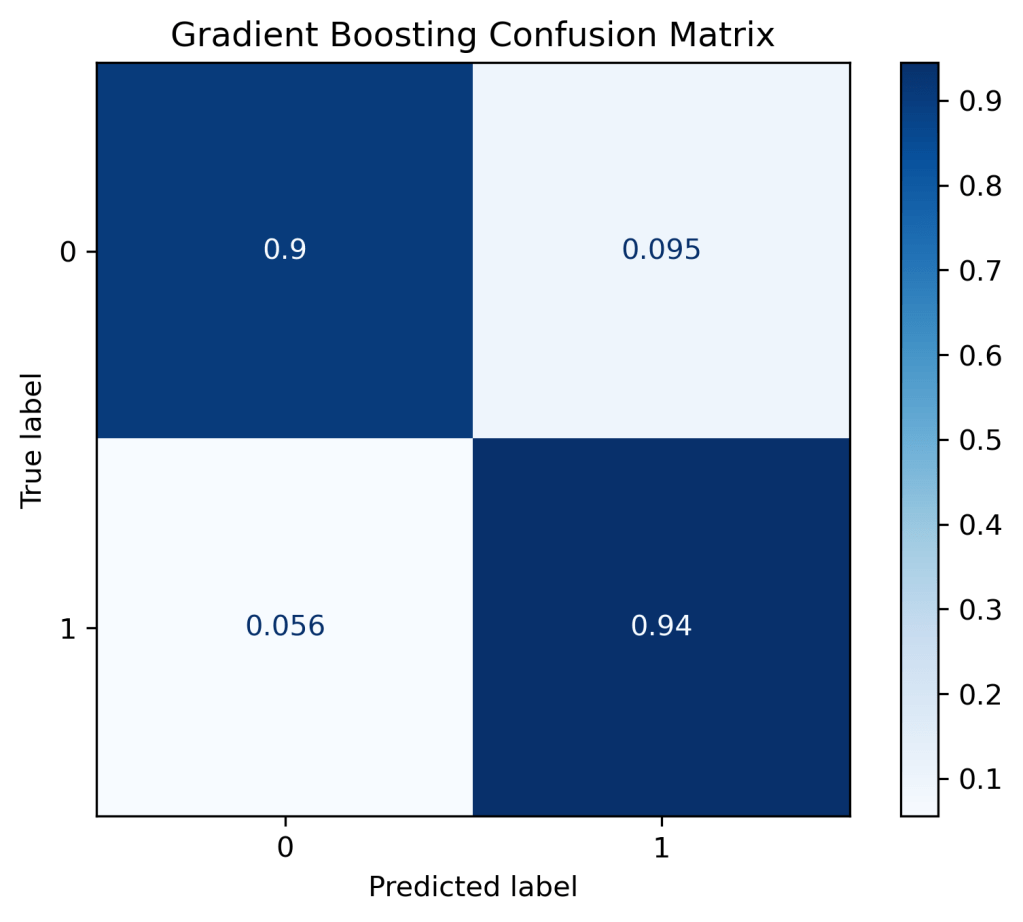
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
y_test_pred_rf = rf.predict(X_test)
print(accuracy_score(y_test_pred_rf, y_test))
print(confusion_matrix(y_test_pred_rf, y_test))
print(classification_report(y_test_pred_rf, y_test))
0.9385964912280702
[[38 3]
[ 4 69]]
precision recall f1-score support
0 0.90 0.93 0.92 41
1 0.96 0.95 0.95 73
accuracy 0.94 114
macro avg 0.93 0.94 0.93 114
weighted avg 0.94 0.94 0.94 114

knn = KNeighborsClassifier(n_neighbors=6)
knn.fit(X_train, y_train)
y_test_pred_knn = knn.predict(X_test)
print(accuracy_score(y_test_pred_knn, y_test))
print(confusion_matrix(y_test_pred_knn, y_test))
print(classification_report(y_test_pred_knn, y_test))
0.9385964912280702
[[39 4]
[ 3 68]]
precision recall f1-score support
0 0.93 0.91 0.92 43
1 0.94 0.96 0.95 71
accuracy 0.94 114
macro avg 0.94 0.93 0.93 114
weighted avg 0.94 0.94 0.94 114
classifier = knn.fit(X_train, y_train)
titles_options = [
(“KNN Confusion Matrix, without normalization”, None),
(“KNN Confusion Matrix”, “true”),
]
for title, normalize in titles_options:
disp = ConfusionMatrixDisplay.from_estimator(
classifier,
X_test,
y_test,
display_labels=class_names,
cmap=plt.cm.Blues,
normalize=normalize,
)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.savefig(‘confmatrixknn.png’, dpi=300, bbox_inches=’tight’)
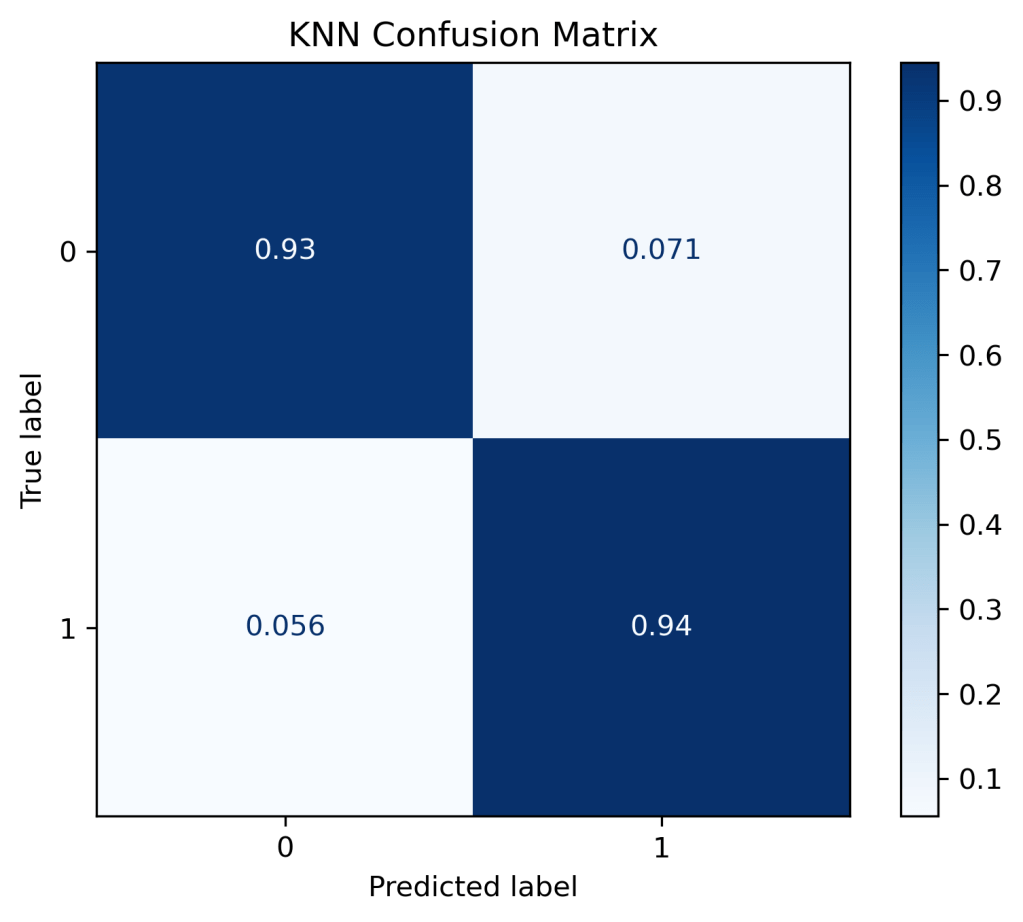
ROC-AUC Score
Let’s summarize our ROC curve analysis with the focus on the AUC score
model = LogisticRegression()
model.fit(X_train, y_train)
probs = model.predict_proba(X_test)
probs = probs[:, 1]
auc = roc_auc_score(y_test, probs)
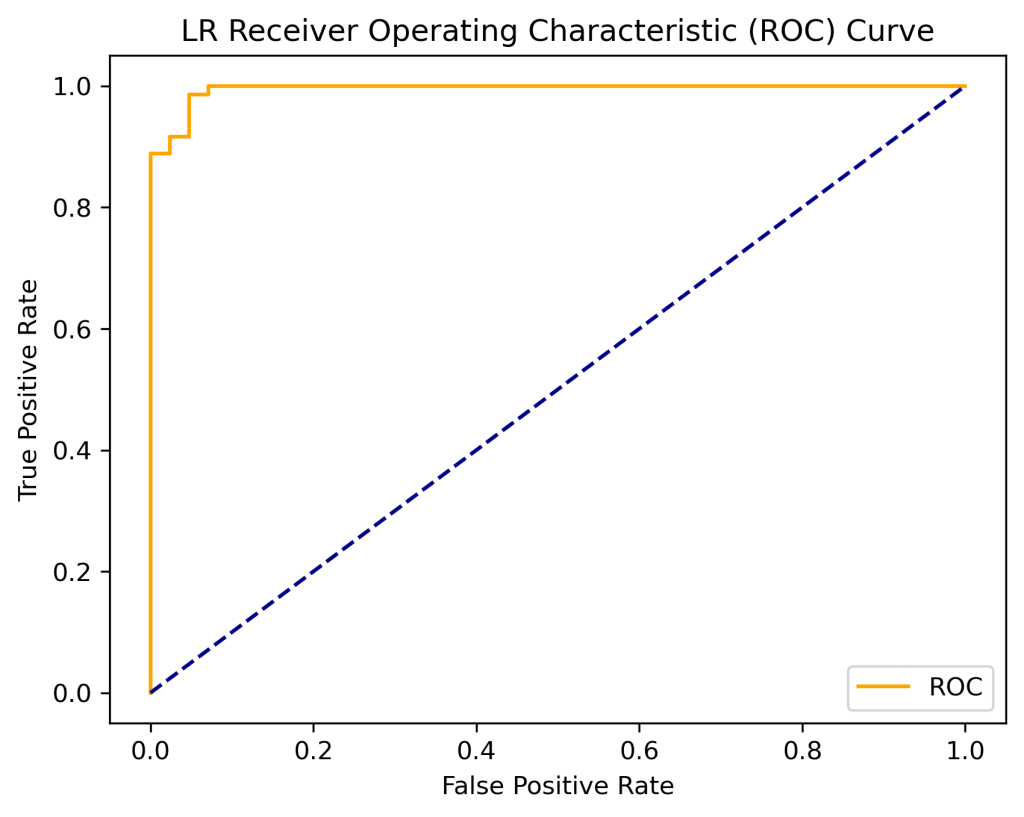
print(‘LR AUC: %.2f’ % auc)
LR AUC: 1.00
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color=’orange’, label=’ROC’)
plt.plot([0, 1], [0, 1], color=’darkblue’, linestyle=’–‘)
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘LR Receiver Operating Characteristic (ROC) Curve’)
plt.legend()
plt.savefig(‘rocauc_lr.png’, dpi=300, bbox_inches=’tight’)
fpr, tpr, thresholds = roc_curve(y_test, probs)
plot_roc_curve(fpr, tpr)
model = ExtraTreesClassifier()
model.fit(X_train, y_train)
probs = model.predict_proba(X_test)
probs = probs[:, 1]
auc = roc_auc_score(y_test, probs)
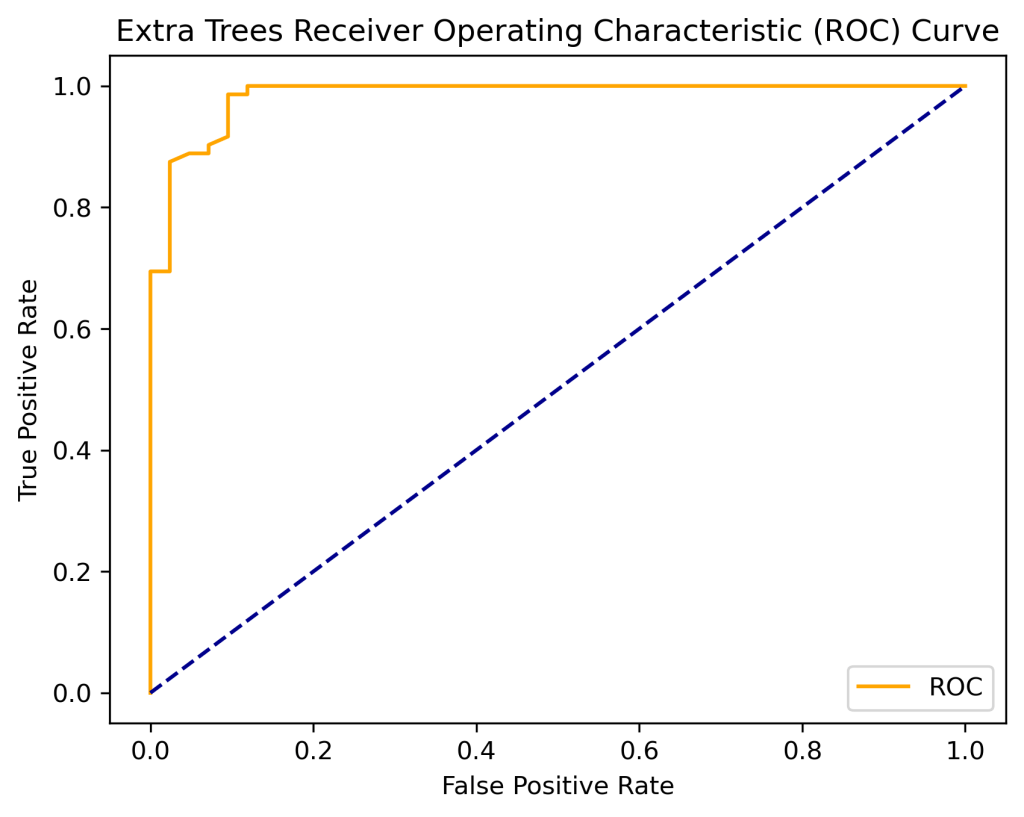
print(‘Extra Trees AUC: %.2f’ % auc)
Extra Trees AUC: 0.98
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color=’orange’, label=’ROC’)
plt.plot([0, 1], [0, 1], color=’darkblue’, linestyle=’–‘)
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘Extra Trees Receiver Operating Characteristic (ROC) Curve’)
plt.legend()
plt.savefig(‘rocauc_xt.png’, dpi=300, bbox_inches=’tight’)
fpr, tpr, thresholds = roc_curve(y_test, probs)
plot_roc_curve(fpr, tpr)
model = GradientBoostingClassifier()
model.fit(X_train, y_train)
probs = model.predict_proba(X_test)
probs = probs[:, 1]
auc = roc_auc_score(y_test, probs)
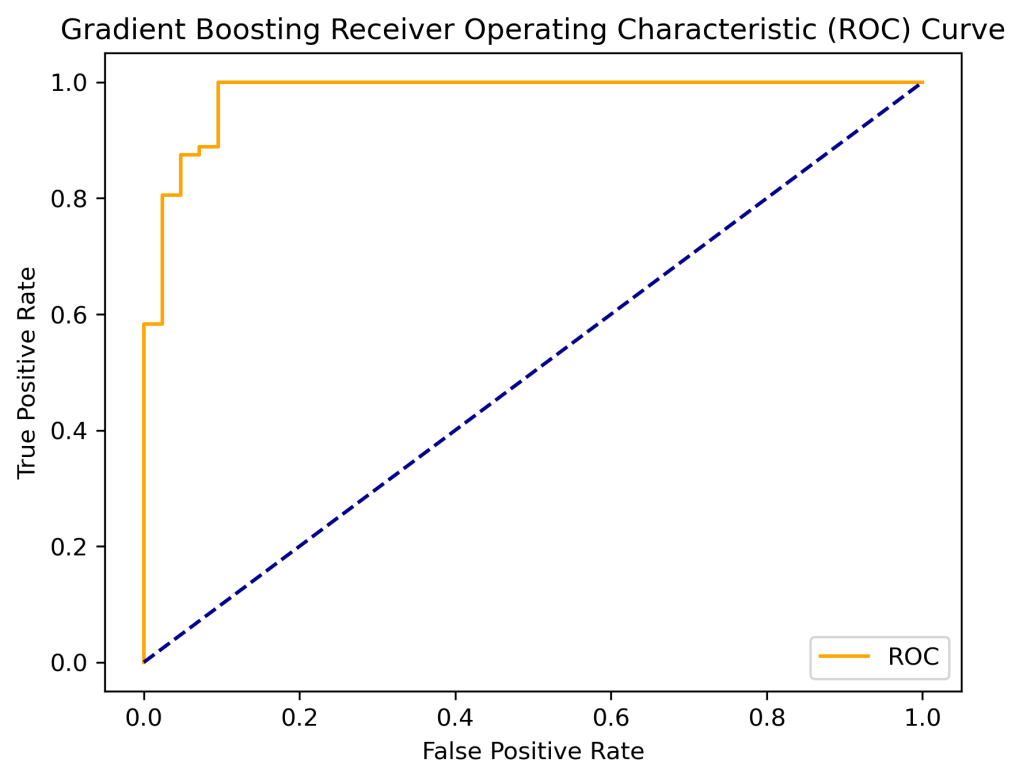
print(‘Gradient Boosting AUC: %.2f’ % auc)
Gradient Boosting AUC: 0.98
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color=’orange’, label=’ROC’)
plt.plot([0, 1], [0, 1], color=’darkblue’, linestyle=’–‘)
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘Gradient Boosting Receiver Operating Characteristic (ROC) Curve’)
plt.legend()
plt.savefig(‘rocauc_gb.png’, dpi=300, bbox_inches=’tight’)
fpr, tpr, thresholds = roc_curve(y_test, probs)
plot_roc_curve(fpr, tpr)
model = RandomForestClassifier()
model.fit(X_train, y_train)
probs = model.predict_proba(X_test)
probs = probs[:, 1]
auc = roc_auc_score(y_test, probs)
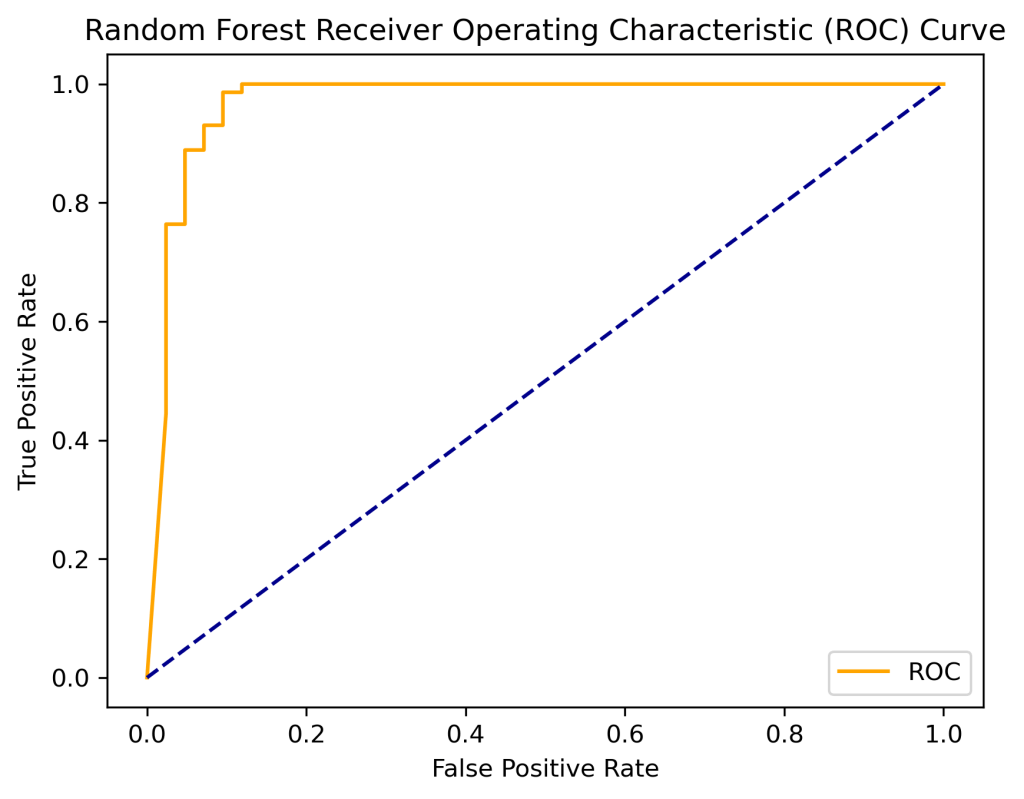
print(‘Random Forest AUC: %.2f’ % auc)
Random Forest AUC: 0.97
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color=’orange’, label=’ROC’)
plt.plot([0, 1], [0, 1], color=’darkblue’, linestyle=’–‘)
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘Random Forest Receiver Operating Characteristic (ROC) Curve’)
plt.legend()
plt.savefig(‘rocauc_rf.png’, dpi=300, bbox_inches=’tight’)
fpr, tpr, thresholds = roc_curve(y_test, probs)
plot_roc_curve(fpr, tpr)
model = KNeighborsClassifier(n_neighbors=6)
model.fit(X_train, y_train)
probs = model.predict_proba(X_test)
probs = probs[:, 1]
auc = roc_auc_score(y_test, probs)
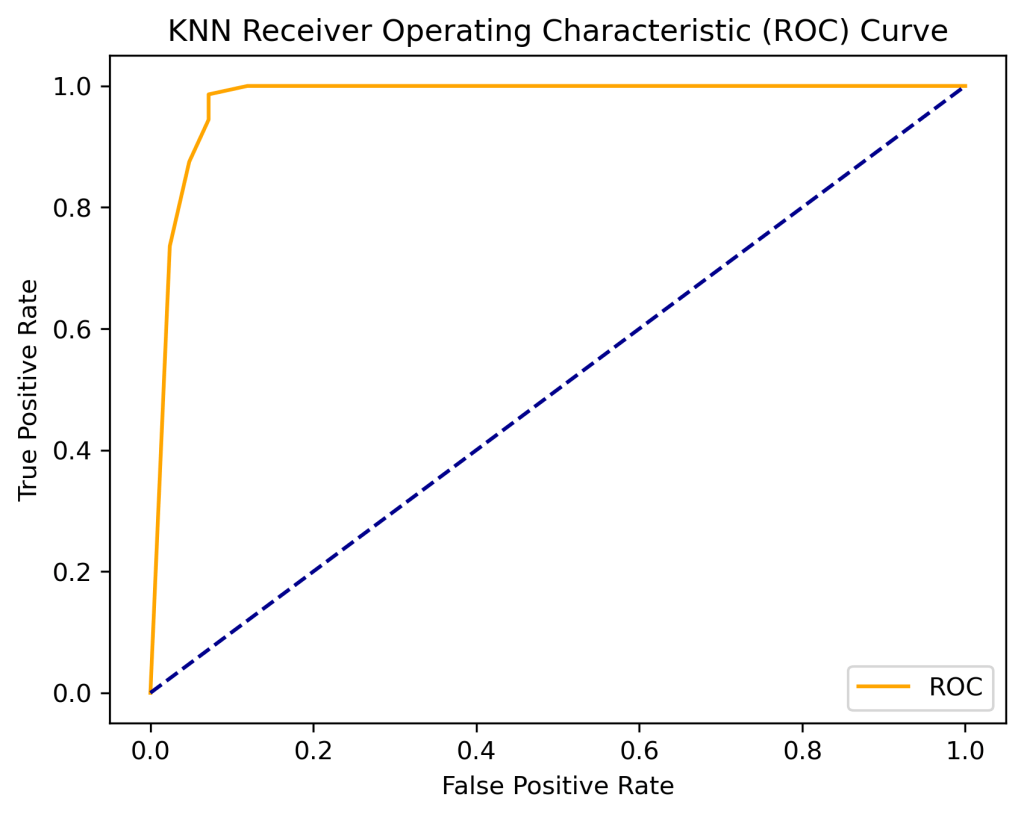
print(‘KNN AUC: %.2f’ % auc)
KNN AUC: 0.98
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color=’orange’, label=’ROC’)
plt.plot([0, 1], [0, 1], color=’darkblue’, linestyle=’–‘)
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘KNN Receiver Operating Characteristic (ROC) Curve’)
plt.legend()
plt.savefig(‘rocauc_knn.png’, dpi=300, bbox_inches=’tight’)
fpr, tpr, thresholds = roc_curve(y_test, probs)
plot_roc_curve(fpr, tpr)
Summary
- Input BC Wisconsin diagnostic dataset with 30 attributes and 569 instances
- ML workflow: reading input data, importing key libraries, Exploratory Data Analysis (EDA), ML data preparation (scaling and test/train/target/features data splitting), ML model training, test data predictions, and classification QC analysis using available metrics and Scikit-Plot curves
- Scope: QC comparison of Scikit-Learn binary classifiers – Logistic Regression (LR), Extra Trees (ET), Gradient Boosting (GB), Random Forest (RF), and KNN
- Most dominant features: worst texture/compactness (RF), mean area (ET), worst concave points (ET), and worst perimeter/area (GB)
- Least dominant features: worst smoothness/symmetry (ET, GB), concave points error (ET, GB), concavity/compactness error (RF), mean fractal dimension (RF), and mean radius (GB)
- Cumulative gains and lift curves (ML performance) are identical for all classifiers
- PCA: Silhouette score 0.116, number of clusters is 3 (elbow plot), 0.982 explained variance ratio for first 1 components
- PCA 2D projection: good separation of classes 0 and 1.
- ML Classification Report:
| No. | Metrics | LR | ET | GB | RF | KNN |
| 1 | Training score 1.0 – overfitting or high variance (learning curves) | No | Yes | Yes | Yes | No |
| 2 | Cross-validation score (train data learning curves, 100 training examples) | 0.99 | 0.96 | 0.95 | 0.95 | 0.97 |
| 3 | Accuracy (train data) | 0.96 | 0.94 | 0.92 | 0.93 | 0.94 |
| 4 | ROC AUC score | 1.00 | 0.98 | 0.98 | 1.00 | 0.98 |
| 5 | F1-score class 1 | 0.97 | 0.95 | 0.94 | 0.94 | 0.95 |
| 6 | Precision class 1 | 0.97 | 0.97 | 0.93 | 0.94 | 0.94 |
| 7 | Recall class 1 | 0.97 | 0.93 | 0.94 | 0.94 | 0.96 |
| 8 | FP (class 1 prediction of class 0) | 0.05 | 0.09 | 0.09 | 0.09 | 0.07 |
| 9 | FN (class 0 prediction of class 1) | 0.03 | 0.03 | 0.05 | 0.04 | 0.05 |
| 10 | Precision-recall curve area class 1 | 0.99 | 0.98 | 0.98 | 0.97 | 0.97 |
| 11 | ROC curve area class 1 | 1.0 | 0.98 | 0.98 | 0.97 | 0.98 |
| 12 | KS test | 0.94 at 0.44 | 0.89 at 0.46 | 0.88 at 0.06 | 0.88 at 0.27 | 0.95 at 0.33 |
| 13 | Calibration curves ranking | 4 | 2 | 5 | 3 | 1 |
- See the code in Github Rep
Explore More
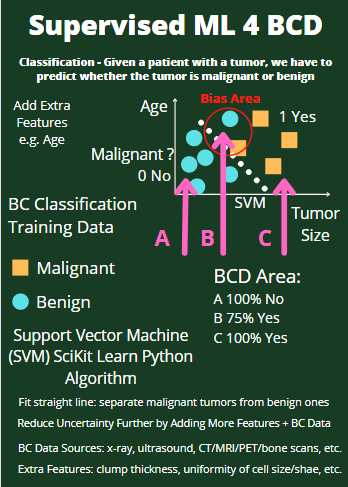
- Supervised ML/AI Breast Cancer Diagnostics (BCD) – The Power of HealthTech
- HealthTech ML/AI Use-Cases
- HealthTech ML/AI Q3 ’22 Round-Up
- A Comparative Analysis of Breast Cancer ML/AI Binary Classifications
- Breast Cancer ML Classification – Logistic Regression vs Gradient Boosting with Hyperparameter Optimization (HPO)
- How to build a basic breast cancer model — machine learning
Infographics









Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.
DonateDonate monthlyDonate yearly

Leave a comment