According to WHO, Breast Cancer (BC) is the leading cause of death among women worldwide. The present study optimizes the use of supervised Machine Learning (ML) algorithms for detecting, analyzing, and classifying BC.
Following previous BC ML/AI case studies (cf. Read More), our objective is to compare Logistic Regression (LR) against Gradient Boosting (GB) Classifier within the Hyperparameter Optimization (HPO) loop given by GridSearchCV.
We use the publicly available BC dataset from the University of Wisconsin Hospitals, Madison, Wisconsin, USA.
Let’s open the Jupyter IDE Notebook
and begin our project by setting the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
and importing the key libraries
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from collections import OrderedDict
from sklearn import datasets
from sklearn.preprocessing import label_binarize, LabelBinarizer
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import classification_report
from sklearn.metrics import roc_curve, auc
DISPLAY_PRECISION = 4
pd.set_option(“display.precision”, DISPLAY_PRECISION)
Let’s load the BC dataset and print its detailed description
dat = datasets.load_breast_cancer()
print(dat.DESCR)
.. _breast_cancer_dataset:
Breast cancer wisconsin (diagnostic) dataset
--------------------------------------------
**Data Set Characteristics:**
:Number of Instances: 569
:Number of Attributes: 30 numeric, predictive attributes and the class
:Attribute Information:
- radius (mean of distances from center to points on the perimeter)
- texture (standard deviation of gray-scale values)
- perimeter
- area
- smoothness (local variation in radius lengths)
- compactness (perimeter^2 / area - 1.0)
- concavity (severity of concave portions of the contour)
- concave points (number of concave portions of the contour)
- symmetry
- fractal dimension ("coastline approximation" - 1)
The mean, standard error, and "worst" or largest (mean of the three
worst/largest values) of these features were computed for each image,
resulting in 30 features. For instance, field 0 is Mean Radius, field
10 is Radius SE, field 20 is Worst Radius.
- class:
- WDBC-Malignant
- WDBC-Benign
:Summary Statistics:
===================================== ====== ======
Min Max
===================================== ====== ======
radius (mean): 6.981 28.11
texture (mean): 9.71 39.28
perimeter (mean): 43.79 188.5
area (mean): 143.5 2501.0
smoothness (mean): 0.053 0.163
compactness (mean): 0.019 0.345
concavity (mean): 0.0 0.427
concave points (mean): 0.0 0.201
symmetry (mean): 0.106 0.304
fractal dimension (mean): 0.05 0.097
radius (standard error): 0.112 2.873
texture (standard error): 0.36 4.885
perimeter (standard error): 0.757 21.98
area (standard error): 6.802 542.2
smoothness (standard error): 0.002 0.031
compactness (standard error): 0.002 0.135
concavity (standard error): 0.0 0.396
concave points (standard error): 0.0 0.053
symmetry (standard error): 0.008 0.079
fractal dimension (standard error): 0.001 0.03
radius (worst): 7.93 36.04
texture (worst): 12.02 49.54
perimeter (worst): 50.41 251.2
area (worst): 185.2 4254.0
smoothness (worst): 0.071 0.223
compactness (worst): 0.027 1.058
concavity (worst): 0.0 1.252
concave points (worst): 0.0 0.291
symmetry (worst): 0.156 0.664
fractal dimension (worst): 0.055 0.208
===================================== ====== ======
:Missing Attribute Values: None
:Class Distribution: 212 - Malignant, 357 - Benign
:Creator: Dr. William H. Wolberg, W. Nick Street, Olvi L. Mangasarian
:Donor: Nick Street
:Date: November, 1995
This is a copy of UCI ML Breast Cancer Wisconsin (Diagnostic) datasets.
https://goo.gl/U2Uwz2
Features are computed from a digitized image of a fine needle
aspirate (FNA) of a breast mass. They describe
characteristics of the cell nuclei present in the image.
Separating plane described above was obtained using
Multisurface Method-Tree (MSM-T) [K. P. Bennett, "Decision Tree
Construction Via Linear Programming." Proceedings of the 4th
Midwest Artificial Intelligence and Cognitive Science Society,
pp. 97-101, 1992], a classification method which uses linear
programming to construct a decision tree. Relevant features
were selected using an exhaustive search in the space of 1-4
features and 1-3 separating planes.
The actual linear program used to obtain the separating plane
in the 3-dimensional space is that described in:
[K. P. Bennett and O. L. Mangasarian: "Robust Linear
Programming Discrimination of Two Linearly Inseparable Sets",
Optimization Methods and Software 1, 1992, 23-34].
This database is also available through the UW CS ftp server:
ftp ftp.cs.wisc.edu
cd math-prog/cpo-dataset/machine-learn/WDBC/
.. topic:: References
- W.N. Street, W.H. Wolberg and O.L. Mangasarian. Nuclear feature extraction
for breast tumor diagnosis. IS&T/SPIE 1993 International Symposium on
Electronic Imaging: Science and Technology, volume 1905, pages 861-870,
San Jose, CA, 1993.
- O.L. Mangasarian, W.N. Street and W.H. Wolberg. Breast cancer diagnosis and
prognosis via linear programming. Operations Research, 43(4), pages 570-577,
July-August 1995.
- W.H. Wolberg, W.N. Street, and O.L. Mangasarian. Machine learning techniques
to diagnose breast cancer from fine-needle aspirates. Cancer Letters 77 (1994)
163-171.
Let’s look at the basic structure of our input data
print(“The sklearn breast cancer dataset keys:”)
print(dat.keys()) # dict_keys([‘target_names’, ‘target’, ‘feature_names’, ‘data’, ‘DESCR’])
print(“—“)
li_classes = [dat.target_names[1], dat.target_names[0]]
li_target = [1 if x==0 else 0 for x in list(dat.target)]
li_ftrs = list(dat.feature_names)
print(“There are 2 target classes:”)
print(“li_classes”, li_classes)
print(“—“)
print(“Target class distribution from a total of %d target values:” % len(li_target))
print(pd.Series(li_target).value_counts())
print(“—“)
df_all = pd.DataFrame(dat.data[:,:], columns=li_ftrs)
print(“Describe dataframe, first 6 columns:”)
print(df_all.iloc[:,:6].describe().to_string())
The sklearn breast cancer dataset keys:
dict_keys(['data', 'target', 'frame', 'target_names', 'DESCR', 'feature_names', 'filename', 'data_module'])
---
There are 2 target classes:
li_classes ['benign', 'malignant']
---
Target class distribution from a total of 569 target values:
0 357
1 212
dtype: int64
---
Describe dataframe, first 6 columns:
mean radius mean texture mean perimeter mean area mean smoothness mean compactness
count 569.0000 569.0000 569.000 569.0000 569.0000 569.0000
mean 14.1273 19.2896 91.969 654.8891 0.0964 0.1043
std 3.5240 4.3010 24.299 351.9141 0.0141 0.0528
min 6.9810 9.7100 43.790 143.5000 0.0526 0.0194
25% 11.7000 16.1700 75.170 420.3000 0.0864 0.0649
50% 13.3700 18.8400 86.240 551.1000 0.0959 0.0926
75% 15.7800 21.8000 104.100 782.7000 0.1053 0.1304
max 28.1100 39.2800 188.500 2501.0000 0.1634 0.3454
Let’s perform test/train data splitting with test_size=0.3
TEST_SIZE_RATIO = 0.3
Setup X and y:
X = df_all
y = pd.Series(li_target)
X_train_0, X_test_0, y_train, y_test = train_test_split(X, y, test_size=TEST_SIZE_RATIO, random_state=0)
print(“X_train_0.shape, y_train.shape”, X_train_0.shape, y_train.shape)
print(“X_test_0.shape, y_test.shape”, X_test_0.shape, y_test.shape)
X_train_0.shape, y_train.shape (398, 30) (398,) X_test_0.shape, y_test.shape (171, 30) (171,)
Let’s build the correlation matrix for the train data using the function
def correlation_matrix(y, X, is_plot=False):
# Calculate and plot the correlation symmetrical matrix
# Return:
# yX – concatenated data
# yX_corr – correlation matrix, pearson correlation of values from -1 to +1
# yX_abs_corr – correlation matrix, absolute values
yX = pd.concat([y, X], axis=1)
yX = yX.rename(columns={0: ‘TARGET’}) # rename first column
print(“Function correlation_matrix: X.shape, y.shape, yX.shape:”, X.shape, y.shape, yX.shape)
print()
# Get feature correlations and transform to dataframe
yX_corr = yX.corr(method=’pearson’)
# Convert to abolute values
yX_abs_corr = np.abs(yX_corr)
if is_plot:
plt.figure(figsize=(10, 10))
plt.imshow(yX_abs_corr, cmap=’RdYlGn’, interpolation=’none’, aspect=’auto’)
plt.colorbar()
plt.xticks(range(len(yX_abs_corr)), yX_abs_corr.columns, rotation=’vertical’)
plt.yticks(range(len(yX_abs_corr)), yX_abs_corr.columns);
plt.suptitle(‘Pearson Correlation Heat Map (absolute values)’, fontsize=15, fontweight=’bold’)
#plt.show()
plt.savefig(‘corrmatrixafter.png’)
return yX, yX_corr, yX_abs_corr
Let’s call this function
yX, yX_corr, yX_abs_corr = correlation_matrix(y_train, X_train_0, is_plot=True)
Function correlation_matrix: X.shape, y.shape, yX.shape: (398, 30) (398,) (398, 31)
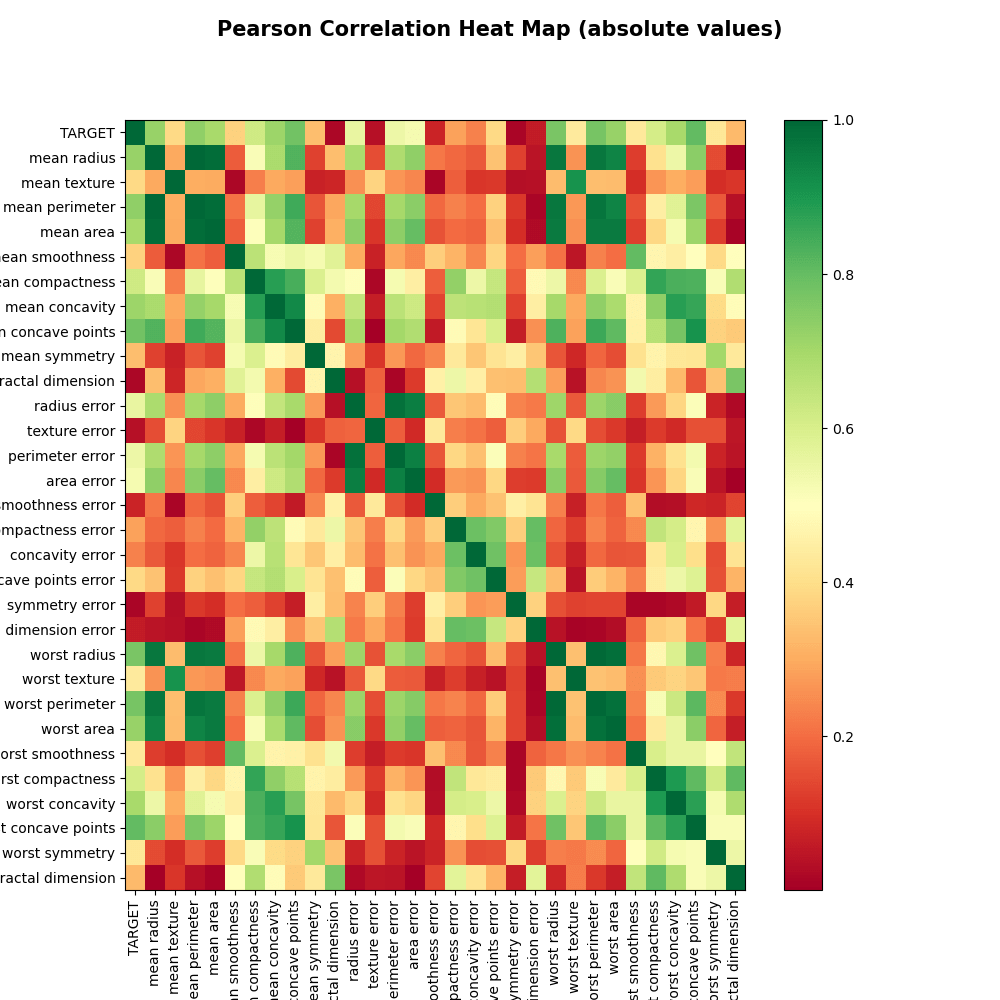
Let’s select the BC model features within the range (0.1, 0.8)
CORRELATION_MIN = 0.1
Sort features by their Pearson correlation with the target value
s_corr_target = yX_abs_corr[‘TARGET’]
s_corr_target_sort = s_corr_target.sort_values(ascending=False)
Only use features with a minimum Pearson correlation with the target of 0.1
s_low_correlation_ftrs = s_corr_target_sort[s_corr_target_sort <= CORRELATION_MIN]
print(“Removed %d low correlation features:” % len(s_low_correlation_ftrs))
for i,v in enumerate(s_low_correlation_ftrs):
print(i,np.round(v, DISPLAY_PRECISION), s_low_correlation_ftrs.index[i])
print(“—“)
s_corr_target_sort = s_corr_target_sort[s_corr_target_sort > CORRELATION_MIN]
print(“Remaining %d feature correlations:” % (len(s_corr_target_sort)-1))
for i,v in enumerate(s_corr_target_sort):
ftr = s_corr_target_sort.index[i]
if ftr == ‘TARGET’:
continue
print(i,np.round(v, DISPLAY_PRECISION), ftr)
Removed 5 low correlation features: 0 0.0778 smoothness error 1 0.0622 fractal dimension error 2 0.0373 texture error 3 0.0187 mean fractal dimension 4 0.0154 symmetry error --- Remaining 25 feature correlations: 1 0.8024 worst concave points 2 0.7794 mean concave points 3 0.7723 worst perimeter 4 0.7661 worst radius 5 0.7318 mean perimeter 6 0.7194 worst area 7 0.719 mean radius 8 0.7111 mean concavity 9 0.6947 worst concavity 10 0.6932 mean area 11 0.6196 mean compactness 12 0.6062 worst compactness 13 0.5574 radius error 14 0.5465 perimeter error 15 0.5264 area error 16 0.4347 worst texture 17 0.4324 worst smoothness 18 0.4255 worst symmetry 19 0.3902 concave points error 20 0.3883 mean texture 21 0.373 mean smoothness 22 0.3361 mean symmetry 23 0.3272 worst fractal dimension 24 0.2823 compactness error 25 0.2322 concavity error
CORRELATION_MAX = 0.8
Remove features that have a low correlation with the target
li_X1_cols = list(set(s_corr_target_sort.index) – set(s_low_correlation_ftrs.index))
li_X1_cols.remove(‘TARGET’)
Build the correlation matrix for the reduced X
X1 = X_train_0[li_X1_cols]
yX1, yX_corr1, yX_abs_corr1 = correlation_matrix(y_train, X1, is_plot=False)
Get all the feature pairs
Xcorr1 = yX_abs_corr1.iloc[1:,1:]
s_pairs = Xcorr1.unstack()
print(“s_pairs.shape”, s_pairs.shape)
s_pairs = np.round(s_pairs, decimals=DISPLAY_PRECISION)
Sort all the pairs by highest correlation values
s_pairs_sorted = s_pairs.sort_values(ascending=False)
s_pairs_sorted = s_pairs_sorted[(s_pairs_sorted != 1) & (s_pairs_sorted > CORRELATION_MAX)] # leave only the top matches that are not identical features
Convert to a list of name tuples e.g. (‘mean radius’, ‘mean perimeter’)
li_corr_pairs = s_pairs_sorted.index.tolist()
print(“len(li_corr_pairs):”, len(li_corr_pairs))
print(“li_corr_pairs[:10]”, li_corr_pairs[:10])
Remove features that have a low correlation with the target
li_X1_cols = list(set(s_corr_target_sort.index) – set(s_low_correlation_ftrs.index))
li_X1_cols.remove(‘TARGET’)
Build the correlation matrix for the reduced X
X1 = X_train_0[li_X1_cols]
yX1, yX_corr1, yX_abs_corr1 = correlation_matrix(y_train, X1, is_plot=False)
Get all the feature pairs
Xcorr1 = yX_abs_corr1.iloc[1:,1:]
s_pairs = Xcorr1.unstack()
print(“s_pairs.shape”, s_pairs.shape)
s_pairs = np.round(s_pairs, decimals=DISPLAY_PRECISION)
Sort all the pairs by highest correlation values
s_pairs_sorted = s_pairs.sort_values(ascending=False)
s_pairs_sorted = s_pairs_sorted[(s_pairs_sorted != 1) & (s_pairs_sorted > CORRELATION_MAX)] # leave only the top matches that are not identical features
Convert to a list of name tuples e.g. (‘mean radius’, ‘mean perimeter’)
li_corr_pairs = s_pairs_sorted.index.tolist()
print(“len(li_corr_pairs):”, len(li_corr_pairs))
print(“li_corr_pairs[:10]”, li_corr_pairs[:10])
Function correlation_matrix: X.shape, y.shape, yX.shape: (398, 25) (398,) (398, 26)
s_pairs.shape (625,)
len(li_corr_pairs): 80
li_corr_pairs[:10] [('mean radius', 'mean perimeter'), ('mean perimeter', 'mean radius'), ('worst perimeter', 'worst radius'), ('worst radius', 'worst perimeter'), ('mean area', 'mean radius'), ('mean radius', 'mean area'), ('mean area', 'mean perimeter'), ('mean perimeter', 'mean area'), ('worst area', 'worst radius'), ('worst radius', 'worst area')]
Let’s build the list of features to remove
li_remove_pair_ftrs = []
li_remove_scores = []
for tup in li_corr_pairs:
s0 = s_corr_target_sort.loc[tup[0]]
s1 = s_corr_target_sort.loc[tup[1]]
remove_ftr = tup[1] if s1 < s0 else tup[0] # get the feature that is less correlated with the target
if remove_ftr not in li_remove_pair_ftrs:
li_remove_pair_ftrs.append(remove_ftr)
di = {‘ftr_0’:tup[0], ‘ftr_1’:tup[1], ‘score_0’:s0, ‘score_1’:s1, ‘FEATURE_TO_REMOVE’:remove_ftr}
li_remove_scores.append(OrderedDict(di))
df_remove_scores = pd.DataFrame(li_remove_scores)
print(“Removing %d features (see last column):” % len(li_remove_pair_ftrs))
print(df_remove_scores.to_string())
print(“—“)
Remove the features that were found in the above procedure
li_X2_cols = list(set(li_X1_cols) – set(li_remove_pair_ftrs))
li_X2_cols.sort()
print(“Remaining %d features:” % (len(li_X2_cols)))
for i,v in enumerate(s_corr_target_sort):
ftr = s_corr_target_sort.index[i]
if ftr in li_X2_cols:
print(i,np.round(v, DISPLAY_PRECISION), ftr)
Removing 16 features (see last column):
ftr_0 ftr_1 score_0 score_1 FEATURE_TO_REMOVE
0 mean radius mean perimeter 0.7190 0.7318 mean radius
1 worst perimeter worst radius 0.7723 0.7661 worst radius
2 mean area mean radius 0.6932 0.7190 mean area
3 worst area worst radius 0.7194 0.7661 worst area
4 radius error perimeter error 0.5574 0.5465 perimeter error
5 worst perimeter mean perimeter 0.7723 0.7318 mean perimeter
6 radius error area error 0.5574 0.5264 area error
7 mean concavity mean concave points 0.7111 0.7794 mean concavity
8 mean texture worst texture 0.3883 0.4347 mean texture
9 mean concave points worst concave points 0.7794 0.8024 mean concave points
10 worst compactness worst concavity 0.6062 0.6947 worst compactness
11 mean concavity mean compactness 0.7111 0.6196 mean compactness
12 mean concavity worst concavity 0.7111 0.6947 worst concavity
13 worst perimeter mean concave points 0.7723 0.7794 worst perimeter
14 worst compactness worst fractal dimension 0.6062 0.3272 worst fractal dimension
15 mean smoothness worst smoothness 0.3730 0.4324 mean smoothness
---
Remaining 9 features:
1 0.8024 worst concave points
13 0.5574 radius error
16 0.4347 worst texture
17 0.4324 worst smoothness
18 0.4255 worst symmetry
19 0.3902 concave points error
22 0.3361 mean symmetry
24 0.2823 compactness error
25 0.2322 concavity error
Let’s calculate the correlation matrix on the subset of features
X2 = X1[li_X2_cols]
print(“After the pair feature reduction, X2.shape:”, X2.shape)
yX2, yX_corr2, yX_abs_corr2 = correlation_matrix(y_train, X2)
Recalculate the correlation matrix to plot the TARGET values grouped by the order of correlation
s_X3_cols = yX_abs_corr2[‘TARGET’].sort_values(ascending=False)
li_X3_cols = s_X3_cols.index.tolist()
print(“Remaining features:”)
print(s_X3_cols)
print(“—“)
li_X3_cols.remove(‘TARGET’)
X3 = X2[li_X3_cols]
print(“After the pair feature reduction, X2.shape:”, X3.shape)
yX3, yX_corr3, yX_abs_corr3 = correlation_matrix(y_train, X3, is_plot=True)
X_train = X3
X_test = X_test_0[li_X3_cols]
After the pair feature reduction, X2.shape: (398, 9) Function correlation_matrix: X.shape, y.shape, yX.shape: (398, 9) (398,) (398, 10) Remaining features: TARGET 1.0000 worst concave points 0.8024 radius error 0.5574 worst texture 0.4347 worst smoothness 0.4324 worst symmetry 0.4255 concave points error 0.3902 mean symmetry 0.3361 compactness error 0.2823 concavity error 0.2322 Name: TARGET, dtype: float64 --- After the pair feature reduction, X2.shape: (398, 9) Function correlation_matrix: X.shape, y.shape, yX.shape: (398, 9) (398,) (398, 10)
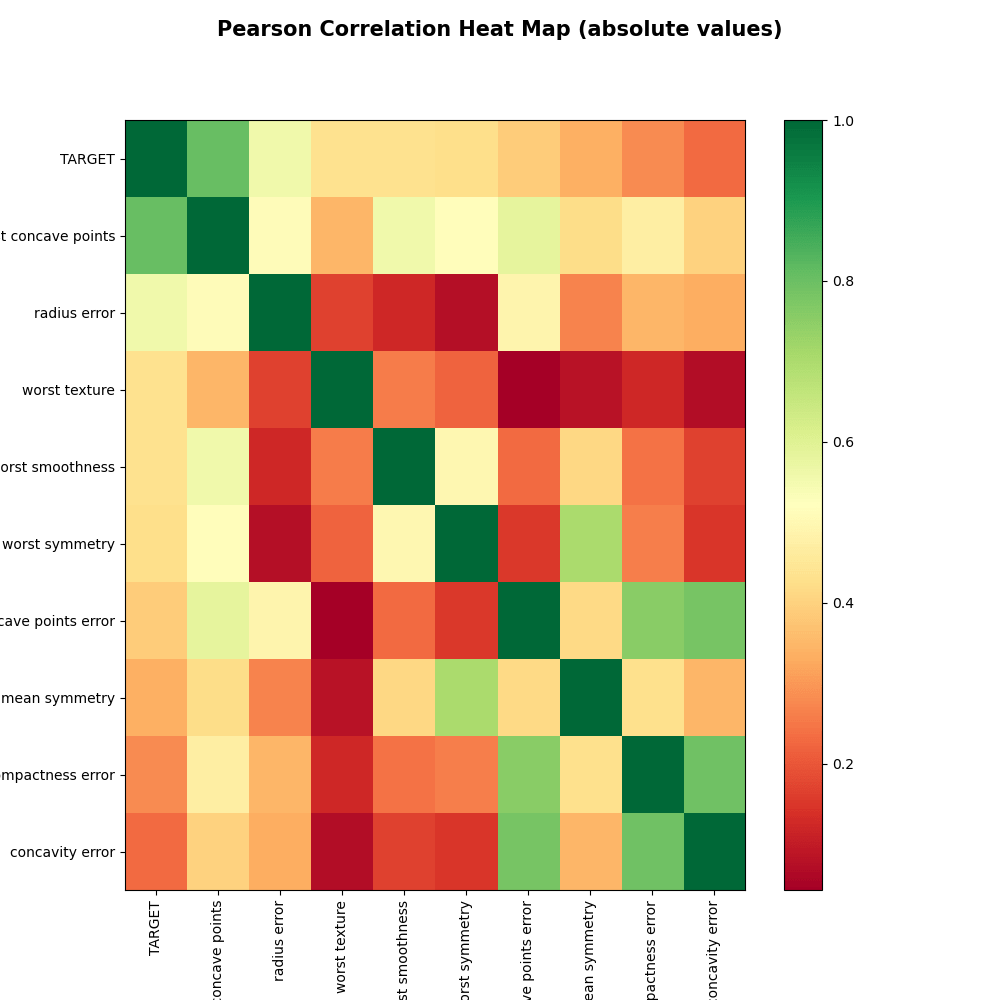
Let’s create the sns joint plot for the 9 remaining dominant features as follows:
sns.jointplot(yX3[‘concavity error’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterconcavityerror.png’)
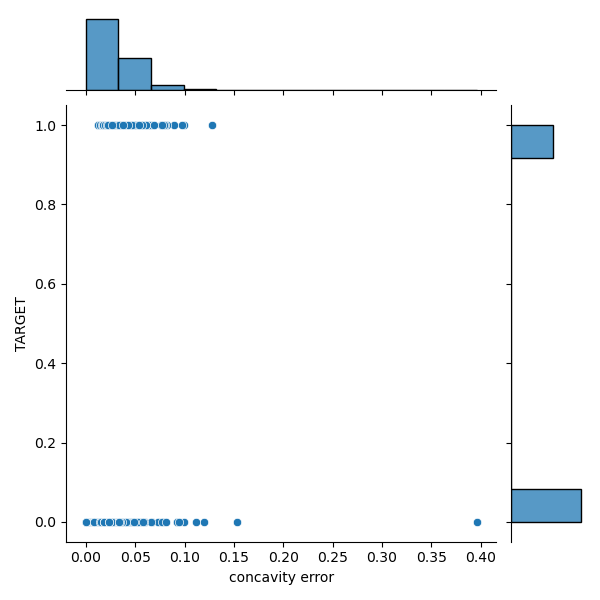
sns.jointplot(yX3[‘compactness error’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscattercompactnesserror.png’)

sns.jointplot(yX3[‘mean symmetry’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscattermeansymmetry.png’)

sns.jointplot(yX3[‘concave points error’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterconcavepointserror.png’)

sns.jointplot(yX3[‘worst symmetry’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterworstsymmetry.png’)
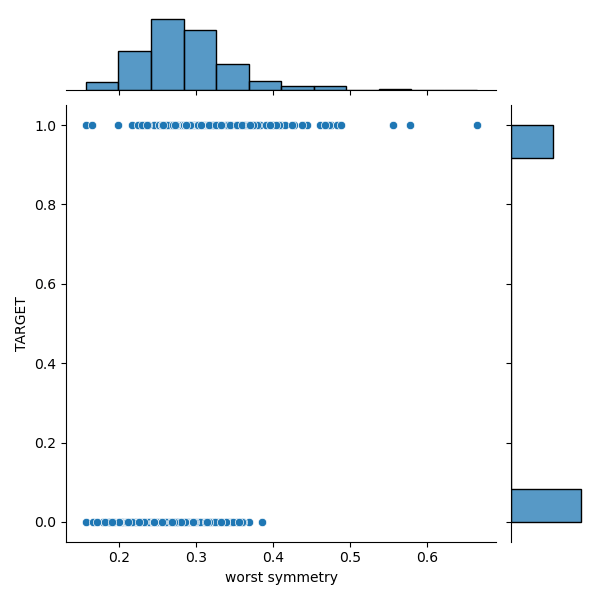
sns.jointplot(yX3[‘worst smoothness’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterworstsmoothness.png’)

sns.jointplot(yX3[‘worst texture’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterworsttexture.png’)
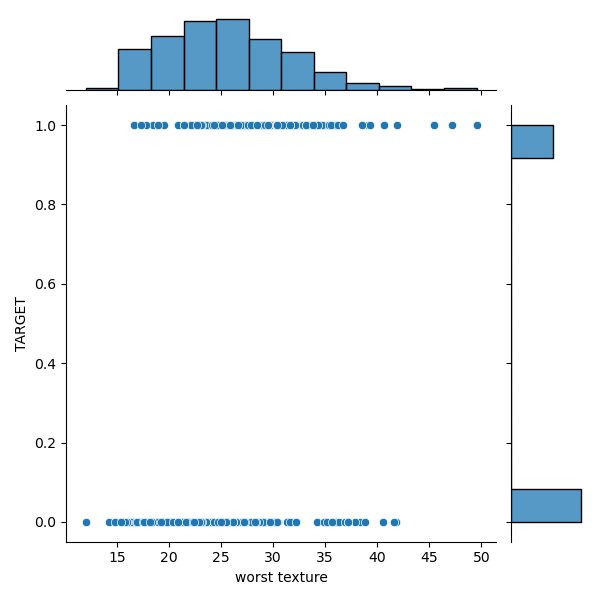
sns.jointplot(yX3[‘radius error’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterradiuserror.png’)

sns.jointplot(yX3[‘worst concave points’], yX3[‘TARGET’], kind=’scatter’, marginal_kws=dict(bins=12, rug=True))
plt.savefig(‘snsscatterworstconcavepoints.png’)

We can see a significant overlap between the M and B distributions in the target-feature domain.
Let’s plot the joint scatter matrix of target and model features
color_map = {0: ‘#0392cf’, 1: ‘#7bc043’} # 0 (negative class): blue, 1 (positive class): green
colors = yX3[‘TARGET’].map(lambda x: color_map.get(x))
pd.plotting.scatter_matrix(yX3, alpha=0.5, color=colors, figsize=(16,18), diagonal=’hist’, hist_kwds={‘bins’:12})
plt.suptitle(‘Scatter matrix of target and model features’)
#plt.show()
plt.savefig(‘scattermatrixjointfeatures.png’)

Let’s print the train data and the class distribution of TARGET for both train and test sets
print(“X_train.shape, y_train.shape”, X_train.shape, y_train.shape)
val_cnts = y_train.value_counts()
print(“Class distribution of positive and negative samples in the train set:”)
print(val_cnts)
print(“Percentage of positive class samples: %s” % “%2f%%” % (100 * val_cnts[1] / len(y_train)))
print(“—“)
print(“X_test.shape, y_test.shape”, X_test.shape, y_test.shape)
val_cnts = y_test.value_counts()
print(“Class distribution of positive and negative samples in the test set:”)
print(val_cnts)
print(“Percentage of positive class samples: %s” % “%2f%%” % (100 * val_cnts[1] / len(y_test)))
X_train.shape, y_train.shape (398, 9) (398,) Class distribution of positive and negative samples in the train set: 0 249 1 149 dtype: int64 Percentage of positive class samples: 37.437186% --- X_test.shape, y_test.shape (171, 9) (171,) Class distribution of positive and negative samples in the test set: 0 108 1 63 dtype: int64 Percentage of positive class samples: 36.842105%
Let’s define the HPO plotting function:
Plot 2d grid search heatmap
Parameters:
grid_search: instance of sklearn’s GridSearchCV
grid_params: dictionary of grid search parameters
x_param: name of x-axis parameter in grid_params
y_param: name of y-axis parameter in grid_params
is_verbose (optional): print results
Return:
grid_search.best_score_: best score found
grid_search.best_estimator_: best estimator found
def plot_2d_grid_search_heatmap(grid_search, grid_params, x_param, y_param, is_verbose=True):
grid_params_x = grid_params[x_param]
grid_params_y = grid_params[y_param]
df_results = pd.DataFrame(grid_search.cv_results_)
ar_scores = np.array(df_results.mean_test_score).reshape(len(grid_params_y), len(grid_params_x))
sns.heatmap(ar_scores, annot=True, fmt=’.3f’, xticklabels=grid_params_x, yticklabels=grid_params_y)
print()
plt.suptitle(‘Grid search heatmap’)
plt.xlabel(x_param)
plt.ylabel(y_param)
plt.savefig(“gridsearchheatmapgbm.png”)
if is_verbose:
print(“grid_search.best_score_:”)
print(grid_search.best_score_)
print()
print(“grid_search.best_estimator_:”)
print(grid_search.best_estimator_)
return grid_search.best_score_, grid_search.best_estimator_
Let’s run the LR classifier with GridSearchCV
grid_lr = {‘C’: [0.001, 0.01, 0.1, 1, 10, 100, 1000], ‘penalty’: [‘l1’, ‘l2’]}
clf_lr = LogisticRegression(class_weight=’balanced’, dual=False,
fit_intercept=True, intercept_scaling=1, max_iter=200,
n_jobs=1, random_state=0, tol=0.0001, verbose=0, warm_start=False)
gs_lr = GridSearchCV(clf_lr, grid_lr, return_train_score=True)
gs_lr.fit(X_train, y_train)
best_score_lr, clf_lr = plot_2d_grid_search_heatmap(gs_lr, grid_lr, ‘C’, ‘penalty’)
print(“intercept_:”)
print(clf_lr.intercept_ )
print()
print(“coef_:”)
print(clf_lr.coef_)
grid_search.best_score_:
0.9548417721518987
grid_search.best_estimator_:
LogisticRegression(C=1000, class_weight='balanced', max_iter=200, n_jobs=1,
random_state=0)
intercept_:
[-20.60103]
coef_:
[[ 78.14404612 15.64423206 0.25956452 -10.02229477 14.54515794
-13.02308961 -13.84545081 -47.97567754 -12.68950602]]

Let’s run the GB classifier with GridSearchCV
grid_gb = {‘min_samples_leaf’: [2, 4, 8, 16], ‘learning_rate’: [0.001, 0.01, 0.1]}
clf_gbm = GradientBoostingClassifier(criterion=’friedman_mse’,
init=None, loss=’deviance’, max_features=’sqrt’, max_leaf_nodes=None,
max_depth=3, min_impurity_decrease=0.0,
min_samples_split=2, min_weight_fraction_leaf=0.0, n_estimators=800,
random_state=0, subsample=1.0, verbose=0, warm_start=False)
gs_gb = GridSearchCV(clf_gbm, grid_gb, verbose=0, return_train_score=True)
gs_gb.fit(X_train, y_train)
best_score_gb, clf_gb = plot_2d_grid_search_heatmap(gs_gb, grid_gb, ‘min_samples_leaf’, ‘learning_rate’)
grid_search.best_score_:
0.9447468354430381
grid_search.best_estimator_:
GradientBoostingClassifier(learning_rate=0.01, max_features='sqrt',
min_samples_leaf=2, n_estimators=800,
random_state=0)

Let’s create the std train data frame
dfimp = pd.DataFrame(np.std(X_train, 0), columns=[‘std’])
and calculate the importance factors to rank them from high to low values of importance
Logistic regression:
dfimp[‘coef’] = clf_lr.coef_.T
dfimp[‘lr_sign_imp’] = (np.std(X_train, 0).ravel() * clf_lr.coef_).T
dfimp[‘lr_imp’] = np.abs(dfimp[‘lr_sign_imp’])
GBM:
dfimp[‘gb_imp’] = pd.Series(clf_gb.feature_importances_, X_train.columns)
Add a column for the correlation with target
dfimp[‘target_corr’] = s_X3_cols.drop(‘TARGET’)
and perform ranking
dfimp[‘lr_rank’] = dfimp[‘lr_imp’].rank(ascending=False)
dfimp[‘gb_rank’] = dfimp[‘gb_imp’].rank(ascending=False)
dfimp[‘target_corr_rank’] = dfimp[‘target_corr’].rank(ascending=False)
dfsort = dfimp.sort_values(‘target_corr_rank’, ascending=True)
print(dfsort.iloc[:20,:].to_string())
std coef lr_sign_imp lr_imp gb_imp target_corr lr_rank gb_rank target_corr_rank worst concave points 0.0656 78.1440 5.1267 5.1267 0.4893 0.8024 1.0 1.0 1.0 radius error 0.2936 15.6442 4.5931 4.5931 0.2137 0.5574 2.0 2.0 2.0 worst texture 6.1397 0.2596 1.5937 1.5937 0.0878 0.4347 3.0 3.0 3.0 worst smoothness 0.0230 -10.0223 -0.2301 0.2301 0.0400 0.4324 8.0 6.0 4.0 worst symmetry 0.0634 14.5452 0.9216 0.9216 0.0286 0.4255 4.0 7.0 5.0 concave points error 0.0059 -13.0231 -0.0762 0.0762 0.0567 0.3902 9.0 5.0 6.0 mean symmetry 0.0280 -13.8455 -0.3880 0.3880 0.0092 0.3361 6.0 9.0 7.0 compactness error 0.0168 -47.9757 -0.8066 0.8066 0.0141 0.2823 5.0 8.0 8.0 concavity error 0.0285 -12.6895 -0.3622 0.3622 0.0605 0.2322 7.0 4.0 9.0
Let’s look at the test data
cols_top = dfsort.index[:7]
print(“[y_test, X_test]”)
yX_test = pd.concat([y_test.iloc[-10:], X_test[cols_top].iloc[-10:,:]], axis=1)
yX_test = yX_test.rename(columns={0: ‘TARGET’})
print(yX_test.to_string())
[y_test, X_test]
TARGET worst concave points radius error worst texture worst smoothness worst symmetry concave points error mean symmetry
334 0 0.0398 0.1840 28.46 0.1222 0.2554 0.0065 0.1539
440 0 0.1555 0.2574 26.87 0.1391 0.2540 0.0177 0.1489
441 1 0.1739 0.5100 35.46 0.1436 0.2500 0.0174 0.1467
137 0 0.0848 0.1759 22.02 0.1190 0.2676 0.0086 0.1734
230 1 0.2543 0.2959 24.89 0.1703 0.3109 0.0138 0.2131
7 1 0.1556 0.5835 28.14 0.1654 0.3196 0.0145 0.2196
408 1 0.1974 0.4537 25.41 0.1482 0.3060 0.0148 0.1992
523 0 0.1284 0.3191 25.63 0.1425 0.2849 0.0135 0.1714
361 0 0.0561 0.2621 29.20 0.1140 0.2637 0.0064 0.1815
553 0 0.0256 0.3013 25.05 0.1103 0.2435 0.0128 0.1692
Let’s created the integrated LR-GBM HPO performance classification report
li_clfs = [‘clf_lr’, ‘clf_gb’]
dfp = pd.DataFrame(index=[‘TARGET’], data=[y_test]).T
for s_clf in li_clfs:
print(“MODEL: ” + s_clf)
print(“————–“)
clf = eval(s_clf)
y_pred = clf.predict(X_test).astype(int) # returns a class decision based on the value of the predicted probability
y_score = clf.predict_proba(X_test) # returns the value of the predicted probability
s_class = "%s_class" % s_clf
s_proba = "%s_proba" % s_clf
s_rank = "%s_rank" % s_clf
dfp[s_class] = y_pred
dfp[s_proba] = y_score[:,1]
dfp[s_rank] = dfp[s_proba].rank(ascending=1).astype(int)
# Print confusion matrix & classification report
# from sklearn.metrics import confusion_matrix
# cm = confusion_matrix(y_test, y_pred)
# Pandas 'crosstab' displays a better formatted confusion matrix than the one in sklearn
cm = pd.crosstab(y_test, y_pred, rownames=['Reality'], colnames=['Predicted'], margins=True)
print(cm)
print()
print("Classification report:")
print(classification_report(y_test, y_pred))
if s_clf == 'clf_lr':
y_score_lr = y_score.copy()
elif s_clf == 'clf_gb':
y_score_gb = y_score.copy()
else:
print('Error')
break
MODEL: clf_lr
--------------
Predicted 0 1 All
Reality
0 101 7 108
1 4 59 63
All 105 66 171
Classification report:
precision recall f1-score support
0 0.96 0.94 0.95 108
1 0.89 0.94 0.91 63
accuracy 0.94 171
macro avg 0.93 0.94 0.93 171
weighted avg 0.94 0.94 0.94 171
MODEL: clf_gb
--------------
Predicted 0 1 All
Reality
0 103 5 108
1 7 56 63
All 110 61 171
Classification report:
precision recall f1-score support
0 0.94 0.95 0.94 108
1 0.92 0.89 0.90 63
accuracy 0.93 171
macro avg 0.93 0.92 0.92 171
weighted avg 0.93 0.93 0.93 171
print(dfp.tail(10).to_string())
TARGET clf_lr_class clf_lr_proba clf_lr_rank clf_gb_class clf_gb_proba clf_gb_rank 334 0 0 0.0006 30 0 0.0079 58 440 0 0 0.3821 101 1 0.7771 113 441 1 1 0.9994 137 1 0.9928 147 137 0 0 0.0020 44 0 0.0032 6 230 1 1 0.9998 147 1 0.9350 119 7 1 1 0.9975 131 1 0.9949 157 408 1 1 0.9990 135 1 0.9924 144 523 0 0 0.4748 105 0 0.2996 103 361 0 0 0.0035 52 0 0.0045 29 553 0 0 0.0001 13 0 0.0041 21
Let’s summarize the statistics
RANK_BINS = 10
rank_cols = [‘i_bin’, ‘i_min_bin’, ‘i_max_bin’, ‘score_min’, ‘score_max’, ‘bin_cnt’, ‘pos_cnt’, ‘pos_rate’]
rank_cols2 = [‘i_bin’, ‘score_min’, ‘score_max’, ‘tnr’, ‘fpr’, ‘fnr’, ‘tpr’, ‘tn’, ‘fp’, ‘fn’, ‘tp’]
print(dfp.shape)
len_test = dfp.shape[0]
len_bin = int(len_test / RANK_BINS)
for s_clf in li_clfs:
i_min_bin = 0
i_max_bin = 0
dfr = pd.DataFrame(columns=rank_cols)
dfr2 = pd.DataFrame(columns=rank_cols2)
s_class = "%s_class" % s_clf
s_proba = "%s_proba" % s_clf
s_rank = "%s_rank" % s_clf
for i in range(RANK_BINS):
if i == RANK_BINS - 1:
i_max_bin = len_test
else:
i_max_bin += len_bin
# Range used for *each* score bin
df_rng = dfp[(dfp[s_rank] >= i_min_bin) & (dfp[s_rank] < i_max_bin)]
score_min = np.min(df_rng[s_proba])
score_max = np.max(df_rng[s_proba])
bin_cnt = df_rng.shape[0]
pos_cnt = len(df_rng[df_rng['TARGET'] == 1])
pos_rate = pos_cnt / bin_cnt
# Range used for *all* score bins up to i_max_bin
df_rng_0 = dfp[dfp[s_rank] < i_max_bin]
df_rng_1 = dfp[dfp[s_rank] >= i_max_bin]
# Positive targets (positive customers)
tp = len(df_rng_1[df_rng_1['TARGET'] == 1])
fn = len(df_rng_0[df_rng_0['TARGET'] == 1])
pos = tp + fn
tpr = tp / pos
fnr = 1 - tpr
# Negative targets
tn = len(df_rng_0[df_rng_0['TARGET'] == 0])
fp = len(df_rng_1[df_rng_1['TARGET'] == 0])
neg = tn + fp
fpr = fp / neg
tnr = 1 - fpr
# Build the dataframe for the summary stats per bin
row = [i, i_min_bin, i_max_bin, score_min, score_max, bin_cnt, pos_cnt, pos_rate]
dfr.loc[i] = row
# Build the dataframe for the ROC stats per bin
row2 = [i, score_min, score_max, tnr, fpr, fnr, tpr, tn, fp, fn, tp]
dfr2.loc[i] = row2
# Prep for next iteration
i_min_bin = i_max_bin
if s_clf == 'clf_lr':
dfr_lr = dfr.copy()
dfr2_lr = dfr2.copy()
elif s_clf == 'clf_gb':
dfr_gb = dfr.copy()
dfr2_gb = dfr2.copy()
else:
print('Error')
break
print(“——-“)
print(“Results of positive count stats per bin”)
print(“Logistic Regression stats:”)
print(dfr_lr.to_string())
print()
print(“GBM stats:”)
print(dfr_gb.to_string())
print()
print(“—“)
print(“Overall stats in the test set:”)
pos_cnt = len(dfp[dfp[‘TARGET’] == 1])
pos_rate = pos_cnt / len_test
print(“pos_cnt”, pos_cnt)
print(“total rows”, len_test)
print(“pos_rate”, pos_rate)
(171, 7) ------- Results of positive count stats per bin Logistic Regression stats: i_bin i_min_bin i_max_bin score_min score_max bin_cnt pos_cnt pos_rate 0 0.0 0.0 17.0 1.4302e-06 0.0002 16.0 0.0 0.0000 1 1.0 17.0 34.0 1.9229e-04 0.0007 17.0 0.0 0.0000 2 2.0 34.0 51.0 7.1944e-04 0.0033 17.0 0.0 0.0000 3 3.0 51.0 68.0 3.5092e-03 0.0103 17.0 0.0 0.0000 4 4.0 68.0 85.0 1.2065e-02 0.0481 17.0 1.0 0.0588 5 5.0 85.0 102.0 4.8669e-02 0.3821 17.0 3.0 0.1765 6 6.0 102.0 119.0 3.9533e-01 0.9029 17.0 6.0 0.3529 7 7.0 119.0 136.0 9.4799e-01 0.9990 17.0 17.0 1.0000 8 8.0 136.0 153.0 9.9925e-01 1.0000 17.0 17.0 1.0000 9 9.0 153.0 171.0 9.9999e-01 1.0000 18.0 18.0 1.0000 GBM stats: i_bin i_min_bin i_max_bin score_min score_max bin_cnt pos_cnt pos_rate 0 0.0 0.0 17.0 0.0023 0.0040 16.0 0.0 0.0000 1 1.0 17.0 34.0 0.0040 0.0047 17.0 0.0 0.0000 2 2.0 34.0 51.0 0.0047 0.0058 17.0 0.0 0.0000 3 3.0 51.0 68.0 0.0059 0.0101 17.0 0.0 0.0000 4 4.0 68.0 85.0 0.0109 0.0341 17.0 2.0 0.1176 5 5.0 85.0 102.0 0.0362 0.2315 17.0 4.0 0.2353 6 6.0 102.0 119.0 0.2659 0.9298 17.0 5.0 0.2941 7 7.0 119.0 136.0 0.9350 0.9880 17.0 16.0 0.9412 8 8.0 136.0 153.0 0.9895 0.9945 17.0 17.0 1.0000 9 9.0 153.0 171.0 0.9946 0.9964 18.0 18.0 1.0000 --- Overall stats in the test set: pos_cnt 63 total rows 171 pos_rate 0.3684210526315789
Let’s compare the percentiles for LR and GBM
s_titles = [‘Logistic Regression’, ‘GBM’]
plt.figure(figsize=(12, 4))
for i, s_clf in enumerate(li_clfs):
s_title = s_titles[i]
if s_clf == 'clf_lr':
dfr = dfr_lr
dfr2 = dfr2_lr
elif s_clf == 'clf_gb':
dfr = dfr_gb
dfr2 = dfr2_gb
else:
print('Error')
break
plt.subplot(1, 2, i+1)
plt.bar(dfr['i_bin'], dfr['pos_rate'])
plt.title('Percentiles for ' + s_title)
# Format charts
plt.grid()
plt.xlabel('Score bin (from low to high scoring bins)')
plt.ylabel('Ratio of positive classes per bin')
plt.legend(loc="lower right")
plt.xlim([0, RANK_BINS])
plt.ylim([0.0, 1])
#plt.show()
plt.savefig(“percentileslrgbm.png”)

It is clear that LR has the highest score and POS.
Let’s check the ROC curves
print(“Results of ROC stats per bin”)
print(“Logistic Regression stats:”)
print(dfr2_lr.to_string())
print()
print(“GBM stats:”)
print(dfr2_gb.to_string())
Results of ROC stats per bin Logistic Regression stats: i_bin score_min score_max tnr fpr fnr tpr tn fp fn tp 0 0.0 1.4302e-06 0.0002 0.1481 0.8519 0.0000 1.0000 16.0 92.0 0.0 63.0 1 1.0 1.9229e-04 0.0007 0.3056 0.6944 0.0000 1.0000 33.0 75.0 0.0 63.0 2 2.0 7.1944e-04 0.0033 0.4630 0.5370 0.0000 1.0000 50.0 58.0 0.0 63.0 3 3.0 3.5092e-03 0.0103 0.6204 0.3796 0.0000 1.0000 67.0 41.0 0.0 63.0 4 4.0 1.2065e-02 0.0481 0.7685 0.2315 0.0159 0.9841 83.0 25.0 1.0 62.0 5 5.0 4.8669e-02 0.3821 0.8981 0.1019 0.0635 0.9365 97.0 11.0 4.0 59.0 6 6.0 3.9533e-01 0.9029 1.0000 0.0000 0.1587 0.8413 108.0 0.0 10.0 53.0 7 7.0 9.4799e-01 0.9990 1.0000 0.0000 0.4286 0.5714 108.0 0.0 27.0 36.0 8 8.0 9.9925e-01 1.0000 1.0000 0.0000 0.6984 0.3016 108.0 0.0 44.0 19.0 9 9.0 9.9999e-01 1.0000 1.0000 0.0000 0.9841 0.0159 108.0 0.0 62.0 1.0
GBM stats: i_bin score_min score_max tnr fpr fnr tpr tn fp fn tp 0 0.0 0.0023 0.0040 0.1481 0.8519 0.0000 1.0000 16.0 92.0 0.0 63.0 1 1.0 0.0040 0.0047 0.3056 0.6944 0.0000 1.0000 33.0 75.0 0.0 63.0 2 2.0 0.0047 0.0058 0.4630 0.5370 0.0000 1.0000 50.0 58.0 0.0 63.0 3 3.0 0.0059 0.0101 0.6204 0.3796 0.0000 1.0000 67.0 41.0 0.0 63.0 4 4.0 0.0109 0.0341 0.7593 0.2407 0.0317 0.9683 82.0 26.0 2.0 61.0 5 5.0 0.0362 0.2315 0.8796 0.1204 0.0952 0.9048 95.0 13.0 6.0 57.0 6 6.0 0.2659 0.9298 0.9907 0.0093 0.1746 0.8254 107.0 1.0 11.0 52.0 7 7.0 0.9350 0.9880 1.0000 0.0000 0.4286 0.5714 108.0 0.0 27.0 36.0 8 8.0 0.9895 0.9945 1.0000 0.0000 0.6984 0.3016 108.0 0.0 44.0 19.0 9 9.0 0.9946 0.9964 1.0000 0.0000 0.9841 0.0159 108.0 0.0 62.0 1.0
Let’s define and call the plot function
def plot_roc_and_precision_recall(y_true, y_score):
# Get ROC curve FPR and TPR from true labels vs score values
fpr, tpr, _ = roc_curve(y_true, y_score)
# Calculate ROC Area Under the Curve (AUC) from FPR and TPR data points
roc_auc = auc(fpr, tpr)
# Calculate precision and recall from true labels vs score values
precision, recall, _ = precision_recall_curve(y_true, y_score)
plt.figure(figsize=(8, 3))
plt.subplot(1,2,1)
lw = 2
plt.plot(fpr, tpr, color=’darkorange’, lw=lw, label=’ROC curve (AUC = %0.4f)’ % roc_auc)
plt.plot([0, 1], [0, 1], color=’navy’, lw=lw, linestyle=’–‘)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel(‘False Positive Rate’)
plt.ylabel(‘True Positive Rate’)
plt.title(‘ROC Curve’)
plt.legend(loc=”lower right”)
plt.grid(True)
plt.subplot(1,2,2)
plt.step(recall, precision, color=’orange’, where=’post’)
# plt.fill_between(recall, precision, step=’post’, alpha=0.5, color=’orange’)
plt.xlabel(‘Recall’)
plt.ylabel(‘Precision’)
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title(‘Precision Recall Curve’)
plt.grid(True)
left = 0.125 # the left side of the subplots of the figure
right = 0.9 # the right side of the subplots of the figure
bottom = 0.1 # the bottom part of the subplots of the figure
top = 0.9 # the top part of the subplots of the figure
wspace = 0.5 # the amount of width reserved for blank space between subplots
hspace = 0.2 # the amount of height reserved for white space between subplots
plt.subplots_adjust(left, bottom, right, top, wspace, hspace)
plt.show()
for s_clf in li_clfs:
print(“MODEL: ” + s_clf)
print(“————–“)
clf = eval(s_clf)
y_pred = clf.predict(X_test).astype(int)
y_score = clf.predict_proba(X_test)
plot_roc_and_precision_recall(y_test, y_score[:,1]) # provide the column for the scores belonging only to the positive class
print()
MODEL: clf_lr --------------

MODEL: clf_gb --------------
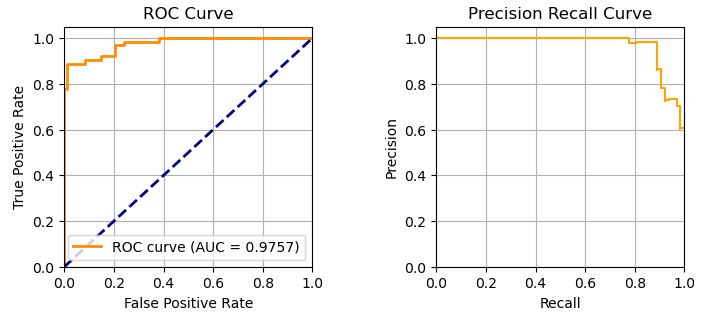
We can see that AUC(LR)>AUC(GBM).
Summary
- Train/test data splitting ratio 70:30%
- Feature engineering yields 9 dominant features
worst concave points 0.8024
radius error 0.5574
worst texture 0.4347
worst smoothness 0.4324
worst symmetry 0.4255
concave points error 0.3902
mean symmetry 0.3361
compactness error 0.2823
concavity error 0.2322
- LR Classification report:
- precision recall f1-score support
- 0 0.96 0.94 0.95 108
- 1 0.89 0.94 0.91 63
- accuracy 0.94 171
- GB Classification report:
- precision recall f1-score support
- 0 0.94 0.95 0.94 108
- 1 0.92 0.89 0.90 63
- accuracy 0.93 171
- Percentiles for LR vs GBM: LR has the highest score.
- AUC(LR)=0.987 > AUC(GBM)=0.975
- Results are consistent with the previous ML R&D study.
Read More
Colab Research Example using scikit-learn: Breast cancer prediction
A Comparative Analysis of Breast Cancer ML/AI Binary Classifications
Supervised ML/AI Breast Cancer Diagnostics (BCD) – The Power of HealthTech
How to find the importance of the features for a logistic regression model?
Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.


Leave a comment