Featured Photo by Myriam Zilles on Unsplash
In the last decade, the impact of Type-2 Diabetes (T2D) has increased to a great extent especially in developing countries. Therefore, early diagnosis and classification of T2D has become an active area of healthtech. Numerous data-driven techniques are available to control T2D. This work presents a comparative use-case study of several exploratory data analysis (EDA) and ML/AI algorithms which process T2D data effectively. The existing Python workflows are analyzed in a comprehensive manner to identify their pros and cons. Their performance assessment is carried out to determine the best approach.
Contents:
Background
- T2D is a common condition that causes the level of sugar (glucose) in the blood to become too high. T2D is responsible for very considerable morbidity, mortality. Furthermore, it is a substantial financial drain health systems and societies.
- Early diagnosis is important to reduce the incidence and mortality rate of diabetes.
- Screening tools have been developed to identify individuals at high risk of developing T2D.
- Most screening tests for T2D in use today were developed using electronically collected data from Electronic Health Record (EHR).
- Traditional screening approaches are based on standard regression techniques.
- It is important to investigate whether using advanced data-based approaches can yield superior results to the currently employed methods.
- The increasing volume of EHR opened the opportunity to develop more accurate AI-driven prediction models that can be continuously updated using available ML algorithms.
State-of-the-Art
Over the last years, various ML/AI techniques (DNN, SVM, KNN, DT, GBT, GBM, RF, LR, etc.) have been used to predict T2D and its complications. However, researchers and developers still face two main challenges when building T2D predictive models:
- There is considerable heterogeneity in previous studies regarding algorithms used, making it challenging to identify the optimal one.
- There is a lack of transparency about the features used in the optimized models, which reduces their interpretability.
- This systematic analysis aimed at providing answers to the above challenges.
- The study followed the earlier review primarily, enriched with the most recent case studies (cf. References and Explore More).
Workflow
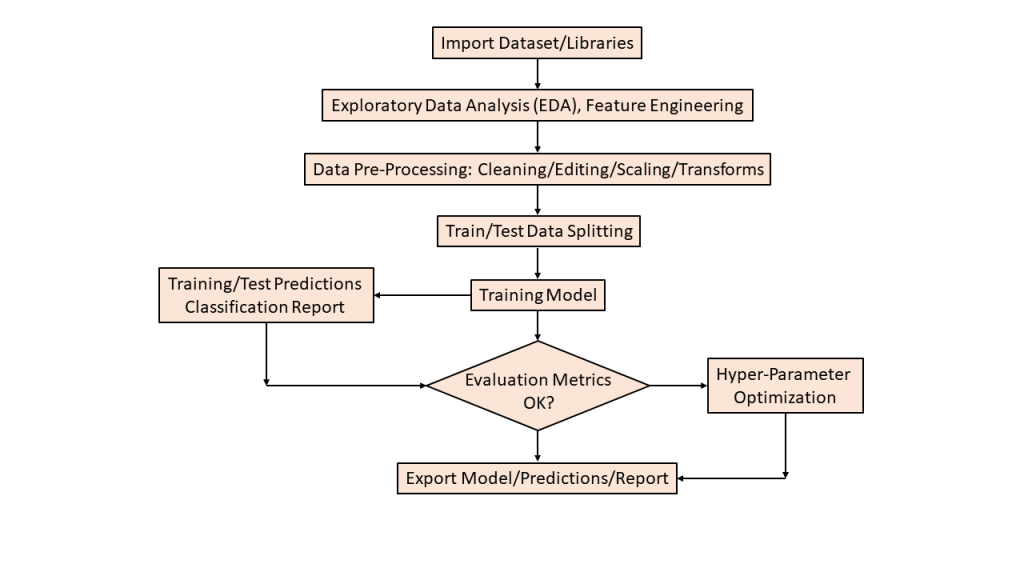
Data
Conventionally, T2D ML/AI studies use the Kaggle PIMA Indian Diabetes (PID) dataset taken from the National Institute of Diabetes and Kidney Diseases center, see UC Irvine Machine Learning Repository. The objective of the dataset is to diagnostically predict whether or not a patient has diabetes, based on certain diagnostic measurements included in the dataset. Several constraints were placed on the selection of these instances from a larger database. In particular, all patients here are females at least 21 years old of Pima Indian heritage.
The AWS Approach
Let’s Import libraries
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns # for data visualization
import matplotlib.pyplot as plt # to plot charts
from collections import Counter
import os
from sklearn.preprocessing import QuantileTransformer
from sklearn.metrics import confusion_matrix, accuracy_score, precision_score
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier, VotingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV, cross_val_score, StratifiedKFold, learning_curve, train_test_split
Let’s import the input dataset
df = pd.read_csv(“diabetes.csv”)
and get familier with dataset structure
df.info()
0 Pregnancies 768 non-null int64 1 Glucose 768 non-null int64 2 BloodPressure 768 non-null int64 3 SkinThickness 768 non-null int64 4 Insulin 768 non-null int64 5 BMI 768 non-null float64 6 DiabetesPedigreeFunction 768 non-null float64 7 Age 768 non-null int64 8 Outcome 768 non-null int64 dtypes: float64(2), int64(7)
Let’s look at top 5 rows
df.head()
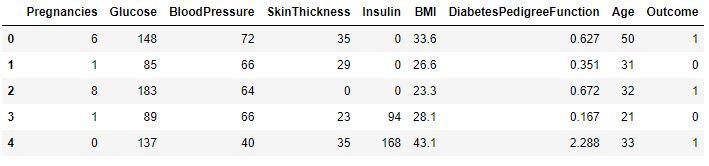
Our target variable is Outcome.
Let’s explore missing values if any
df.isnull().sum()
Pregnancies 0 Glucose 0 BloodPressure 0 SkinThickness 0 Insulin 0 BMI 0 DiabetesPedigreeFunction 0 Age 0 Outcome 0 dtype: int64
There are no missing values.
Let’s review the dataset descriptive statistics
df.describe()

and look at the correlation matrix
plt.figure(figsize=(13,10))
sns.heatmap(df.corr(),annot=True, fmt = “.2f”, cmap = “coolwarm”)

Let’s explore Pregnancies vs Outcome
plt.figure(figsize=(13,6))
g = sns.kdeplot(df[“Pregnancies”][df[“Outcome”] == 1],
color=”Red”, shade = True)
g = sns.kdeplot(df[“Pregnancies”][df[“Outcome”] == 0],
ax =g, color=”Green”, shade= True)
g.set_xlabel(“Pregnancies”)
g.set_ylabel(“Frequency”)
g.legend([“Positive”,”Negative”])
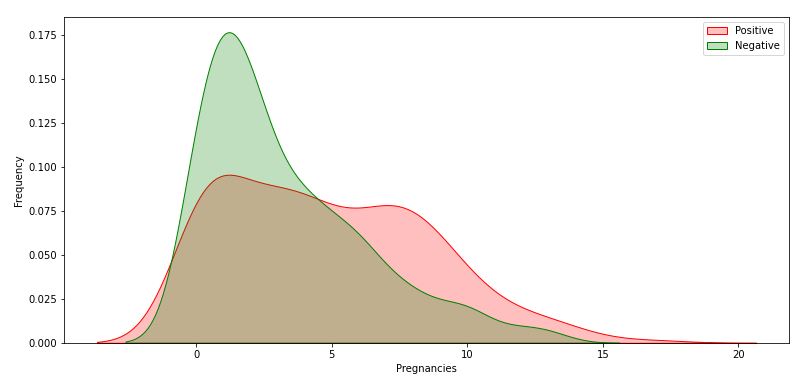

Let’s explore Glucose vs Outcome
plt.figure(figsize=(10,6))
sns.violinplot(data=df, x=”Outcome”, y=”Glucose”,
split=True, inner=”quart”, linewidth=1)

Let’s explore Glucose vs Outcome empirical distributions
plt.figure(figsize=(13,6))
g = sns.kdeplot(df[“Glucose”][df[“Outcome”] == 1], color=”Red”, shade = True)
g = sns.kdeplot(df[“Glucose”][df[“Outcome”] == 0], ax =g, color=”Green”, shade= True)
g.set_xlabel(“Glucose”)
g.set_ylabel(“Frequency”)
g.legend([“Positive”,”Negative”])

Let’s combine Glucose vs BMI vs Age
plt.figure(figsize=(20,10))
sns.scatterplot(data=df, x=”Glucose”, y=”BMI”, hue=”Age”, size=”Age”)

Let’s detect outliers
def detect_outliers(df,n,features):
outlier_indices = []
“””
Detect outliers from given list of features. It returns a list of the indices
according to the observations containing more than n outliers according
to the Tukey method
“””
# iterate over features(columns)
for col in features:
Q1 = np.percentile(df[col], 25)
Q3 = np.percentile(df[col],75)
IQR = Q3 – Q1
# outlier step
outlier_step = 1.5 * IQR
# Determine a list of indices of outliers for feature col
outlier_list_col = df[(df[col] < Q1 - outlier_step) | (df[col] > Q3 + outlier_step )].index
# append the found outlier indices for col to the list of outlier indices
outlier_indices.extend(outlier_list_col)
# select observations containing more than 2 outliers
outlier_indices = Counter(outlier_indices)
multiple_outliers = list( k for k, v in outlier_indices.items() if v > n )
return multiple_outliers
Let’s group outliers to drop using the above function
outliers_to_drop = detect_outliers(df, 2 ,[“Pregnancies”, ‘Glucose’, ‘BloodPressure’, ‘BMI’, ‘DiabetesPedigreeFunction’, ‘SkinThickness’, ‘Insulin’, ‘Age’])
We are ready to drop those outliers
df.drop(df.loc[outliers_to_drop].index, inplace=True)
Let’s apply the QuantileTransformer to change the probability distribution of numeric variables to improve the performance of predictive models
q = QuantileTransformer()
X = q.fit_transform(df)
transformedDF = q.transform(X)
transformedDF = pd.DataFrame(X)
transformedDF.columns =[‘Pregnancies’, ‘Glucose’, ‘BloodPressure’, ‘SkinThickness’, ‘Insulin’, ‘BMI’, ‘DiabetesPedigreeFunction’, ‘Age’, ‘Outcome’]
Let’s look at top 5 rows of the transformed dataset
transformedDF.head()

Let’s separate features and labels
features = df.drop([“Outcome”], axis=1)
labels = df[“Outcome”]
Let’s perform training/test data split with test_size=0.30
x_train, x_test, y_train, y_test = train_test_split(features, labels, test_size=0.30, random_state=7)
Let’s evaluate ML models using the function
def evaluate_model(models):
“””
Takes a list of models and returns chart of cross validation scores using mean accuracy
“””
# Cross validate model with Kfold stratified cross val
kfold = StratifiedKFold(n_splits = 10)
result = []
for model in models :
result.append(cross_val_score(estimator = model, X = x_train, y = y_train, scoring = "accuracy", cv = kfold, n_jobs=4))
cv_means = []
cv_std = []
for cv_result in result:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
result_df = pd.DataFrame({
"CrossValMeans":cv_means,
"CrossValerrors": cv_std,
"Models":[
"LogisticRegression",
"DecisionTreeClassifier",
"AdaBoostClassifier",
"SVC",
"RandomForestClassifier",
"GradientBoostingClassifier",
"KNeighborsClassifier"
]
})
# Generate chart
bar = sns.barplot(x = "CrossValMeans", y = "Models", data = result_df, orient = "h")
bar.set_xlabel("Mean Accuracy")
bar.set_title("Cross validation scores")
return result_df
Let’s perform modeling to compare different algorithms
random_state = 30
models = [
LogisticRegression(random_state = random_state, solver=’liblinear’),
DecisionTreeClassifier(random_state = random_state),
AdaBoostClassifier(DecisionTreeClassifier(random_state = random_state), random_state = random_state, learning_rate = 0.2),
SVC(random_state = random_state),
RandomForestClassifier(random_state = random_state),
GradientBoostingClassifier(random_state = random_state),
KNeighborsClassifier(),
]
evaluate_model(models)
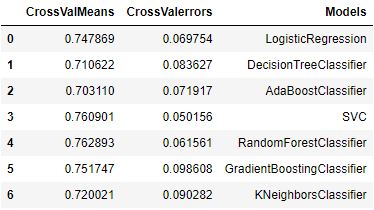

Let’s implement hyper-parameter optimization (HPO) via GridSearchCV and print out the classification report
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
def analyze_grid_result(grid_result):
”’
Analysis of GridCV result and predicting with test dataset
Show classification report at last
”’
# Best parameters and accuracy
print(“Tuned hyperparameters: (best parameters) “, grid_result.best_params_)
print(“Accuracy :”, grid_result.best_score_)
means = grid_result.cv_results_["mean_test_score"]
stds = grid_result.cv_results_["std_test_score"]
for mean, std, params in zip(means, stds, grid_result.cv_results_["params"]):
print("%0.3f (+/-%0.03f) for %r" % (mean, std * 2, params))
print()
print("Detailed classification report:")
y_true, y_pred = y_test, grid_result.predict(x_test)
print(classification_report(y_true, y_pred))
print()
Let’s define models and parameters for LogisticRegression
model = LogisticRegression(solver=’liblinear’)
solvers = [‘newton-cg’, ‘liblinear’]
penalty = [‘l2’]
c_values = [100, 10, 1.0, 0.1, 0.01]
Let’s consider GridSearchCV HPO
grid = dict(solver = solvers, penalty = penalty, C = c_values)
cv = StratifiedKFold(n_splits = 50, random_state = 1, shuffle = True)
grid_search = GridSearchCV(estimator = model, param_grid = grid, cv = cv, scoring = ‘accuracy’, error_score = 0)
logi_result = grid_search.fit(x_train, y_train)
Let’s print out the Logistic Regression HPO Result
analyze_grid_result(logi_result)
Tuned hyperparameters: (best parameters) {'C': 10, 'penalty': 'l2', 'solver': 'liblinear'}
Accuracy : 0.7883636363636363
0.788 (+/-0.260) for {'C': 100, 'penalty': 'l2', 'solver': 'newton-cg'}
0.788 (+/-0.260) for {'C': 100, 'penalty': 'l2', 'solver': 'liblinear'}
0.788 (+/-0.260) for {'C': 10, 'penalty': 'l2', 'solver': 'newton-cg'}
0.788 (+/-0.250) for {'C': 10, 'penalty': 'l2', 'solver': 'liblinear'}
0.785 (+/-0.253) for {'C': 1.0, 'penalty': 'l2', 'solver': 'newton-cg'}
0.743 (+/-0.267) for {'C': 1.0, 'penalty': 'l2', 'solver': 'liblinear'}
0.773 (+/-0.238) for {'C': 0.1, 'penalty': 'l2', 'solver': 'newton-cg'}
0.705 (+/-0.281) for {'C': 0.1, 'penalty': 'l2', 'solver': 'liblinear'}
0.773 (+/-0.232) for {'C': 0.01, 'penalty': 'l2', 'solver': 'newton-cg'}
0.696 (+/-0.264) for {'C': 0.01, 'penalty': 'l2', 'solver': 'liblinear'}
Detailed classification report:
precision recall f1-score support
0 0.78 0.84 0.81 147
1 0.68 0.58 0.62 83
accuracy 0.75 230
macro avg 0.73 0.71 0.72 230
weighted avg 0.74 0.75 0.74 230
Let’s look at the GridSearchCV
model = SVC()
tuned_parameters = [
{“kernel”: [“rbf”], “gamma”: [1e-3, 1e-4], “C”: [1, 10, 100, 1000]},
{“kernel”: [“linear”], “C”: [1, 10, 100, 1000]},
]
cv = StratifiedKFold(n_splits = 2, random_state = 1, shuffle = True)
grid_search = GridSearchCV(estimator = model, param_grid = tuned_parameters, cv = cv, scoring = ‘accuracy’, error_score = 0)
scv_result = grid_search.fit(x_train, y_train)
analyze_grid_result(scv_result)
Tuned hyperparameters: (best parameters) {'C': 10, 'kernel': 'linear'}
Accuracy : 0.7775797976410084
0.712 (+/-0.061) for {'C': 1, 'gamma': 0.001, 'kernel': 'rbf'}
0.735 (+/-0.053) for {'C': 1, 'gamma': 0.0001, 'kernel': 'rbf'}
0.677 (+/-0.035) for {'C': 10, 'gamma': 0.001, 'kernel': 'rbf'}
0.716 (+/-0.016) for {'C': 10, 'gamma': 0.0001, 'kernel': 'rbf'}
0.658 (+/-0.020) for {'C': 100, 'gamma': 0.001, 'kernel': 'rbf'}
0.707 (+/-0.042) for {'C': 100, 'gamma': 0.0001, 'kernel': 'rbf'}
0.656 (+/-0.001) for {'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
0.667 (+/-0.046) for {'C': 1000, 'gamma': 0.0001, 'kernel': 'rbf'}
0.770 (+/-0.025) for {'C': 1, 'kernel': 'linear'}
0.778 (+/-0.010) for {'C': 10, 'kernel': 'linear'}
0.778 (+/-0.005) for {'C': 100, 'kernel': 'linear'}
0.759 (+/-0.033) for {'C': 1000, 'kernel': 'linear'}
Detailed classification report:
precision recall f1-score support
0 0.78 0.84 0.81 147
1 0.67 0.57 0.61 83
accuracy 0.74 230
macro avg 0.72 0.70 0.71 230
weighted avg 0.74 0.74 0.74 230
Let’s look at the RandomForestClassifier HPO
model = RandomForestClassifier(random_state=42)
tuned_parameters = {
‘n_estimators’: [200, 500],
‘max_features’: [‘auto’, ‘sqrt’, ‘log2’],
‘max_depth’ : [4,5,6,7,8],
‘criterion’ :[‘gini’, ‘entropy’]
}
cv = StratifiedKFold(n_splits = 2, random_state = 1, shuffle = True)
grid_search = GridSearchCV(estimator = model, param_grid = tuned_parameters, cv = cv, scoring = ‘accuracy’, error_score = 0)
grid_result = grid_search.fit(x_train, y_train)
analyze_grid_result(grid_result)
Tuned hyperparameters: (best parameters) {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'log2', 'n_estimators': 200}
Accuracy : 0.7663648051875454
0.759 (+/-0.025) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'auto', 'n_estimators': 200}
0.761 (+/-0.029) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'auto', 'n_estimators': 500}
0.759 (+/-0.025) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'sqrt', 'n_estimators': 200}
0.761 (+/-0.029) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'sqrt', 'n_estimators': 500}
0.751 (+/-0.018) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'log2', 'n_estimators': 200}
0.757 (+/-0.022) for {'criterion': 'gini', 'max_depth': 4, 'max_features': 'log2', 'n_estimators': 500}
0.751 (+/-0.025) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'auto', 'n_estimators': 200}
0.759 (+/-0.025) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'auto', 'n_estimators': 500}
0.751 (+/-0.025) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'sqrt', 'n_estimators': 200}
0.759 (+/-0.025) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'sqrt', 'n_estimators': 500}
0.751 (+/-0.025) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'log2', 'n_estimators': 200}
0.753 (+/-0.014) for {'criterion': 'gini', 'max_depth': 5, 'max_features': 'log2', 'n_estimators': 500}
0.763 (+/-0.033) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'auto', 'n_estimators': 200}
0.757 (+/-0.029) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'auto', 'n_estimators': 500}
0.763 (+/-0.033) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'sqrt', 'n_estimators': 200}
0.757 (+/-0.029) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'sqrt', 'n_estimators': 500}
0.759 (+/-0.025) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'log2', 'n_estimators': 200}
0.753 (+/-0.029) for {'criterion': 'gini', 'max_depth': 6, 'max_features': 'log2', 'n_estimators': 500}
0.755 (+/-0.033) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'auto', 'n_estimators': 200}
0.751 (+/-0.025) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'auto', 'n_estimators': 500}
0.755 (+/-0.033) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'sqrt', 'n_estimators': 200}
0.751 (+/-0.025) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'sqrt', 'n_estimators': 500}
0.748 (+/-0.018) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'log2', 'n_estimators': 200}
0.746 (+/-0.007) for {'criterion': 'gini', 'max_depth': 7, 'max_features': 'log2', 'n_estimators': 500}
0.751 (+/-0.033) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'auto', 'n_estimators': 200}
0.751 (+/-0.040) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'auto', 'n_estimators': 500}
0.751 (+/-0.033) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'sqrt', 'n_estimators': 200}
0.751 (+/-0.040) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'sqrt', 'n_estimators': 500}
0.750 (+/-0.021) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'log2', 'n_estimators': 200}
0.757 (+/-0.044) for {'criterion': 'gini', 'max_depth': 8, 'max_features': 'log2', 'n_estimators': 500}
0.763 (+/-0.040) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'auto', 'n_estimators': 200}
0.759 (+/-0.040) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'auto', 'n_estimators': 500}
0.763 (+/-0.040) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'sqrt', 'n_estimators': 200}
0.759 (+/-0.040) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'sqrt', 'n_estimators': 500}
0.757 (+/-0.029) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'log2', 'n_estimators': 200}
0.763 (+/-0.010) for {'criterion': 'entropy', 'max_depth': 4, 'max_features': 'log2', 'n_estimators': 500}
0.757 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'auto', 'n_estimators': 200}
0.757 (+/-0.036) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'auto', 'n_estimators': 500}
0.757 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'sqrt', 'n_estimators': 200}
0.757 (+/-0.036) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'sqrt', 'n_estimators': 500}
0.766 (+/-0.010) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'log2', 'n_estimators': 200}
0.764 (+/-0.014) for {'criterion': 'entropy', 'max_depth': 5, 'max_features': 'log2', 'n_estimators': 500}
0.765 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'auto', 'n_estimators': 200}
0.763 (+/-0.033) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'auto', 'n_estimators': 500}
0.765 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'sqrt', 'n_estimators': 200}
0.763 (+/-0.033) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'sqrt', 'n_estimators': 500}
0.764 (+/-0.007) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'log2', 'n_estimators': 200}
0.759 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 6, 'max_features': 'log2', 'n_estimators': 500}
0.761 (+/-0.014) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'auto', 'n_estimators': 200}
0.757 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'auto', 'n_estimators': 500}
0.761 (+/-0.014) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'sqrt', 'n_estimators': 200}
0.757 (+/-0.022) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'sqrt', 'n_estimators': 500}
0.757 (+/-0.029) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'log2', 'n_estimators': 200}
0.753 (+/-0.014) for {'criterion': 'entropy', 'max_depth': 7, 'max_features': 'log2', 'n_estimators': 500}
0.755 (+/-0.010) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'auto', 'n_estimators': 200}
0.755 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'auto', 'n_estimators': 500}
0.755 (+/-0.010) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'sqrt', 'n_estimators': 200}
0.755 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'sqrt', 'n_estimators': 500}
0.748 (+/-0.040) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'log2', 'n_estimators': 200}
0.763 (+/-0.033) for {'criterion': 'entropy', 'max_depth': 8, 'max_features': 'log2', 'n_estimators': 500}
Detailed classification report:
precision recall f1-score support
0 0.78 0.83 0.80 147
1 0.66 0.58 0.62 83
accuracy 0.74 230
macro avg 0.72 0.70 0.71 230
weighted avg 0.73 0.74 0.74 230
Let’s test logi_result predictions
y_pred = logi_result.predict(x_test)
print(classification_report(y_test, y_pred))
precision recall f1-score support
0 0.78 0.84 0.81 147
1 0.68 0.58 0.62 83
accuracy 0.75 230
macro avg 0.73 0.71 0.72 230
weighted avg 0.74 0.75 0.74 230
Let’s add our predictions to the test dataset
x_test[‘pred’] = y_pred
print(x_test)
Pregnancies Glucose BloodPressure SkinThickness Insulin BMI
236 7 181.0 84.0 21 192.0 35.9
715 7 187.0 50.0 33 392.0 33.9
766 1 126.0 60.0 23 30.5 30.1
499 6 154.0 74.0 32 193.0 29.3
61 8 133.0 72.0 23 30.5 32.9
.. ... ... ... ... ... ...
189 5 139.0 80.0 35 160.0 31.6
351 4 137.0 84.0 23 30.5 31.2
120 0 162.0 76.0 56 100.0 53.2
108 3 83.0 58.0 31 18.0 34.3
637 2 94.0 76.0 18 66.0 31.6
DiabetesPedigreeFunction Age pred
236 0.586 51 1
715 0.826 34 1
766 0.349 47 0
499 0.839 39 1
61 0.270 39 1
.. ... ... ...
189 0.361 25 0
351 0.252 30 0
120 0.759 25 1
108 0.336 25 0
637 0.649 23 0
[230 rows x 9 columns]
Let’s plot histograms
plt.hist(y_test)
plt.hist(y_pred)

Conclusions
- Input dataset: the Kaggle PIMA Indian Diabetes (PID)
- Data Pre-Processing: drop outliers and apply QuantileTransformer
- Correlation matrix feature ranking: 1 – Glucose, 2 – BMI, 3 – Age.
- 0.7 < Cross-Validation Mean Errors (Models) < 0.8
- Binary Classification Models: LR, KNN, GB, RF, DT, AdaBoost, and SVC.
- SVC HPO via GridSearchCV: Accuracy : 0.77
precision recall f1-score support
0 0.78 0.84 0.81 147
1 0.67 0.57 0.61 83
- RandomForestClassifier HPO via GridSearchCV: Accuracy : 0.74
precision recall f1-score support
0 0.78 0.83 0.80 147
1 0.66 0.58 0.62 83
References
Machine learning and deep learning predictive models for type 2 diabetes: a systematic review
Early detection of type 2 diabetes mellitus using machine learning-based prediction models
Explore More
HealthTech ML/AI Q3 ’22 Round-Up
The Application of ML/AI in Diabetes
Diabetes Prediction using ML/AI in Python
Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.


Leave a comment