This blog was inspired by the recent tests with Machine Learning (ML) hyper-parameter optimization (HPO) and cross-validation scores for enhanced prediction of Type-2 diabetes (T2D).
The basic workflow includes RandomizedSearchCV HPO and accuracy metrics:
- Import libraries and data
- Data Pre-Processing, Cleaning, Scaling, Resampling and Transformations
- Exploratory Data Analysis (EDA)
- Feature Engineering (FE)
- Model Building – GaussianNB
- RandomForestClassifier with RandomizedSearchCV
- DecisionTreeClassifier
- Compute confusion matrix, accuracy score
The advanced workflow includes EDA, FE, HPO, and complete X-validation:
- Import libraries and data
- Feature Engineering and Exploratory Data Analysis
- Replace missing values by median values
- Creating metrics for evaluations
- Model building with GBC and HPO with GridSearchCV
- X-validation and full classification report
- Export the best model with pickle
Both workflows compare several ML algorithms, SMOTE oversampling, scaling and various performance metrics.
Basic Workflow
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
Importing essential libraries
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
import seaborn as sns
and importing the input dataset
df = pd.read_csv(‘diabetes.csv’)
df.head()
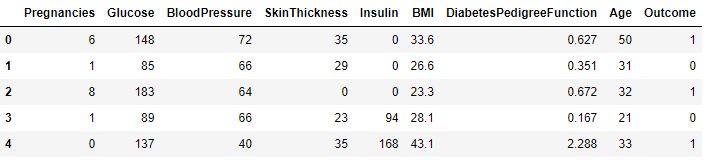
We rename DiabetesPedigreeFunction as DPF
df = df.rename(columns={‘DiabetesPedigreeFunction’:’DPF’})
while replacing the 0 values from [‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’] by NaN
df_copy = df.copy(deep=True)
df_copy[[‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’]] = df_copy[[‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’]].replace(0,np.nan)
Print No. of rows and columns
print(f’Total Rows {df_copy.shape[0]}’)
print(f’Total Rows {df_copy.shape[1]}’)
Total Rows 768 Total Rows 9
We collect all columns having missing values and at what percentage
columns_with_na=[features for features in df_copy.columns if df_copy[features].isnull().sum()>0]
for feature in columns_with_na:
print(feature, np.round(df_copy[feature].isnull().mean(), 4), ‘ % missing values’)
Glucose 0.0065 % missing values BloodPressure 0.0456 % missing values SkinThickness 0.2956 % missing values Insulin 0.487 % missing values BMI 0.0143 % missing values
Let’s plot the counts of missing values per Outcome
for feature in columns_with_na:
data = df_copy.copy()
data[feature] = np.where(data[feature].isnull(), 1, 0)
data.groupby(feature)[‘Outcome’].value_counts().plot.bar()
plt.title(feature)
plt.show()
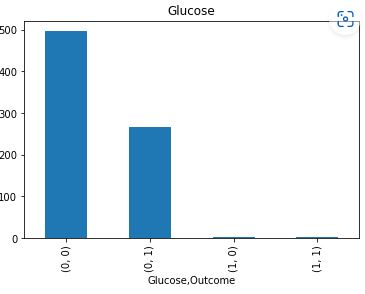
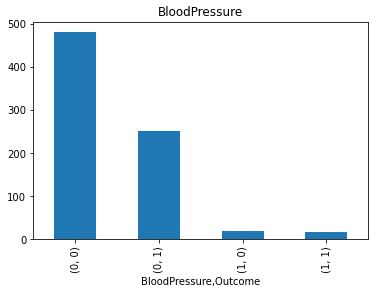
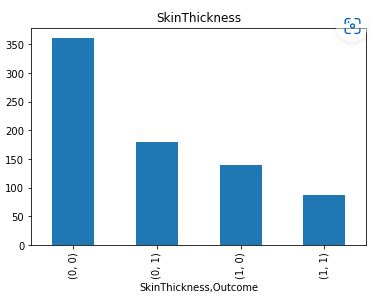
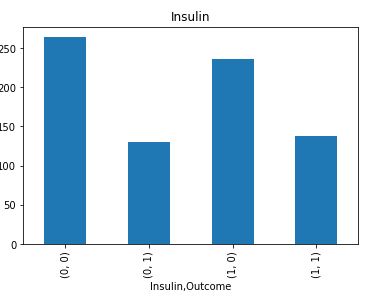
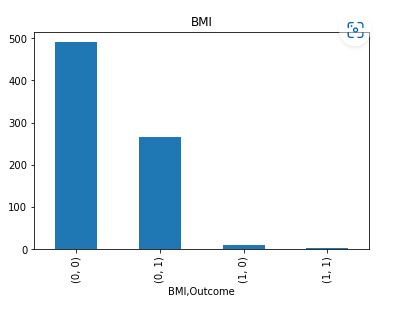
It is clear that Insulin and Skin Thickness have the largest number of missing values for both values of Outcome.
Get all the columns having numerical values
num_data = [features for features in df_copy.columns if df_copy[features].dtypes != ‘O’]
print(f’Number of numerical columns is {len(num_data)}’)
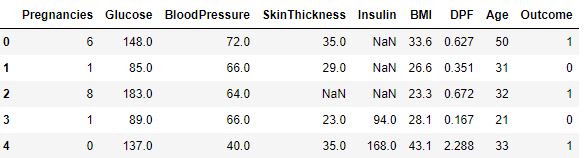
df_copy[num_data].head()
Number of numerical columns is 9
Check if any numerical columns are discrete
discrete_columns = [feature for feature in df_copy.columns if len(df_copy[feature].unique()) < 20 and feature not in [‘Outcome’] ]
print(f’Number of discrete Columns is {len(discrete_columns)}’)
for i in discrete_columns:
print(f'{i} has {len(df_copy[i].unique())} discrete values’)
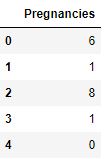
df_copy[discrete_columns].head()
Number of discrete Columns is 1 Pregnancies has 17 discrete values
Check distribution of the discrete data
for feature in discrete_columns:
dt=df_copy.copy()
dt.groupby(feature)[‘Outcome’].value_counts().plot.bar()
plt.xlabel(feature)
plt.ylabel(‘Outcome’)
plt.title(feature)
plt.show()
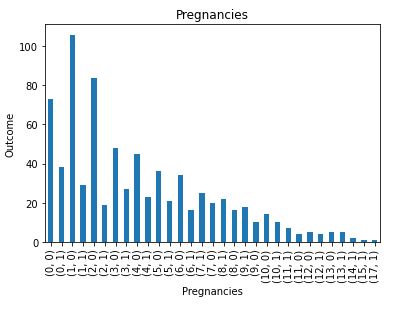
Check if any numerical columns are continuous
continuous_columns = [feature for feature in num_data if feature not in discrete_columns and feature not in [‘Outcome’] ]
print(f’Number of Continous Columns is {len(continuous_columns)}’)
df_copy[continuous_columns].head()
Number of Continuous Columns is 7
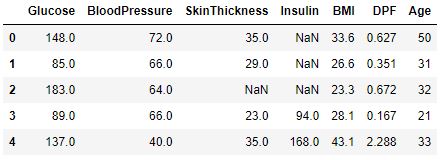
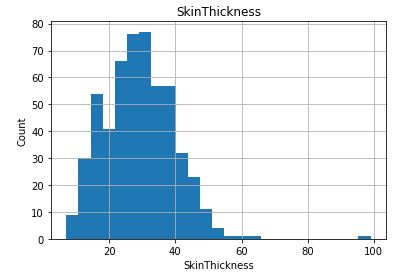
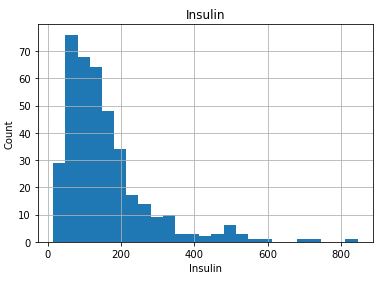
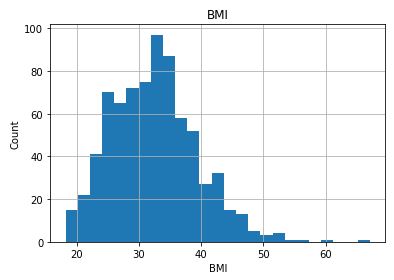
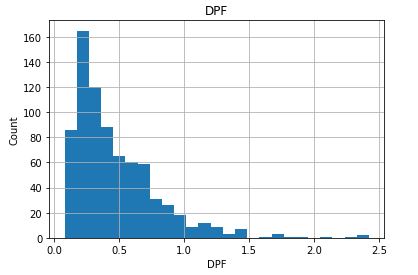
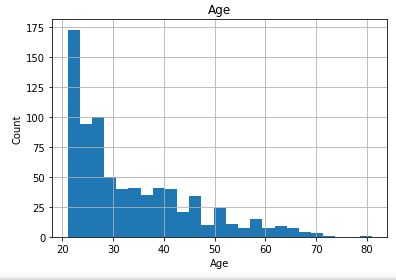
Check distribution of the Continuous data to see their outliers
for feature in continuous_columns:
dt = df_copy.copy()
dt[feature].hist(bins=25)
plt.xlabel(feature)
plt.ylabel(“Count”)
plt.title(feature)
plt.show()
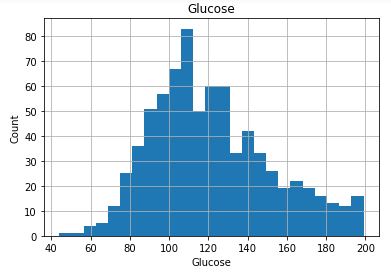
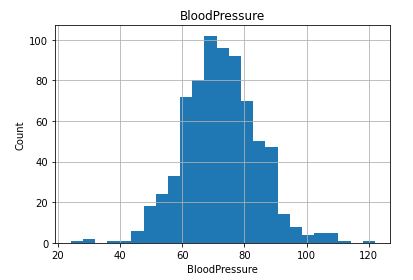
Collect all columns having missing values and at what percentage
columns_with_na=[features for features in df_copy.columns if df_copy[features].isnull().sum()>0]
for feature in columns_with_na:
print(feature, np.round(df_copy[feature].isnull().mean(), 4), ‘ % missing values’)
Glucose 0.0065 % missing values BloodPressure 0.0456 % missing values SkinThickness 0.2956 % missing values Insulin 0.487 % missing values BMI 0.0143 % missing values
Column editing: replace NaN values with median of that column and make new column as df_copy[feature_nan]. Give it value 1 if NaN is present else 0
for feature in columns_with_na:
median_value = df_copy[feature].mean()
df_copy[feature+’nan’] = np.where(df_copy[feature].isnull(), 1, 0)
df_copy[feature].fillna(median_value, inplace = True)
df_copy[columns_with_na].isnull().sum()
Glucose 0 BloodPressure 0 SkinThickness 0 Insulin 0 BMI 0 dtype: int64
df_copy.head()
| Pregnancies | Glucose | BloodPressure | SkinThickness | Insulin | BMI | DPF | Age | Outcome | Glucosenan | BloodPressurenan | SkinThicknessnan | Insulinnan | BMInan |
|---|
Remove any duplicate entry
final_df =df_copy.loc[:,~df_copy.columns.duplicated()]
final_df.shape
(768, 14)
Let’s import StandardScaler and SMOTE
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
from imblearn import over_sampling
from imblearn.over_sampling import SMOTE
oversample = SMOTE()
Model Building:
from sklearn.model_selection import train_test_split
X0 = final_df.drop(columns=’Outcome’)
let’s transform our data
X = X0
y = final_df[‘Outcome’]
X, y = oversample.fit_resample(X, y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=0)
Naive Bayes:
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB(var_smoothing=1e-06)
classifier.fit(X_train, y_train)
GaussianNB(var_smoothing=1e-06)
y_pred = classifier.predict(X_test)
y_pred
Let’s call confusion_matrix and accuracy_score
from sklearn.metrics import confusion_matrix, accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print(accuracy_score(y_test, y_pred))
[[77 28] [18 77]] 0.77
Random Forest Model + HPO:
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import RandomizedSearchCV
clf = RandomForestClassifier()
n_estimators = [10, 20, 30, 50, 100]
max_depth = [2, 3, 5, 7, 10]
Define the grid of hyperparameters to search
hyperparameter_grid = {
‘n_estimators’: n_estimators,
‘max_depth’:max_depth,
}
random_cv = RandomizedSearchCV(estimator=clf,param_distributions=hyperparameter_grid,
cv=5, n_iter=5,scoring = ‘neg_mean_absolute_error’,n_jobs = 4,verbose = 5,
return_train_score = True,random_state=42)
random_cv.fit(X_train,y_train)
random_cv.best_estimator_
Fitting 5 folds for each of 5 candidates, totalling 25 fits
Out[93]:
RandomForestClassifier(max_depth=10, n_estimators=50)
clf = RandomForestClassifier(max_depth=5, n_estimators=50)
clf.fit(X_train, y_train)
RandomForestClassifier(max_depth=5, n_estimators=50)
y_pred = clf.predict(X_test)
y_pred
Let’s check confusion_matrix and accuracy_score
from sklearn.metrics import confusion_matrix, accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print(accuracy_score(y_test, y_pred))
[[80 25] [ 8 87]] 0.835
We can see that RandomForestClassifier+RandomizedSearchCV is more accurate than GaussianNB.
Decision Tree + HPO:
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
classifier = DecisionTreeClassifier()
from scipy.stats import randint
from sklearn.model_selection import RandomizedSearchCV
parameters = {“max_depth”:[2,3, None],
“max_features”:randint(1,9),
“min_samples_leaf”:randint(1,8),
“criterion”: [“gini”, “entropy”]
}
tree_cv = RandomizedSearchCV(classifier, parameters, cv = 5)
tree_cv.fit(X_train, y_train)
tree_cv.best_estimator_
DecisionTreeClassifier(max_features=7, min_samples_leaf=3)
classifier = DecisionTreeClassifier(max_depth=15, max_features=8, min_samples_leaf=4)
classifier.fit(X_train, y_train)
DecisionTreeClassifier(max_depth=15, max_features=8, min_samples_leaf=4)
y_pred = classifier.predict(X_test)
y_pred
Let’s check confusion_matrix and accuracy_score
[[85 20] [20 75]] 0.8
Creating a pickle file for the clf model
import pickle
filename = ‘diabetes-model.pkl’
pickle.dump(clf, open(filename, ‘wb’))
In-Depth EDA+RF Analysis
Importing libraries and data
import numpy as np
import pandas as pd
import pickle
import seaborn as sns
df = pd.read_csv(‘diabetes.csv’)
and checking the content
df.head(5)
Data Visualization + EDA:
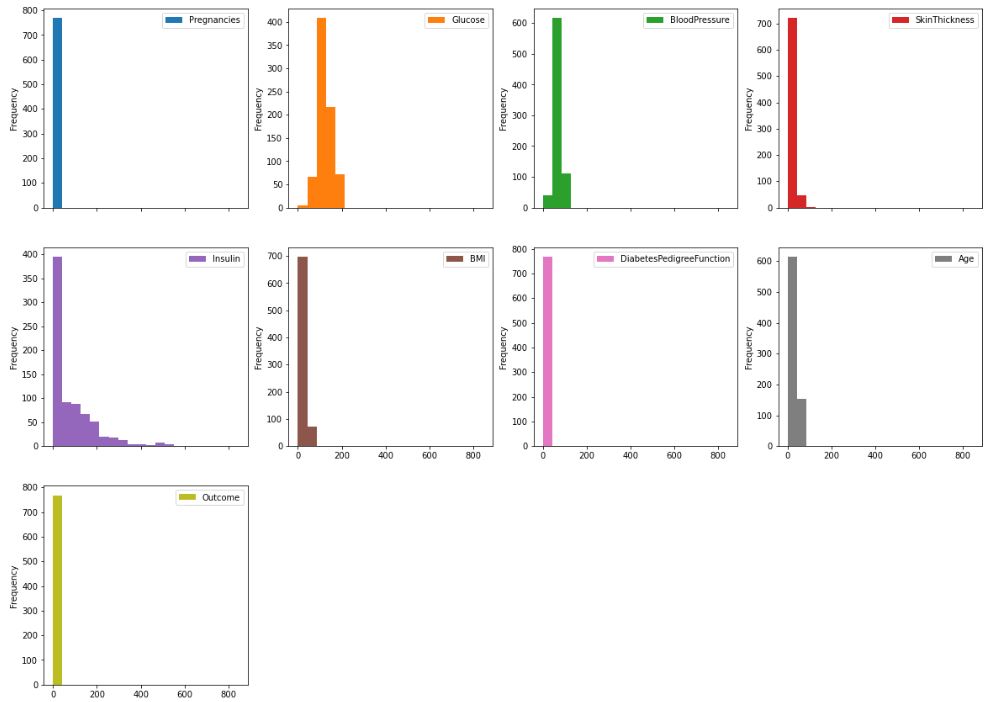
Histograms
df.plot.hist(subplots=True, layout=(4,4), figsize=(20, 20), bins=20)
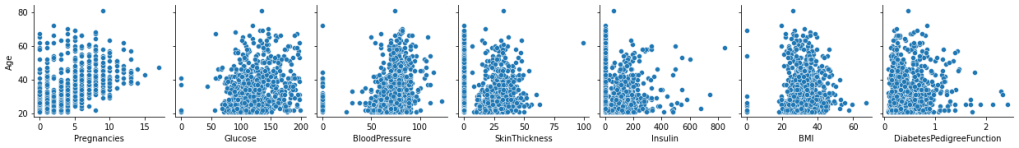
fig=sns.pairplot(
df,
x_vars=[‘Pregnancies’,’Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’,’DiabetesPedigreeFunction’],
y_vars=[‘Age’],
)
fig.savefig(“hcpairplot.png”)
df.boxplot(column=[‘Pregnancies’,’Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’,’DiabetesPedigreeFunction’,’Age’], figsize=(30,20), fontsize=24)
plt.savefig(‘hcboxplot.png’)
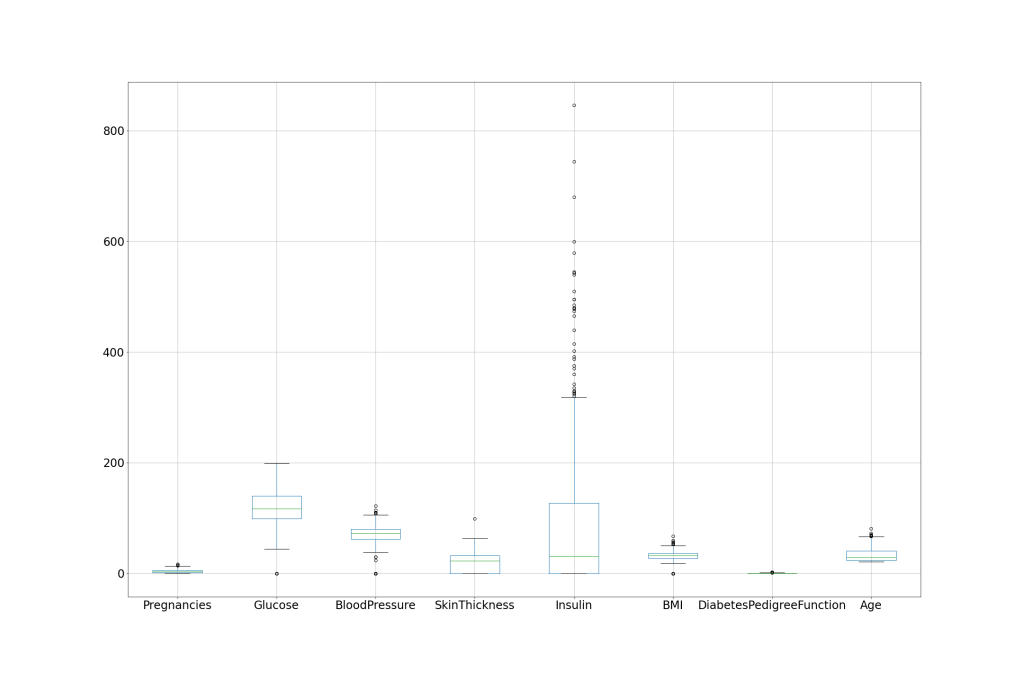
We can see that Insulin has the largest number of outliers.
Renaming DiabetesPedigreeFunction as DPF
df = df.rename(columns={‘DiabetesPedigreeFunction’:’DPF’})
Replacing the 0 values from [‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’] by NaN
df_copy = df.copy(deep=True)
df_copy[[‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’]] = df_copy[[‘Glucose’,’BloodPressure’,’SkinThickness’,’Insulin’,’BMI’]].replace(0,np.NaN)
Replacing NaN value by mean, median depending upon distribution
df_copy[‘Glucose’].fillna(df_copy[‘Glucose’].mean(), inplace=True)
df_copy[‘BloodPressure’].fillna(df_copy[‘BloodPressure’].mean(), inplace=True)
df_copy[‘SkinThickness’].fillna(df_copy[‘SkinThickness’].median(), inplace=True)
df_copy[‘Insulin’].fillna(df_copy[‘Insulin’].median(), inplace=True)
df_copy[‘BMI’].fillna(df_copy[‘BMI’].median(), inplace=True)
Random Forest Classifier Model:
from sklearn.model_selection import train_test_split
X = df.drop(columns=’Outcome’)
y = df[‘Outcome’]
X1=scaler.fit_transform(X)
X, y = oversample.fit_resample(X1, y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=0)
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(n_estimators = 50,random_state=0)
classifier.fit(X_train, y_train)
import pickle
filename = ‘RFmodel.pkl’
with open(filename, ‘wb’) as file:
pickle.dump(classifier, file)
Predicting the Test set results
y_pred = classifier.predict(X_test)
df2 = pd.DataFrame({‘Actual’:y_test,’Predicted’:y_pred})
df2
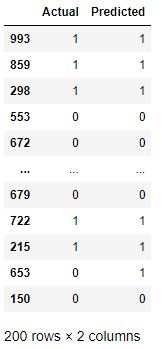
df2.plot.hist(alpha=0.5)
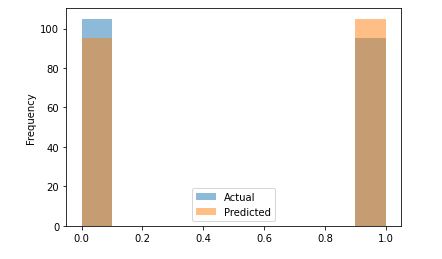
Let’s call confusion_matrix and accuracy_score
from sklearn.metrics import confusion_matrix, accuracy_score
print(“Confusion Matrix:”, confusion_matrix(y_test, y_pred))
print(“Accuracy:”,accuracy_score(y_test,y_pred))
Confusion Matrix: [[86 19] [ 9 86]] Accuracy: 0.86
Let’s check other metrics
from sklearn import metrics
print(‘Root Mean Squared Error (RMSE):’, np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
print(‘Mean Absolute Error (MAE):’, metrics.mean_absolute_error(y_test, y_pred))
print(‘Mean Squared Error (MSE):’, metrics.mean_squared_error(y_test, y_pred))
Root Mean Squared Error (RMSE): 0.37416573867739417 Mean Absolute Error (MAE): 0.14 Mean Squared Error (MSE): 0.14
Advanced Workflow
Importing libraries and data as 768 rows × 9 columns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import pickle
%matplotlib inline
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split,cross_validate,GridSearchCV
data = pd.read_csv(‘diabetes1.csv’)
data.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 768 entries, 0 to 767 Data columns (total 9 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 Pregnancies 768 non-null int64 1 Glucose 768 non-null int64 2 BloodPressure 768 non-null int64 3 SkinThickness 768 non-null int64 4 Insulin 768 non-null int64 5 BMI 768 non-null float64 6 DiabetesPedigreeFunction 768 non-null float64 7 Age 768 non-null int64 8 Outcome 768 non-null int64 dtypes: float64(2), int64(7) memory usage: 54.1 KB
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
from imblearn import over_sampling
from imblearn.over_sampling import SMOTE
oversample = SMOTE()
Let’s look at the pairplot
fig=sns.pairplot(data, hue=’Outcome’,vars = data.columns[1:-1])
fig.savefig(“hcpxcorrpairplot.png”)
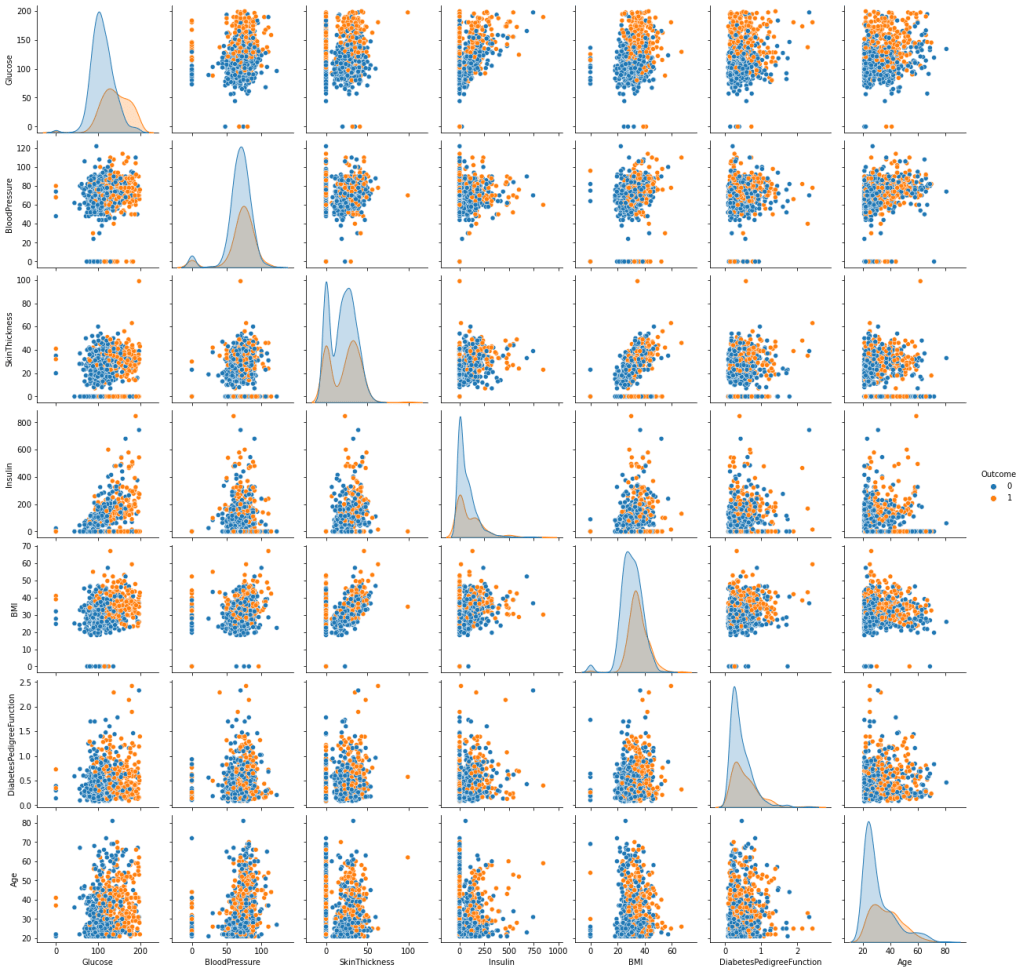

Let’s look at the correlation matrix C(*,*)
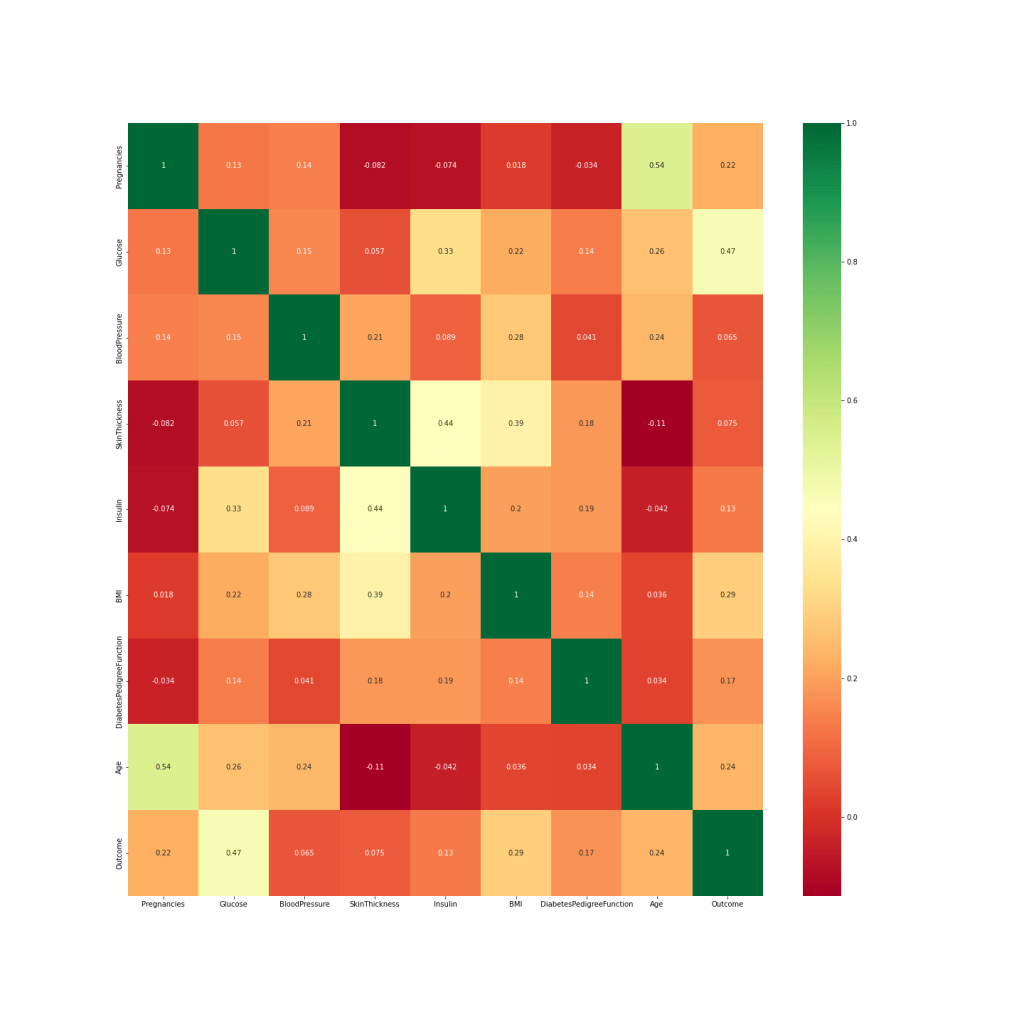
We can see that C(Glucose, Outcome)=0.47 (highest correlation), while C(BP, Outcome)=0.065 (lowest correlation). At the same time, C(Age, Pregnancies)=0.54, so we can drop one of these features such as Pregnancies.
We can check Outcome
sns.countplot(data[‘Outcome’])
and correlation values consistent with the above matrix C plot
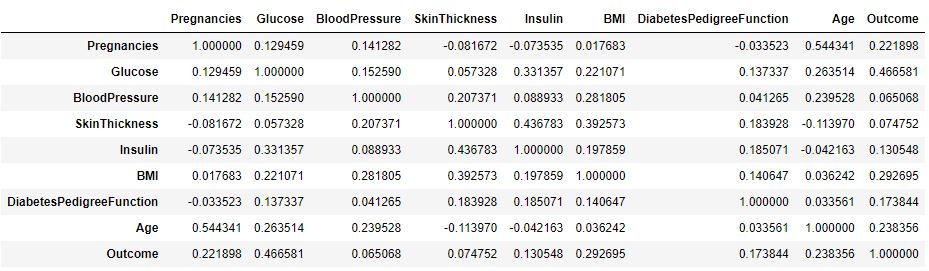
Check missing values
print(“total number of rows : {0}”.format(len(data)))
print(“number of rows missing Glucose: {0}”.format(len(data.loc[data[‘Glucose’] == 0])))
print(“number of rows missing diastolic_bp: {0}”.format(len(data.loc[data[‘BloodPressure’] == 0])))
print(“number of rows missing insulin: {0}”.format(len(data.loc[data[‘Insulin’] == 0])))
print(“number of rows missing bmi: {0}”.format(len(data.loc[data[‘BMI’] == 0])))
print(“number of rows missing diab_pred: {0}”.format(len(data.loc[data[‘DiabetesPedigreeFunction’] == 0])))
print(“number of rows missing age: {0}”.format(len(data.loc[data[‘Age’] == 0])))
print(“number of rows missing skin: {0}”.format(len(data.loc[data[‘SkinThickness’] == 0])))
total number of rows : 768 number of rows missing Glucose: 5 number of rows missing diastolic_bp: 35 number of rows missing insulin: 374 number of rows missing bmi: 11 number of rows missing diab_pred: 0 number of rows missing age: 0 number of rows missing skin: 227
Let’s replace missing values
df = data.copy()
for column in [‘Glucose’,’SkinThickness’,’Insulin’]:
median_0 = data[column][data[‘Outcome’]==0].median()
median_1 = data[column][data[‘Outcome’]==1].median()
df[column][df['Outcome']==0] = data[column][df['Outcome']==0].fillna(median_0)
df[column][df['Outcome']==1] = data[column][df['Outcome']==1].fillna(median_1)
df.BloodPressure.fillna(df.BloodPressure.median(),inplace=True)
df.BMI.fillna(df.BMI.median(),inplace=True)
Let’s create X (features) and y (target) objects
X = df.drop(‘Outcome’,axis=1)
X
y = df.Outcome
y
Let’s apply our data transformations and splitting
from sklearn.model_selection import train_test_split with test_size=0.20
X1=scaler.fit_transform(X)
X, y = oversample.fit_resample(X1, y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=0)
Creating metrics for evaluations
from sklearn import metrics
f1 = metrics.make_scorer(metrics.f1_score)
accuracy = metrics.make_scorer(metrics.accuracy_score)
precision = metrics.make_scorer(metrics.precision_score)
recall = metrics.make_scorer(metrics.recall_score)
auc = metrics.make_scorer(metrics.roc_auc_score)
scoring = {
“accuracy”:accuracy,
“precision”:precision,
“recall”: recall,
“f1”:f1,
}
def printResults(cv):
print(“Accuracy {:.3f} ({:.3f})”.format(cv[“test_accuracy”].mean(), cv[“test_accuracy”].std()))
print(“Precision {:.3f} ({:.3f})”.format(cv[“test_precision”].mean(), cv[“test_precision”].std()))
print(“Recall {:.3f} ({:.3f})”.format(cv[“test_recall”].mean(), cv[“test_recall”].std()))
print(“F1 {:.3f} ({:.3f})”.format(cv[“test_f1”].mean(), cv[“test_f1”].std()))
Creating our ML Model
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
gbc = GradientBoostingClassifier()
gbc.fit(X_train,y_train)
GradientBoostingClassifier()
Making test predictions
y_pred = gbc.predict(X_test)
Let’s check results
print(metrics.classification_report(y_test, y_pred))
precision recall f1-score support
0 0.76 0.71 0.74 96
1 0.75 0.80 0.77 104
accuracy 0.76 200
macro avg 0.76 0.75 0.75 200
weighted avg 0.76 0.76 0.75 200
cm = metrics.confusion_matrix(y_test, y_pred,normalize=’all’)
sns.heatmap(cm, annot=True, cmap=”Blues”);
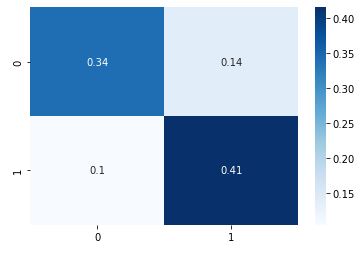
cv_gbc = cross_validate(gbc, X, y, scoring=scoring, cv=5)
printResults(cv_gbc)
Accuracy 0.787 (0.069) Precision 0.779 (0.063) Recall 0.800 (0.089) F1 0.789 (0.072)
Tuning our ML Model
params = {
‘loss’: [‘deviance’,’exponential’],
‘learning_rate’: [0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0],
‘n_estimators’: [100,200,300,400,500,600,700,800,800,1000],
}
gs = GridSearchCV(estimator = gbc,param_grid=params,cv=5)
gs.fit(X,y)
GridSearchCV(cv=5, estimator=GradientBoostingClassifier(),
param_grid={'learning_rate': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7,
0.8, 0.9, 1.0],
'loss': ['deviance', 'exponential'],
'n_estimators': [100, 200, 300, 400, 500, 600, 700,
800, 800, 1000]})
gs.best_score_
0.806
gs.best_params_
{'learning_rate': 0.7, 'loss': 'exponential', 'n_estimators': 100}
gbc_best = GradientBoostingClassifier(learning_rate=0.2,loss=’deviance’,n_estimators=200)
gbc_best.fit(X_train,y_train)
GradientBoostingClassifier(learning_rate=0.2, n_estimators=200)
Let’s make predictions
y_pred = gbc_best.predict(X_test)
print(metrics.classification_report(y_test, y_pred))
precision recall f1-score support
0 0.77 0.77 0.77 96
1 0.79 0.79 0.79 104
accuracy 0.78 200
macro avg 0.78 0.78 0.78 200
weighted avg 0.78 0.78 0.78 200
cm = metrics.confusion_matrix(y_test, y_pred,normalize=’all’)
sns.heatmap(cm, annot=True, cmap=”Blues”);
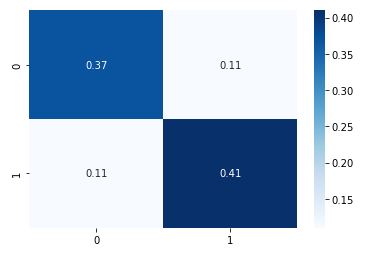
cv_gbc_best = cross_validate(gbc_best, X, y, cv=5, scoring=scoring)
printResults(cv_gbc_best)
Accuracy 0.788 (0.056) Precision 0.783 (0.030) Recall 0.794 (0.121) F1 0.785 (0.069)
Let’s invoke pickle
import pickle
filename = ‘diabetes-model.pkl’
pickle.dump(gbc_best,open(filename,’wb’))
loaded_model = pickle.load(open(‘diabetes_model_improved.pkl’, ‘rb’))
loaded_model.fit(X_train,y_train)
GradientBoostingClassifier(n_estimators=400)
y_pred = loaded_model.predict(X_test)
print(metrics.classification_report(y_test, y_pred))
precision recall f1-score support
0 0.75 0.72 0.73 96
1 0.75 0.78 0.76 104
accuracy 0.75 200
macro avg 0.75 0.75 0.75 200
weighted avg 0.75 0.75 0.75 200
cm = metrics.confusion_matrix(y_test, y_pred,normalize=’all’)
sns.heatmap(cm, annot=True, cmap=”Blues”);
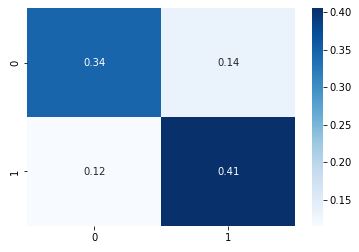
Summary
- The objective of the project is to diagnostically predict whether or not a patient has diabetes, based on certain diagnostic measurements included in the input dataset from Kaggle.
- Data editing: we replaced missing values with mean/median feature values and removed outliers.
- We applied StandardScaler and SMOTE data transformations before splitting.
- RandomForestClassifier+RandomizedSearchCV (accuracy ~86%) is more accurate than GaussianNB and DecisionTreeClassifier
- GradientBoostingClassifier + GridSearchCV
Accuracy 0.788 (0.056)
Precision 0.783 (0.030)
Recall 0.794 (0.121)
F1 0.785 (0.069)
- The model created as a result of HPO became the model with the best Cross Validation Score value, as shown above.
Explore More
HealthTech ML/AI Q3 ’22 Round-Up
The Application of ML/AI in Diabetes
Diabetes Prediction using ML/AI in Python
Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.


Leave a comment