This project was inspired by the multi-classification proof-of-concept research in the area of credit risk analytics (CRA). CRA provides a guide for risk managers looking to efficiently build credit risk management models. In recent studies, credit rating models have been constructed by relying on a number of financial ratios, including correlations between them.
The input dataset ratings.csv can be downloaded from the Github repository. The corresponding CRA code is also available.
Featured Photo by Jonathan Cooper on Unsplash.
Workflow:
- Import libraries and download input data
- Data Editing and Exploratory Data Analysis (EDA)
- Feature Engineering (FE) and importance scores
- Training/Testing multi-classification models
- ML validation and QC performance analysis
- Saving the best model and classification report
Contents:
Preparation Phase + EDA
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
import the key libraries
import pandas as pd
import numpy as np
import scikitplot as skplt
import sklearn
from sklearn.datasets import load_digits, load_boston, load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor, GradientBoostingClassifier, ExtraTreesClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
import itertools
import matplotlib.pyplot as plt
import sys
import warnings
warnings.filterwarnings(“ignore”)
print(“Scikit Plot Version : “, skplt.version)
print(“Scikit Learn Version : “, sklearn.version)
print(“Python Version : “, sys.version)
%matplotlib inline
Scikit Plot Version : 0.3.7 Scikit Learn Version : 1.0.2 Python Version : 3.9.12 (main, Apr 4 2022, 05:22:27)
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
and download the input dataset
df = pd.read_csv(“ratings.csv”)
print(df.columns)
Index(['spid', 'rating', 'COMMEQTA', 'LLPLOANS', 'COSTTOINCOME', 'ROE',
'LIQASSTA', 'SIZE'],
dtype='object')
df.head()

df.shape
(5000, 8) df.info() <class 'pandas.core.frame.DataFrame'> RangeIndex: 5000 entries, 0 to 4999 Data columns (total 8 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 spid 5000 non-null int64 1 rating 5000 non-null int64 2 COMMEQTA 5000 non-null float64 3 LLPLOANS 5000 non-null float64 4 COSTTOINCOME 5000 non-null float64 5 ROE 5000 non-null float64 6 LIQASSTA 5000 non-null float64 7 SIZE 5000 non-null float64 dtypes: float64(6), int64(2) memory usage: 312.6 KB
Let’s drop the column spid
df.drop([‘spid’], axis=1, inplace=True)
df.columns
Index(['rating', 'COMMEQTA', 'LLPLOANS', 'COSTTOINCOME', 'ROE', 'LIQASSTA',
'SIZE'],
dtype='object')
Let's create the sns pairplot

Let’s separate the feature and target columns
X = df.loc[:, df.columns != “rating”]
y = df.loc[:, df.columns == “rating”]
and split the data into training/testing subsets with test_size=0.2
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
Building Model + FE + QC
Let’s look at KNeighborsClassifier
training_accuracy = []
test_accuracy = []
try n_neighbors from 1 to 10
neighbors_settings = range(1, 11)
for n_neighbors in neighbors_settings:
fit the classification model with the above parameters
knn = KNeighborsClassifier(n_neighbors=n_neighbors)
knn.fit(X_train, y_train)
check the training set accuracy
training_accuracy.append(knn.score(X_train, y_train))
and the test set accuracy
test_accuracy.append(knn.score(X_test, y_test))
plt.plot(neighbors_settings, training_accuracy, label=”training accuracy”)
plt.plot(neighbors_settings, test_accuracy, label=”test accuracy”)
plt.ylabel(“Accuracy”)
plt.xlabel(“n_neighbors”)
plt.legend()
plt.savefig(“knn_compare_model”)
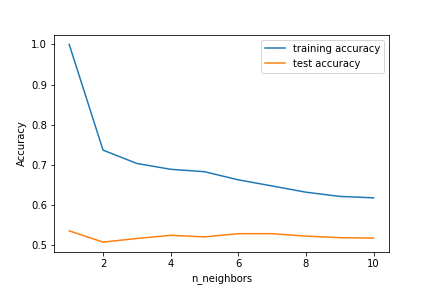
Let’s run the model with the optimal set n_neighbors=4
knn = KNeighborsClassifier(n_neighbors=4)
knn.fit(X_train, y_train)
print(“Accuracy of K-NN classifier on training set: {:.2f}”.format(knn.score(X_train, y_train)))
print(“Accuracy of K-NN classifier on test set: {:.2f}”.format(knn.score(X_test, y_test)))
Accuracy of K-NN classifier on training set: 0.69 Accuracy of K-NN classifier on test set: 0.53
let’s look at the KNN learning curve
skplt.estimators.plot_learning_curve(knn, X, y,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”KNeighborsClassifier() Learning Curve”);

Let’s compare it to LogisticRegression
logreg = LogisticRegression(random_state=42).fit(X_train, y_train)
print(“Training set score: {:.3f}”.format(logreg.score(X_train, y_train)))
print(“Test set score: {:.3f}”.format(logreg.score(X_test, y_test)))
Training set score: 0.462 Test set score: 0.463
Let’s check it against DecisionTreeClassifier
tree1 = DecisionTreeClassifier(random_state=42)
tree1.fit(X_train, y_train)
print(“Accuracy on training set: {:.3f}”.format(tree1.score(X_train, y_train)))
print(“Accuracy on test set: {:.3f}”.format(tree1.score(X_test, y_test)))
Accuracy on training set: 1.000 Accuracy on test set: 0.708
Let’s update the parameteres to avoid training set overfitting
tree2 = DecisionTreeClassifier(max_depth=10, random_state=1)
tree2.fit(X_train, y_train)
print(“Accuracy on training set: {:.3f}”.format(tree2.score(X_train, y_train)))
print(“Accuracy on test set: {:.3f}”.format(tree2.score(X_test, y_test)))
Accuracy on training set: 0.812 Accuracy on test set: 0.742
The corresponding learning curve looks as follows:
skplt.estimators.plot_learning_curve(tree2, X, y,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”DecisionTreeClassifier() Learning Curve”);

This is the best result so far.
Let’s plot Feature importances
df_features = [x for i,x in enumerate(X.columns) if i!= len(X.columns) ]
print(“Feature importances:\n{}”.format(tree2.feature_importances_))
Feature importances: [0.09359034 0.26028331 0.1721998 0.17193909 0.09117593 0.21081153]
def plot_feature_importances_credit(model):
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align=’center’)
plt.yticks(np.arange(n_features), df_features)
plt.xlabel(“Feature importance”)
plt.ylabel(“Feature”)
plt.ylim(-1, n_features)
plot_feature_importances_credit(tree2)
plt.savefig(‘feature_importance.png’)

We can see that LLPLOANS appears to be the most dominant feature.
Let’s look at RandomForestClassifier
rf = RandomForestClassifier(n_estimators=100, random_state=42)
rf.fit(X_train, y_train)
print(“Accuracy on training set: {:.3f}”.format(rf.score(X_train, y_train)))
print(“Accuracy on test set: {:.3f}”.format(rf.score(X_test, y_test)))
Accuracy on training set: 1.000 Accuracy on test set: 0.799
Let’s update the parameters
rf1 = RandomForestClassifier(max_depth=18, n_estimators=100, random_state=1)
rf1.fit(X_train, y_train)
print(“Accuracy on training set: {:.3f}”.format(rf1.score(X_train, y_train)))
print(“Accuracy on test set: {:.3f}”.format(rf1.score(X_test, y_test)))
Accuracy on training set: 0.978 Accuracy on test set: 0.800
It appears that RandomForestClassifier is more accurate than DecisionTreeClassifier.
Let’s plot feature importances
plot_feature_importances_credit(rf1)

Let’s run GradientBoostingClassifier
gb2 = GradientBoostingClassifier(learning_rate=0.2, random_state=1)
gb2.fit(X_train, y_train)
print(“Accuracy on training set: {:.3f}”.format(gb2.score(X_train, y_train)))
print(“Accuracy on test set: {:.3f}”.format(gb2.score(X_test, y_test)))
Accuracy on training set: 0.970 Accuracy on test set: 0.817
The feature importance chart is
plot_feature_importances_credit(gb2)

Let’s apply the SVC algorithm to the data scaled by StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.fit_transform(X_test)
svc2 = SVC(C=500, random_state=42)
svc2.fit(X_train_scaled, y_train)
print(“Accuracy on training set: {:.2f}”.format(svc2.score(X_train_scaled, y_train)))
print(“Accuracy on test set: {:.2f}”.format(svc2.score(X_test_scaled, y_test)))
Accuracy on training set: 0.90 Accuracy on test set: 0.71
Let’s compare it to MLPClassifier
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.fit_transform(X_test)
mlp3 = MLPClassifier(max_iter=5000, random_state=42)
mlp3.fit(X_train_scaled, y_train)
print(“Accuracy on training set: {:.3f}”.format(mlp3.score(X_train_scaled, y_train)))
print(“Accuracy on test set: {:.3f}”.format(mlp3.score(X_test_scaled, y_test)))
Accuracy on training set: 0.836 Accuracy on test set: 0.764
Let’s look at the NN weight coefficients
plt.figure(figsize=(20, 5))
plt.imshow(mlp3.coefs_[0], interpolation=”none”, cmap=’viridis’)
plt.yticks(range(len(X.columns)), df_features)
plt.xlabel(“Columns in weight matrix”)
plt.ylabel(“Input feature”)
plt.colorbar()

Finally, let’s try LogisticRegression
logreg100 = LogisticRegression(C=10, random_state=42).fit(X_train, y_train)
print(“Training set accuracy: {:.3f}”.format(logreg100.score(X_train, y_train)))
print(“Test set accuracy: {:.3f}”.format(logreg100.score(X_test, y_test)))
Training set accuracy: 0.488 Test set accuracy: 0.480
Let’s get the summary of tests accuracy
algorithms = [“k-Nearest Neighbors”, “Logistic Regression”, “Decision Trees”, “Random Forest”,
“Gradient Boosting”, “Support Vector Machine”, “Deep Learning”]
tests_accuracy = [knn.score(X_test, y_test), logreg100.score(X_test, y_test), tree2.score(X_test, y_test),
rf1.score(X_test, y_test), gb2.score(X_test, y_test), svc2.score(X_test_scaled, y_test),
mlp3.score(X_test_scaled, y_test)]
compare_algorithms = pd.DataFrame({ “Algorithms”: algorithms, “Tests Accuracy”: tests_accuracy })
compare_algorithms.sort_values(by = “Tests Accuracy”, ascending = False)

Let’s plot this table as the bar chart
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=(8,8))
sns.barplot(x = “Tests Accuracy”, y = “Algorithms”, data = compare_algorithms)
plt.show()

The Gradient Boosting (GB) algorithm gives us the sense of achieving the highest test accuracy of 81.7%.
Performance
Let’s check the GB model performance
model = gb2
model
GradientBoostingClassifier(learning_rate=0.2, random_state=1)
Let’s make test GB predictions
predictions = model.predict(X_test)
and compute the GB confusion matrix
cm = confusion_matrix(y_true=y_test, y_pred=predictions)
Let’s define the following plot function
def plot_confusion_matrix(cm, classes,
normalize=False,
title=’Confusion matrix’,
cmap=plt.cm.Blues):
“””
This function prints and plots the confusion matrix.
Normalization can be applied by setting normalize=True.
“””
plt.imshow(cm, interpolation=’nearest’, cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
Let’s plot the GB confusion matrix with the labels
cm_plot_labels = [‘1′,’2′,’3′,’4′,’5′,’6′,’7′,’8′,’9′,’10’]
plot_confusion_matrix(cm=cm, classes=cm_plot_labels, title=’Confusion Matrix’)
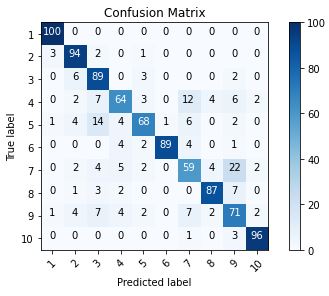

Let’s print our predictions
print(“\033[1m The result is telling us that we have: “,(cm[0,0] + cm[1,1] + cm[2,2] + cm[3,3] + cm[4,4] +
cm[5,5] + cm[6,6] + cm[7,7] + cm[8,8] + cm[9,9] ),”correct predictions.”)
print(“\033[1m The result is telling us that we have: “, ( cm.sum() – (cm[0,0] + cm[1,1] + cm[2,2] +
cm[3,3] + cm[4,4] + cm[5,5] + cm[6,6] + cm[7,7] +
cm[8,8] + cm[9,9] )),”incorrect predictions.”)
print(“\033[1m We have a total predictions of: “,( cm.sum()) )
The result is telling us that we have: 817 correct predictions (81.7%). The result is telling us that we have: 183 incorrect predictions (18.3%). We have a total predictions of: 1000
Let’s print the multi-classification report
print(classification_report(y_test, predictions))
precision recall f1-score support
1 0.95 1.00 0.98 100
2 0.83 0.94 0.88 100
3 0.71 0.89 0.79 100
4 0.77 0.64 0.70 100
5 0.84 0.68 0.75 100
6 0.99 0.89 0.94 100
7 0.66 0.59 0.62 100
8 0.90 0.87 0.88 100
9 0.62 0.71 0.66 100
10 0.94 0.96 0.95 100
accuracy 0.82 1000
macro avg 0.82 0.82 0.82 1000
weighted avg 0.82 0.82 0.82 1000
Let’s plot the GB learning curve
skplt.estimators.plot_learning_curve(gb2, X, y,
cv=7, shuffle=True, scoring=”accuracy”,
n_jobs=-1, figsize=(6,4), title_fontsize=”large”, text_fontsize=”large”,
title=”GradientBoostingClassifier() Learning Curve”);

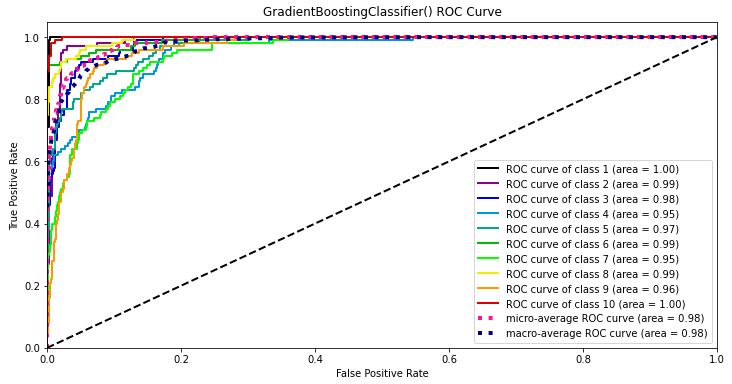
Let’s plot the GB ROC curve
Y_test_probs = gb2.predict_proba(X_test)
skplt.metrics.plot_roc_curve(y_test, Y_test_probs,
title=”GradientBoostingClassifier() ROC Curve”, figsize=(12,6));
skplt.metrics.plot_precision_recall_curve(y_test, Y_test_probs,
title=”GradientBoostingClassifier() Precision-Recall Curve”, figsize=(12,6));

Let’s look at the GB elbow plot
skplt.cluster.plot_elbow_curve(KMeans(random_state=1),
X_train,
cluster_ranges=range(2, 20),
figsize=(8,6));

Let’s perform the KMeans cluster analysis
kmeans = KMeans(n_clusters=10, random_state=1)
kmeans.fit(X_train, y_train)
cluster_labels = kmeans.predict(X_test)
skplt.metrics.plot_silhouette(X_test, cluster_labels,
figsize=(8,6));

Let’s look at the PCA components
pca = PCA(random_state=1)
pca.fit(X_train)
skplt.decomposition.plot_pca_component_variance(pca, figsize=(8,6));

Summary
- Machine Learning (ML) algorithms have been used to classify multiple outcomes in credit ratings datasets, including random forests (RF), KNeighbors (KNN), decision tree (DT), Gradient Boosting (GB), logistic regression (LR), artificial neural networks (ANN), and support vector machine (SVM).
- The Gradient Boosting (GB) algorithm has given us the sense of achieving the highest test accuracy of 81.7%.
- The GB multi-classification report consists of 10 labels:
label precision recall f1-score support
1 0.95 1.00 0.98 100
2 0.83 0.94 0.88 100
3 0.71 0.89 0.79 100
4 0.77 0.64 0.70 100
5 0.84 0.68 0.75 100
6 0.99 0.89 0.94 100
7 0.66 0.59 0.62 100
8 0.90 0.87 0.88 100
9 0.62 0.71 0.66 100
10 0.94 0.96 0.95 100
accuracy 0.82 1000
macro avg 0.82 0.82 0.82 1000
weighted avg 0.82 0.82 0.82 1000
- The micro-average GB ROC curve area is 0.98
- The micro-average GB precision-recall curve area is 0.907
- The optimal number of clusters is 6 (see the elbow plot)
- The Silhouette analysis score is 0.345
- The explained variance ratio for first PCA components is 0.946.
- We have derived FE importance coefficients that weight the company’s financial ratios and reflect the borrower’s relevant credit rating or probability of default.
The proposed ML approach can be used to assess the probability of default or to determine the relevant credit rating well beyond classical operating profitability and internal liquidity ratios. Potential stakeholders of this technology are those responsible for providing credit in banks and insurance companies. Shareholders, investors and executives are also required to assess the risk and financing policy of their investments or enterprises. This technology enables them to examine the degree of risk of their clients.
Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.


Leave a comment