Operating rooms (ORs) are some of the most valuable hospital assets, generating a large part of hospital revenue. For efficient utilization of ORs, accurate schedules of assigned block time and sequences of patient cases need to be made [1]. Statistical models have been developed using datasets to predict daily surgical volumes weeks in advance. The quest was motivated by the need to make real-time adjustments to staff capacity and reallocation of the OR block time based on predicted future demand.

Following previous studies [3,4], we focus on the VUMC dataset for evaluation of our statistical models. The dataset represents a 48-week surgery schedule for VUMC recorded from October 10, 2011 to September 14, 2012.
Our method consists of the ANOVA null-hypothesis test for the total number of surgeries to see if these variables change from day to day with 99% confidence. The null hypothesis H0 is the statement that the total number of surgeries does not depend upon the weekday (DOW). The alternative hypothesis states that the total number of surgeries does vary from day to day.
Next, the Ordinary Least Squares (OLS) linear regression formula [3,4] is applied to our target variable Actual Surgery as a function of 28 independent variables, T-28 to T-1.
The OLS regression results are assessed using the following metrics [4]: (adjusted) R2, F-statistic, Log-Likelihood, AIC, BIC, std err, MSE, MAE, and RMSE.
Contents:
Workflow
- Import/install libraries
- Read the input dataset
- Exploratory Data Analysis (EDA)
- ANOVA H0-H1 test
- Build OLS regression model
- Model performance QC
- Report summary
Ipynb Script
Let’s set the working directory YOURPATH
os.chdir(‘YOURPATH’)
and import the libraries of interest
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from sklearn import metrics
import warnings
warnings.filterwarnings(‘ignore’)
Let’s read the Excel dataset
df = pd.read_excel(‘Dataset.xlsx’)
and check the content
df.head()

df.shape
(241, 19)
df.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 241 entries, 0 to 240 Data columns (total 19 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 SurgDate 241 non-null datetime64[ns] 1 DOW 241 non-null object 2 T - 28 241 non-null int64 3 T - 21 241 non-null int64 4 T - 14 241 non-null int64 5 T - 13 241 non-null int64 6 T - 12 241 non-null int64 7 T - 11 241 non-null int64 8 T - 10 241 non-null int64 9 T - 9 241 non-null int64 10 T - 8 241 non-null int64 11 T - 7 241 non-null int64 12 T - 6 241 non-null int64 13 T - 5 241 non-null int64 14 T - 4 241 non-null int64 15 T - 3 241 non-null int64 16 T - 2 241 non-null int64 17 T - 1 241 non-null int64 18 Actual 241 non-null int64 dtypes: datetime64[ns](1), int64(17), object(1) memory usage: 35.9+ KB
Let’s get the descriptive statistics info
df.describe()
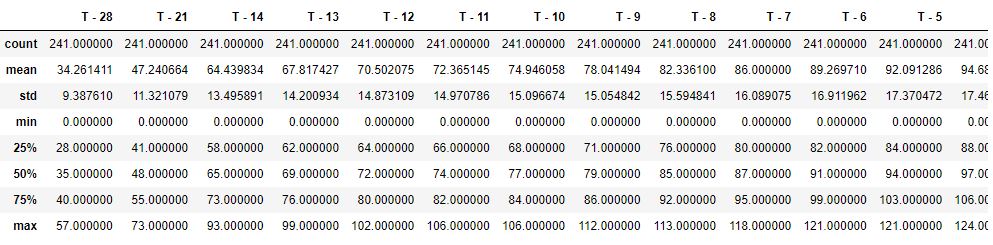
Let’s calculate correlations
df.corr()
| T – 28 | T – 21 | T – 14 | T – 13 | T – 12 | T – 11 | T – 10 | T – 9 | T – 8 | T – 7 | T – 6 | T – 5 | T – 4 | T – 3 | T – 2 | T – 1 | Actual | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T – 28 | 1.000000 | 0.894700 | 0.766981 | 0.761258 | 0.764272 | 0.769680 | 0.744281 | 0.718607 | 0.697891 | 0.669865 | 0.669421 | 0.679711 | 0.685468 | 0.686128 | 0.655022 | 0.629432 | 0.608290 |
| T – 21 | 0.894700 | 1.000000 | 0.871427 | 0.862506 | 0.849120 | 0.839669 | 0.821875 | 0.807351 | 0.794639 | 0.769279 | 0.771311 | 0.766765 | 0.766230 | 0.763745 | 0.742956 | 0.718364 | 0.702459 |
| T – 14 | 0.766981 | 0.871427 | 1.000000 | 0.975593 | 0.940374 | 0.918844 | 0.913420 | 0.924774 | 0.919929 | 0.900452 | 0.890108 | 0.863536 | 0.846024 | 0.845696 | 0.848112 | 0.821478 | 0.800877 |
| T – 13 | 0.761258 | 0.862506 | 0.975593 | 1.000000 | 0.977337 | 0.955026 | 0.941554 | 0.940412 | 0.931122 | 0.914450 | 0.911955 | 0.895554 | 0.878267 | 0.870565 | 0.862705 | 0.835040 | 0.812730 |
| T – 12 | 0.764272 | 0.849120 | 0.940374 | 0.977337 | 1.000000 | 0.986618 | 0.962074 | 0.941533 | 0.922158 | 0.904064 | 0.912807 | 0.919413 | 0.910958 | 0.893899 | 0.876955 | 0.847387 | 0.818714 |
| T – 11 | 0.769680 | 0.839669 | 0.918844 | 0.955026 | 0.986618 | 1.000000 | 0.979289 | 0.947764 | 0.918142 | 0.896470 | 0.906488 | 0.920257 | 0.923938 | 0.908863 | 0.885674 | 0.851878 | 0.819855 |
| T – 10 | 0.744281 | 0.821875 | 0.913420 | 0.941554 | 0.962074 | 0.979289 | 1.000000 | 0.973322 | 0.935192 | 0.912204 | 0.918598 | 0.922247 | 0.927982 | 0.926197 | 0.907966 | 0.871200 | 0.842193 |
| T – 9 | 0.718607 | 0.807351 | 0.924774 | 0.940412 | 0.941533 | 0.947764 | 0.973322 | 1.000000 | 0.971532 | 0.955061 | 0.945678 | 0.933364 | 0.925826 | 0.924528 | 0.922874 | 0.895139 | 0.872890 |
| T – 8 | 0.697891 | 0.794639 | 0.919929 | 0.931122 | 0.922158 | 0.918142 | 0.935192 | 0.971532 | 1.000000 | 0.984829 | 0.969236 | 0.948335 | 0.930065 | 0.920350 | 0.927708 | 0.909233 | 0.887675 |
| T – 7 | 0.669865 | 0.769279 | 0.900452 | 0.914450 | 0.904064 | 0.896470 | 0.912204 | 0.955061 | 0.984829 | 1.000000 | 0.984542 | 0.960000 | 0.938392 | 0.925503 | 0.934284 | 0.918124 | 0.895779 |
| T – 6 | 0.669421 | 0.771311 | 0.890108 | 0.911955 | 0.912807 | 0.906488 | 0.918598 | 0.945678 | 0.969236 | 0.984542 | 1.000000 | 0.983981 | 0.963228 | 0.946564 | 0.950649 | 0.927954 | 0.898900 |
| T – 5 | 0.679711 | 0.766765 | 0.863536 | 0.895554 | 0.919413 | 0.920257 | 0.922247 | 0.933364 | 0.948335 | 0.960000 | 0.983981 | 1.000000 | 0.984911 | 0.964317 | 0.959692 | 0.937331 | 0.902827 |
| T – 4 | 0.685468 | 0.766230 | 0.846024 | 0.878267 | 0.910958 | 0.923938 | 0.927982 | 0.925826 | 0.930065 | 0.938392 | 0.963228 | 0.984911 | 1.000000 | 0.984158 | 0.968785 | 0.943132 | 0.906040 |
| T – 3 | 0.686128 | 0.763745 | 0.845696 | 0.870565 | 0.893899 | 0.908863 | 0.926197 | 0.924528 | 0.920350 | 0.925503 | 0.946564 | 0.964317 | 0.984158 | 1.000000 | 0.983117 | 0.950928 | 0.913242 |
| T – 2 | 0.655022 | 0.742956 | 0.848112 | 0.862705 | 0.876955 | 0.885674 | 0.907966 | 0.922874 | 0.927708 | 0.934284 | 0.950649 | 0.959692 | 0.968785 | 0.983117 | 1.000000 | 0.970063 | 0.936430 |
| T – 1 | 0.629432 | 0.718364 | 0.821478 | 0.835040 | 0.847387 | 0.851878 | 0.871200 | 0.895139 | 0.909233 | 0.918124 | 0.927954 | 0.937331 | 0.943132 | 0.950928 | 0.970063 | 1.000000 | 0.964727 |
| Actual | 0.608290 | 0.702459 | 0.800877 | 0.812730 | 0.818714 | 0.819855 | 0.842193 | 0.872890 | 0.887675 | 0.895779 | 0.898900 | 0.902827 | 0.906040 | 0.913242 | 0.936430 | 0.964727 | 1.000000 |
We can plot the correlation matrix as an image
f = plt.figure(figsize=(19, 15))
plt.matshow(df.corr(), fignum=f.number)
plt.xticks(range(df.select_dtypes([‘number’]).shape[1]), df.select_dtypes([‘number’]).columns, fontsize=14, rotation=45)
plt.yticks(range(df.select_dtypes([‘number’]).shape[1]), df.select_dtypes([‘number’]).columns, fontsize=14)
cb = plt.colorbar()
cb.ax.tick_params(labelsize=14)
plt.title(‘Correlation Matrix’, fontsize=16);
plt.savefig(‘corrmatrix.png’)
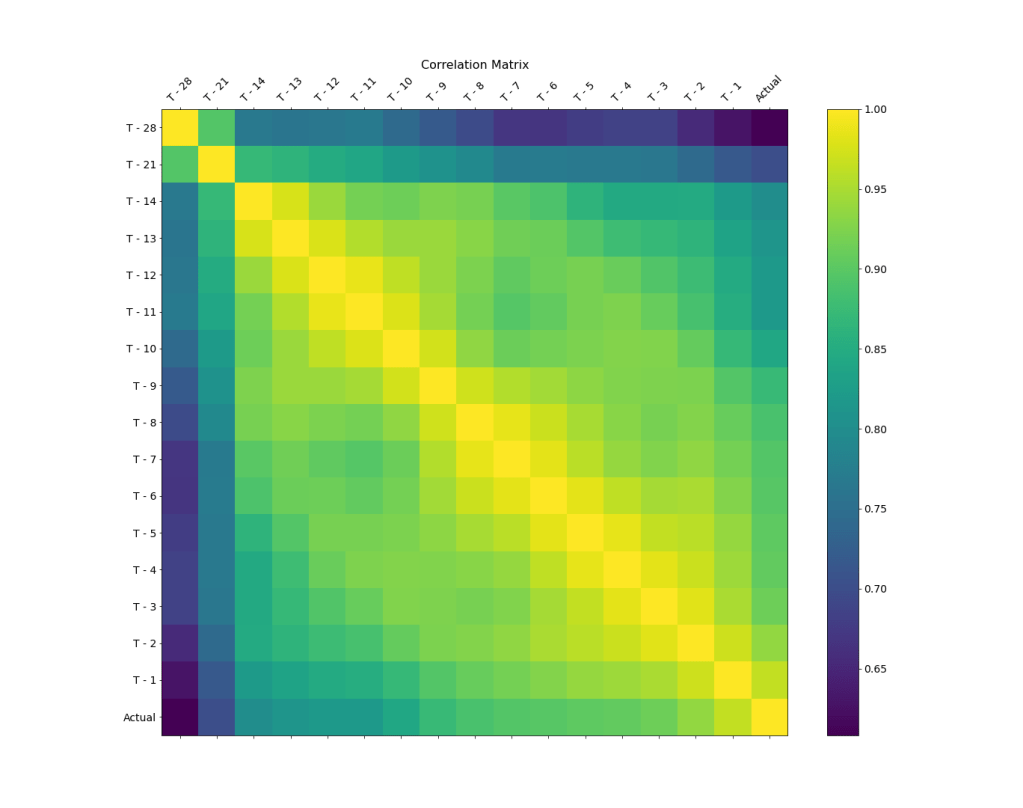
or a lower triangular matrix with annotations

We can see that the surgery volume correlation value decreases as the gap from the surgery day increases.
Let’s look at the mean and std of the Actual data grouped by DOW
df.groupby(‘DOW’)[‘Actual’].mean(), df.groupby(‘DOW’)[‘Actual’].std()
(DOW Fri 105.612245 Mon 116.255319 Thu 124.083333 Tue 119.081633 Wed 117.041667 Name: Actual, dtype: float64, DOW Fri 26.357175 Mon 18.456138 Thu 10.379672 Tue 10.864385 Wed 11.240047 Name: Actual, dtype: float64)
Let’s look at the mean and std of the T-1 data grouped by DOW
df.groupby(‘DOW’)[‘T – 1’].mean(), df.groupby(‘DOW’)[‘T – 1’].std()
(DOW Fri 97.428571 Mon 110.787234 Thu 117.583333 Tue 114.020408 Wed 110.416667 Name: T - 1, dtype: float64, DOW Fri 25.922159 Mon 18.863279 Thu 10.394011 Tue 10.296621 Wed 11.100764 Name: T - 1, dtype: float64)
Let’s calculate the number of surgeries per weekday
df[‘DOW’].value_counts()
Tue 49 Fri 49 Wed 48 Thu 48 Mon 47 Name: DOW, dtype: int64
Let’s create the barplot Actual-DOW
df2 = df[[‘DOW’, ‘Actual’]]
sns.color_palette(“flare”, as_cmap=True)
ax = sns.barplot(x=’DOW’,y=’Actual’, data=df)
ax.set(ylim=(80, 130))
plt.savefig(‘barplotactualdow.png’)

The corresponding boxplot is as follows
sns.color_palette(“flare”, as_cmap=True)
ax=sns.boxplot(x=’DOW’,y=’Actual’, data=df)
ax.set(ylim=(80, 150))
plt.savefig(‘boxplotactualdow.png’)
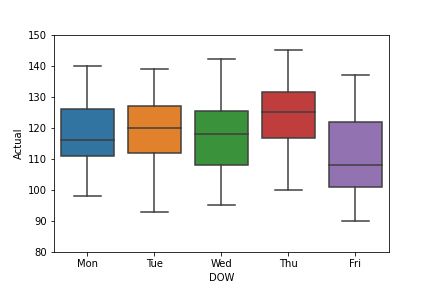
We can also look at the violine plot
sns.color_palette(“flare”, as_cmap=True)
ax=sns.violinplot(x=’DOW’,y=’Actual’, data=df)
plt.savefig(‘violineplotactualdow.png’)
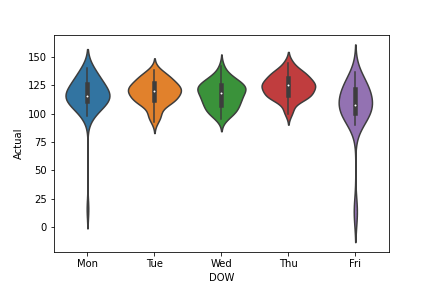
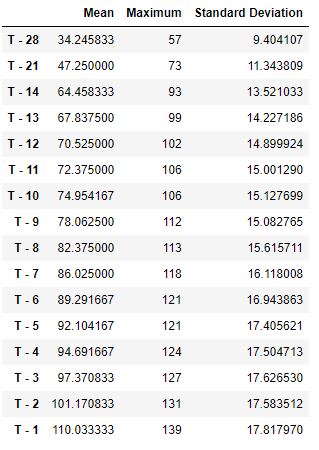
Let’s create the table new with 3 columns [‘Mean’, ‘Maximum’, ‘Standard Deviation’]
test=df.loc[1:242, ‘T – 28′:’T – 1’]
mean = test.mean()
maximum = test.max()
std = test.std()
a = pd.DataFrame(mean)
b = pd.DataFrame(maximum)
c = pd.DataFrame(std)
new = pd.concat([a,b,c], axis = 1)
new.columns = [‘Mean’, ‘Maximum’, ‘Standard Deviation’]
new
Let’s perform the OLS fitting while running the ANOVA H0-H1 test
import statsmodels.api as sm
from statsmodels.formula.api import ols
model = ols(‘Actual ~ DOW’, data=df).fit()
aov_table = sm.stats.anova_lm(model, typ=2)
aov_table
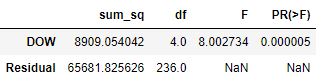
We reject the H0 hypothesis at the 99% confidence since p-value is less than 0.05.
Let’s perform multiple pairwise comparison of mean surgery volumes using the Tukey HSD test. The idea is to examine the difference between daily surgery volumes by taking various combinations of any two (randomly selected) weekdays
from statsmodels.stats.multicomp import pairwise_tukeyhsd
tukey = pairwise_tukeyhsd(endog=df[‘Actual’], groups=df[‘DOW’], alpha=0.05)
print(tukey)
Multiple Comparison of Means - Tukey HSD, FWER=0.05 ===================================================== group1 group2 meandiff p-adj lower upper reject ----------------------------------------------------- Fri Mon 10.6431 0.017 1.2798 20.0063 True Fri Thu 18.4711 0.0 9.1577 27.7844 True Fri Tue 13.4694 0.0008 4.2042 22.7346 True Fri Wed 11.4294 0.0077 2.1161 20.7428 True Mon Thu 7.828 0.1529 -1.5829 17.2389 False Mon Tue 2.8263 0.9212 -6.537 12.1896 False Mon Wed 0.7863 0.9994 -8.6246 10.1972 False Thu Tue -5.0017 0.5789 -14.3151 4.3117 False Thu Wed -7.0417 0.2376 -16.4029 2.3196 False Tue Wed -2.04 0.9747 -11.3533 7.2734 False -----------------------------------------------------
This test confirms significant differences between various combinations of daily surgery volumes by considering any two (randomly selected) weekdays.
We are ready to build and deploy our regression model(s).
Let’s prepare the data by excluding the ouliers (cf. boxplot and violineplot)
df1 = df.drop(columns = [‘SurgDate’, ‘DOW’])
from scipy import stats
df1 = df1[(np.abs(stats.zscore(df1)) < 3).all(axis=1)]
Let’s begin with the T-1, T-2, and T-3 variables x being related to the target variable y as follows
x = df[[‘T – 3’, ‘T – 2’, ‘T – 1’]]
y = df[[‘Actual’]]
First, let’s try sklearn linear_model
from sklearn import linear_model
import statsmodels.api as sm
regr = linear_model.LinearRegression()
regr.fit(x, y)
print(‘Intercept: \n’, regr.intercept_)
print(‘Coefficients: \n’, regr.coef_)
Intercept: [10.96140663] Coefficients: [[-0.14256711 0.15693554 0.94016588]]
Second, we invoke OLS statsmodels
model = sm.OLS(y, x).fit()
predictions = model.predict(x)
print_model = model.summary()
print(print_model)
OLS Regression Results
=======================================================================================
Dep. Variable: Actual R-squared (uncentered): 0.998
Model: OLS Adj. R-squared (uncentered): 0.998
Method: Least Squares F-statistic: 4.535e+04
Date: Mon, 27 Jun 2022 Prob (F-statistic): 0.00
Time: 11:39:33 Log-Likelihood: -725.88
No. Observations: 241 AIC: 1458.
Df Residuals: 238 BIC: 1468.
Df Model: 3
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
T - 3 -0.1908 0.099 -1.926 0.055 -0.386 0.004
T - 2 0.1453 0.127 1.144 0.254 -0.105 0.395
T - 1 1.0908 0.069 15.891 0.000 0.956 1.226
==============================================================================
Omnibus: 4.980 Durbin-Watson: 1.916
Prob(Omnibus): 0.083 Jarque-Bera (JB): 4.972
Skew: 0.245 Prob(JB): 0.0833
Kurtosis: 3.506 Cond. No. 89.7
==============================================================================
Notes:
[1] R² is computed without centering (uncentered) since the model does not contain a constant.
[2] Standard Errors assume that the covariance matrix of the errors is correctly specified.
We can also fit the simple linear regression model
from sklearn.linear_model import LinearRegression
regressorObject=LinearRegression()
regressorObject.fit(x,y)
Predict the number of surgeries:
predict = df1.loc[2:242, ‘T – 3′:’T – 1’]
y_pred_test_data=regressorObject.predict(predict)
Predicted = pd.DataFrame(y_pred_test_data, columns = [‘Predicted’])
result = pd.concat([df, Predicted], ignore_index=True, axis=1)
result.columns = [‘SurgDate’, ‘DOW’, ‘T – 28’, ‘T – 21’, ‘T – 14’, ‘T – 13’, ‘T – 12’, ‘T – 11′,’ T – 10′, ‘T – 9’,
‘T – 8’, ‘T – 7’, ‘T – 6’, ‘T – 5’, ‘T – 4’, ‘T – 3’, ‘T – 2’, ‘T – 1’, ‘Actual’, ‘Predicted’]
result[‘Predicted’] = round(result[‘Predicted’])
final = result.dropna()
final.tail(10)

Let’s calculate RMS
from sklearn.metrics import mean_squared_error
from math import sqrt
y_actual = final[‘Actual’]
y_predicted = final[‘Predicted’]
rms = sqrt(mean_squared_error(y_actual, y_predicted))
rms
20.653782302625718
Let’s calculate other performance metrics
y = final[‘Actual’]
yhat = final[‘Predicted’]
d = y – yhat
mse_f = np.mean(d2)
mae_f = np.mean(abs(d)) rmse_f = np.sqrt(mse_f) r2_f = 1-(sum(d2)/sum((y-np.mean(y))**2))
print(“Results by manual calculation:”)
print(“MAE:”,mae_f)
print(“MSE:”, mse_f)
print(“RMSE:”, rmse_f)
print(“R-Squared:”, r2_f)
Results by manual calculation: MAE: 14.953191489361702 MSE: 426.5787234042553 RMSE: 20.653782302625718 R-Squared: -0.3528353839078937
Recall that the above model was applied to the three variables ‘T – 3′:’T – 1’.
Now let us consider 16 variables. We want to predict results for 3 days before the Surgery Date
x = df1[[‘T – 28’, ‘T – 21’, ‘T – 14’, ‘T – 13’, ‘T – 12’, ‘T – 11’, ‘T – 10’, ‘T – 9’, ‘T – 8’, ‘T – 7’,
‘T – 6’, ‘T – 5’, ‘T – 4’, ‘T – 3’, ‘T – 2’, ‘T – 1’]]
y = df1[[‘Actual’]]
by predicting results for 3 days before the Surgery Date.
Let’s re-apply our regression models
With sklearn linear_model
from sklearn import linear_model
import statsmodels.api as sm
regr = linear_model.LinearRegression()
regr.fit(x, y)
print(‘Intercept: \n’, regr.intercept_)
print(‘Coefficients: \n’, regr.coef_)
Intercept: [14.84638197] Coefficients: [[-0.03590406 0.0856059 -0.10080439 0.07939623 0.08764874 -0.20750371 0.03761113 0.10250478 0.00932406 0.12867575 -0.11420405 -0.03460241 0.06443631 -0.15945052 0.12085577 0.87903786]]
With OLS statsmodels
model = sm.OLS(y, x).fit()
predictions = model.predict(x)
print_model = model.summary()
print(print_model)
OLS Regression Results
=======================================================================================
Dep. Variable: Actual R-squared (uncentered): 0.998
Model: OLS Adj. R-squared (uncentered): 0.998
Method: Least Squares F-statistic: 8646.
Date: Mon, 27 Jun 2022 Prob (F-statistic): 4.06e-299
Time: 11:58:15 Log-Likelihood: -705.05
No. Observations: 237 AIC: 1442.
Df Residuals: 221 BIC: 1498.
Df Model: 16
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
T - 28 -0.0402 0.079 -0.507 0.612 -0.196 0.116
T - 21 0.0955 0.084 1.142 0.254 -0.069 0.260
T - 14 -0.1061 0.126 -0.845 0.399 -0.354 0.141
T - 13 0.1548 0.189 0.821 0.413 -0.217 0.526
T - 12 -0.0594 0.214 -0.278 0.781 -0.480 0.362
T - 11 -0.0900 0.210 -0.429 0.669 -0.504 0.324
T - 10 -0.0144 0.168 -0.086 0.932 -0.345 0.316
T - 9 0.1131 0.150 0.756 0.451 -0.182 0.408
T - 8 -0.0078 0.152 -0.052 0.959 -0.307 0.291
T - 7 0.2021 0.178 1.133 0.259 -0.149 0.554
T - 6 -0.1933 0.193 -1.002 0.317 -0.573 0.187
T - 5 -0.0897 0.182 -0.492 0.623 -0.448 0.269
T - 4 0.1104 0.183 0.605 0.546 -0.249 0.470
T - 3 -0.1945 0.159 -1.227 0.221 -0.507 0.118
T - 2 0.1490 0.138 1.083 0.280 -0.122 0.420
T - 1 1.0405 0.072 14.501 0.000 0.899 1.182
==============================================================================
Omnibus: 6.221 Durbin-Watson: 1.995
Prob(Omnibus): 0.045 Jarque-Bera (JB): 7.642
Skew: 0.204 Prob(JB): 0.0219
Kurtosis: 3.779 Cond. No. 314.
==============================================================================
Notes:
[1] R² is computed without centering (uncentered) since the model does not contain a constant.
[2] Standard Errors assume that the covariance matrix of the errors is correctly specified.
Fitting Simple Linear regression data model
from sklearn.linear_model import LinearRegression
regressorObject=LinearRegression()
regressorObject.fit(x,y)
Predicting the number of surgeries
predict = df1.loc[2:242, ‘T – 28′:’T – 1’]
y_pred_test_data=regressorObject.predict(predict)
Predicted = pd.DataFrame(y_pred_test_data, columns = [‘Predicted’])
result2 = pd.concat([df, Predicted], ignore_index=True, axis=1)
result2.columns = [‘SurgDate’, ‘DOW’, ‘T – 28’, ‘T – 21’, ‘T – 14’, ‘T – 13’, ‘T – 12’, ‘T – 11′,’ T – 10′, ‘T – 9’,
‘T – 8’, ‘T – 7’, ‘T – 6’, ‘T – 5’, ‘T – 4’, ‘T – 3’, ‘T – 2’, ‘T – 1’, ‘Actual’, ‘Predicted’]
result2[‘Predicted’] = round(result2[‘Predicted’])
final2 = result2.dropna()
round(result[‘Actual’].mean()), round(result2[‘Predicted’].mean())
(116, 118)
We can see that the Average Daily Error of surgeries is ~2 Surgeries.
Let’s check RMS
from sklearn.metrics import mean_squared_error
from math import sqrt
y_actual = final2[‘Actual’]
y_predicted = final2[‘Predicted’]
rms = sqrt(mean_squared_error(y_actual, y_predicted))
rms
20.367475114829315
Let’s calculate other performance metrics
y = final2[‘Actual’]
yhat = final2[‘Predicted’]
d = y – yhat
mse_f = np.mean(d2) mae_f = np.mean(abs(d)) rmse_f = np.sqrt(mse_f) r2_f = 1-(sum(d2)/sum((y-np.mean(y))**2))
print(“Results by manual calculation:”)
print(“MAE:”,mae_f)
print(“MSE:”, mse_f)
print(“RMSE:”, rmse_f)
print(“R-Squared:”, r2_f)
Results by manual calculation: MAE: 14.697872340425532 MSE: 414.83404255319147 RMSE: 20.367475114829315 R-Squared: -0.3155887540215563
Predicting results for 7 days before the surgery date.
Let’s set x and y
x = df1[[‘T – 28’, ‘T – 21’, ‘T – 14’, ‘T – 13’, ‘T – 12’, ‘T – 11’, ‘T – 10’, ‘T – 9’, ‘T – 8’]]
y = df1[[‘Actual’]]
Fitting Simple Linear regression data model
from sklearn.linear_model import LinearRegression
regressorObject=LinearRegression()
regressorObject.fit(x,y)
Predicting the number of surgeries
predict = df1.loc[2:242, ‘T – 28′:’T – 8’]
y_pred_test_data=regressorObject.predict(predict)
Predicted = pd.DataFrame(y_pred_test_data, columns = [‘Predicted’])
result3 = pd.concat([df, Predicted], ignore_index=True, axis=1)
result3.columns = [‘SurgDate’, ‘DOW’, ‘T – 28’, ‘T – 21’, ‘T – 14’, ‘T – 13’, ‘T – 12’, ‘T – 11′,’ T – 10′, ‘T – 9’,
‘T – 8’, ‘T – 7’, ‘T – 6’, ‘T – 5’, ‘T – 4’, ‘T – 3’, ‘T – 2’, ‘T – 1’, ‘Actual’, ‘Predicted’]
result3[‘Predicted’] = round(result3[‘Predicted’])
final3 = result3.dropna()
round(result[‘Actual’].mean()), round(final3[‘Predicted’].mean())
(116, 118)
We see that the Average Daily Error is ~2 Surgeries.
Let’s check the performance
y = final3[‘Actual’]
yhat = final3[‘Predicted’]
d = y – yhat
mse_f = np.mean(d2) mae_f = np.mean(abs(d)) rmse_f = np.sqrt(mse_f) r2_f = 1-(sum(d2)/sum((y-np.mean(y))**2))
print(“Results by manual calculation:”)
print(“MAE:”,mae_f)
print(“MSE:”, mse_f)
print(“RMSE:”, rmse_f)
print(“R-Squared:”, r2_f)
Results by manual calculation: MAE: 13.565957446808511 MSE: 375.6085106382979 RMSE: 19.380622039508893 R-Squared: -0.19119040826349143
Let’s plot actual versus predicted values
plt.figure(figsize=[15,8])
plt.grid(True)
plt.plot(result[‘Actual’],label=’Actual’)
plt.plot(result[‘Predicted’],label=’Predicted’)
plt.ylim(80, 150)
plt.legend(loc=2)
plt.savefig(‘predictedactualdata.png’)

And the % relative error is
plt.figure(figsize=[15,8])
plt.grid(True)
plt.plot(100*abs((result[‘Actual’]-result[‘Predicted’])/result[‘Actual’]))
plt.ylim([0, 40])
plt.savefig(‘predictrelerror.png’)

Summary
Our input data have 4 outliers (11/25/2011, 12/23/2011, 12/26/2011, and 12/30/2011).
Our data analysis shows that Fridays/Thursdays have the lowest/highest number of surgeris.
The surgery volume correlation value between DOW and the actual Surgery Date (SD) decreases as the gap between DOW and SD increases.
The ANOVE test rejects our null hypothesis H0 with 99% confidence.
Our results are statistically significant since p<<0.05.
We have performed multiple pairwise comparison (Tukey HSD) test to confirm the acceptance of our H1 hypothesis.
Predicting surgery volumes for 7 days before the Surgery Date yields the best result, as shown below:
| Metric | Base Model | 3 Days Forecast | 7 Days Forecast |
| MAE | 14.95 | 14.69 | 13.56 |
| MSE | 426.57 | 414.83 | 375.60 |
| RMSE | 20.65 | 20.36 | 19.38 |
| R2 | -0.35 | -0.31 | -0.19 |
This project supports earlier studies [1, 2] in that it underlines the importance of the implementation of automatic registration systems that integrate into the work processes in the OR to collect more and better data.
References
[1] Edelman et al, Front Med (Lausanne). 2017; 4: 85.
Published online 2017 Jun 19. doi: 10.3389/fmed.2017.00085
[2] Eun et al, Decision Sciences, 07 August 2020
[3] Rashi Desai, 2021, Vanderbilt University Medical Center: Effective Surgery Schedule. Github Repository.
[4] Cem ÖZÇELİK, 2022, DATA SCIENCE PROJECT: PREDICTING THE AMOUNT OF SURGERY, Published in MLearning.ai.

Leave a comment