After over two years of living with Covid-19, we are learning to adapt to a world with this disease.
2022 ends with looming risk of a new coronavirus variant, health experts warn.
In this post, we explore the statistics on the coronavirus pandemic for every country in the world. The goal is to compare the latest number of confirmed deaths and recovered people of COVID-19 cases Country/Region – Province/State wise. In addition, we invoke COVID-19 Vaccine Sentiment Analysis using Twitter Data for the data science research.
Table of Contents:
- EDA
- Bokeh Plots
- Plotly Impact Analysis
- Vaccine Sentiment Analysis
- Conclusions
- Explore More
- Embed Socials
- Infographic
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
EDA
Referring to the Squarespace Exploratory Data Analysis (EDA), let’s look at the latest number of confirmed deaths and recovered people of Novel Coronavirus cases Country wise.
Let’s import the key libraries and the input dataset
import pandas as pd
covid_data= pd.read_csv(‘https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/03-17-2020.csv’)
print(covid_data)
print(“\nDataset information:”)
print(covid_data.info())
print(“\nMissing data information:”)
print(covid_data.isna().sum())
Province/State Country/Region Last Update Confirmed Deaths \
0 Hubei China 2020-03-17T11:53:10 67799 3111
1 NaN Italy 2020-03-17T18:33:02 31506 2503
2 NaN Iran 2020-03-17T15:13:09 16169 988
3 NaN Spain 2020-03-17T20:53:02 11748 533
4 NaN Germany 2020-03-17T18:53:02 9257 24
.. ... ... ... ... ...
307 Wales United Kingdom 2020-03-17T11:53:10 0 5
308 NaN Nauru 2020-03-17T11:53:10 0 0
309 Niue New Zealand 2020-03-17T11:53:10 0 0
310 NaN Tuvalu 2020-03-17T11:53:10 0 0
311 Pitcairn Islands United Kingdom 2020-03-17T11:53:10 0 0
Recovered Latitude Longitude
0 56003 30.9756 112.2707
1 2941 41.8719 12.5674
2 5389 32.4279 53.6880
3 1028 40.4637 -3.7492
4 67 51.1657 10.4515
.. ... ... ...
307 0 52.1307 -3.7837
308 0 -0.5228 166.9315
309 0 -19.0544 -169.8672
310 0 -7.1095 177.6493
311 0 -24.3768 -128.3242
[312 rows x 8 columns]
Dataset information:
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 312 entries, 0 to 311
Data columns (total 8 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 Province/State 154 non-null object
1 Country/Region 312 non-null object
2 Last Update 312 non-null object
3 Confirmed 312 non-null int64
4 Deaths 312 non-null int64
5 Recovered 312 non-null int64
6 Latitude 309 non-null float64
7 Longitude 309 non-null float64
dtypes: float64(2), int64(3), object(3)
memory usage: 19.6+ KB
None
Missing data information:
Province/State 158
Country/Region 0
Last Update 0
Confirmed 0
Deaths 0
Recovered 0
Latitude 3
Longitude 3
dtype: int64
Let’s create the Active column
covid_data[‘Active’] = covid_data[‘Confirmed’] – covid_data[‘Deaths’] – covid_data[‘Recovered’]
result = covid_data.groupby(‘Country/Region’)[‘Confirmed’, ‘Deaths’, ‘Recovered’, ‘Active’].sum().reset_index()
print(result)
Country/Region Confirmed Deaths Recovered Active 0 Afghanistan 26 0 1 25 1 Albania 55 1 0 54 2 Algeria 60 4 12 44 3 Andorra 39 0 1 38 4 Antarctica 0 0 0 0 .. ... ... ... ... ... 163 Uzbekistan 10 0 0 10 164 Venezuela 33 0 0 33 165 Vietnam 66 0 16 50 166 Winter Olympics 2022 0 0 0 0 167 occupied Palestinian territory 0 0 0 0 [168 rows x 5 columns]
Let’s plot Deaths vs Confirmed cases
resultdeaths = result.sort_values(‘Deaths’, ascending=False)
resultdeaths10=resultdeaths
import pandas as pd
import plotly.express as px
state_fig = px.scatter(resultdeaths10, x=’Confirmed’, y=’Deaths’, title=’Top COVID-19 Deaths vs Confirmed’, text=’Deaths’,trendline=”ols”)
state_fig.show()

Let’s plot Active vs Recovered
resultdeaths = result.sort_values(‘Active’, ascending=False)
resultdeaths10=resultdeaths
import pandas as pd
import plotly.express as px
state_fig = px.scatter(resultdeaths10, x=’Recovered’, y=’Active’, title=’Top COVID-19 Active vs Recovered’, text=’Active’)
state_fig.show()

Let’s plot Deaths vs Country/Region
resultdeaths = covid_data.groupby(‘Country/Region’)[‘Deaths’].sum().reset_index().sort_values(‘Deaths’, ascending=False)
resultdeaths10=resultdeaths.head()
import pandas as pd
import plotly.express as px
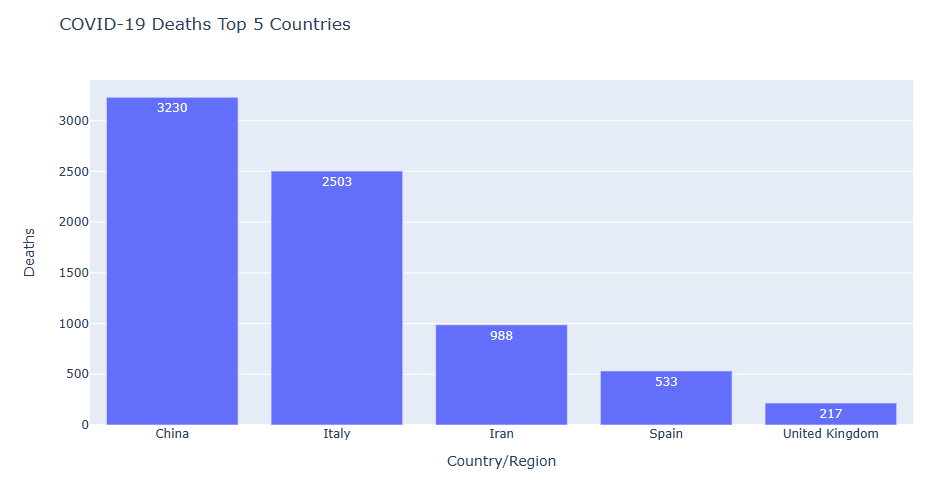
state_fig = px.bar(resultdeaths10, x=’Country/Region’, y=’Deaths’, title=’COVID-19 Deaths Top 5 Countries’, text=’Deaths’)
state_fig.show()
Let’s group our data
data = covid_data.groupby([‘Country/Region’, ‘Province/State’])[‘Confirmed’, ‘Deaths’, ‘Recovered’].max()
pd.set_option(‘display.max_rows’, None)
print(data)
Confirmed Deaths Recovered
Country/Region Province/State
Australia Australian Capital Territory 2 0 0
From Diamond Princess 0 0 0
New South Wales 210 4 4
Northern Territory 1 0 0
Queensland 78 0 8
South Australia 29 0 3
Tasmania 7 0 0
Victoria 94 0 8
Western Australia 31 1 0
Canada Alberta 74 0 0
British Columbia 103 4 4
Grand Princess 8 0 0
Manitoba 8 0 0
New Brunswick 8 0 0
Newfoundland and Labrador 3 0 0
Nova Scotia 7 0 0
Ontario 507 1 5
Prince Edward Island 1 0 0
Quebec 128 1 0
Saskatchewan 7 0 0
China Anhui 990 6 984
Beijing 456 8 369
Chongqing 576 6 570
Fujian 296 1 295
Gansu 133 2 91
Guangdong 1364 8 1307
Guangxi 253 2 248
Guizhou 147 2 144
Hainan 168 6 161
Hebei 318 6 310
Heilongjiang 482 13 456
Henan 1273 22 1250
Hong Kong 162 4 88
Hubei 67799 3111 56003
Hunan 1018 4 1014
Inner Mongolia 75 1 73
Jiangsu 631 0 631
Jiangxi 935 1 934
Jilin 93 1 92
Liaoning 125 1 120
Macau 12 0 10
Ningxia 75 0 75
Qinghai 18 0 18
Shaanxi 246 3 236
Shandong 761 7 746
Shanghai 358 3 325
Shanxi 133 0 133
Sichuan 540 3 520
Tianjin 136 3 133
Tibet 1 0 1
Unknown 0 0 0
Xinjiang 76 3 73
Yunnan 176 2 172
Zhejiang 1232 1 1216
Cruise Ship Diamond Princess 696 7 325
Denmark Denmark 977 4 1
Faroe Islands 47 0 0
France France 7652 148 12
French Guiana 7 0 0
French Polynesia 3 0 0
Guadeloupe 6 0 0
Mayotte 1 0 0
Reunion 9 0 0
Saint Barthelemy 3 0 0
St Martin 2 0 0
Malaysia Johor 0 0 0
Kedah 0 0 0
Kelantan 0 0 0
Melaka 0 0 0
Negeri Sembilan 0 0 0
Pahang 0 0 0
Perak 0 0 0
Perlis 0 0 0
Pulau Pinang 0 0 0
Sabah 0 0 0
Sarawak 0 0 0
Selangor 0 0 0
Terengganu 0 0 0
Unknown 0 0 0
W.P. Kuala Lumpur 0 0 0
W.P. Labuan 0 0 0
W.P. Putrajaya 0 0 0
Netherlands Curacao 3 0 0
Netherlands 1705 43 2
New Zealand Cook Islands 0 0 0
Niue 0 0 0
US Alabama 39 0 0
Alaska 3 0 0
Arizona 20 0 1
Arkansas 22 0 0
California 698 12 6
Colorado 160 2 0
Connecticut 68 0 0
Delaware 16 0 0
Diamond Princess 47 0 0
District of Columbia 22 0 0
Florida 216 6 0
Georgia 146 1 0
Grand Princess 21 0 0
Guam 3 0 0
Hawaii 10 0 0
Idaho 8 0 0
Illinois 161 1 2
Indiana 30 2 0
Iowa 23 0 0
Kansas 18 1 0
Kentucky 26 1 1
Louisiana 196 4 0
Maine 32 0 0
Maryland 60 0 3
Massachusetts 218 0 1
Michigan 65 0 0
Minnesota 60 0 0
Mississippi 21 0 0
Missouri 11 0 0
Montana 9 0 0
Nebraska 21 0 0
Nevada 56 1 0
New Hampshire 26 0 0
New Jersey 267 3 1
New Mexico 23 0 0
New York 1706 13 0
North Carolina 64 0 0
North Dakota 3 0 0
Ohio 67 0 0
Oklahoma 19 0 0
Oregon 66 1 0
Pennsylvania 112 0 0
Puerto Rico 5 0 0
Rhode Island 23 0 0
South Carolina 47 1 0
South Dakota 11 1 0
Tennessee 74 0 0
Texas 110 1 0
Utah 51 0 0
Vermont 12 0 0
Virgin Islands 2 0 0
Virginia 67 2 0
Washington 1076 55 1
West Virginia 1 0 0
Wisconsin 72 0 1
Wyoming 11 0 0
Ukraine Unknown 0 0 0
United Kingdom Cayman Islands 1 1 0
Channel Islands 0 0 0
England 0 198 0
Gibraltar 3 0 1
Guernsey 0 0 0
Jersey 0 0 0
Northern Ireland 0 0 0
Pitcairn Islands 0 0 0
Scotland 0 11 0
Unknown 1950 2 52
Wales 0 5 0
Let’s select China
c_data = covid_data[covid_data[‘Country/Region’]==’China’]
c_data = c_data[[‘Province/State’, ‘Confirmed’, ‘Deaths’, ‘Recovered’]]
result = c_data.sort_values(by=’Confirmed’, ascending=False)
result = result.reset_index(drop=True)
print(result)
Province/State Confirmed Deaths Recovered 0 Hubei 67799 3111 56003 1 Guangdong 1364 8 1307 2 Henan 1273 22 1250 3 Zhejiang 1232 1 1216 4 Hunan 1018 4 1014 5 Anhui 990 6 984 6 Jiangxi 935 1 934 7 Shandong 761 7 746 8 Jiangsu 631 0 631 9 Chongqing 576 6 570 10 Sichuan 540 3 520 11 Heilongjiang 482 13 456 12 Beijing 456 8 369 13 Shanghai 358 3 325 14 Hebei 318 6 310 15 Fujian 296 1 295 16 Guangxi 253 2 248 17 Shaanxi 246 3 236 18 Yunnan 176 2 172 19 Hainan 168 6 161 20 Hong Kong 162 4 88 21 Guizhou 147 2 144 22 Tianjin 136 3 133 23 Gansu 133 2 91 24 Shanxi 133 0 133 25 Liaoning 125 1 120 26 Jilin 93 1 92 27 Xinjiang 76 3 73 28 Inner Mongolia 75 1 73 29 Ningxia 75 0 75 30 Qinghai 18 0 18 31 Macau 12 0 10 32 Tibet 1 0 1 33 Unknown 0 0 0
Let’s plot Confirmed per Province
resultdeaths10=result
import pandas as pd
import plotly.express as px
state_fig = px.bar(resultdeaths10, x=’Province/State’, y=’Confirmed’, title=’COVID-19 Confirmed Top 10 Provinces’)
state_fig.show()

Let’s plot Deaths per Province
resultdeaths10=result
import pandas as pd
import plotly.express as px
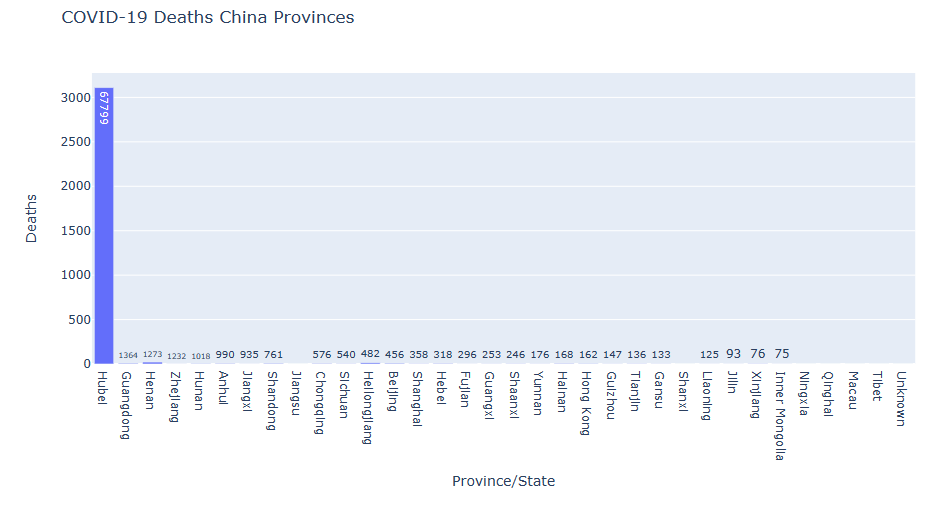
state_fig = px.bar(resultdeaths10, x=’Province/State’, y=’Deaths’, title=’COVID-19 Deaths China Provinces’)
state_fig.show()
Let’s plot Recovered per Province
resultdeaths10=result
import pandas as pd
import plotly.express as px
state_fig = px.bar(resultdeaths10, x=’Province/State’, y=’Recovered’, title=’COVID-19 Recovered China Provinces’)
state_fig.show()

Let’s select subset Deaths>0 vs Country/Region
data = covid_data.groupby(‘Country/Region’)[‘Confirmed’, ‘Deaths’, ‘Recovered’].sum().reset_index()
result = data[data[‘Deaths’]>0][[‘Country/Region’, ‘Deaths’]]
print(result)
Country/Region Deaths 1 Albania 1 2 Algeria 4 6 Argentina 2 9 Australia 5 10 Austria 3 11 Azerbaijan 1 12 Bahrain 1 16 Belgium 10 21 Brazil 1 23 Bulgaria 2 27 Canada 6 30 China 3230 37 Cruise Ship 7 41 Denmark 4 42 Dominican Republic 1 43 Ecuador 2 44 Egypt 4 49 Finland 4 50 France 148 54 Germany 24 56 Greece 5 60 Guatemala 1 63 Guyana 1 66 Hungary 1 67 Iceland 1 68 India 3 69 Indonesia 5 70 Iran 988 71 Iraq 11 72 Ireland 2 74 Italy 2503 76 Japan 29 83 Korea, South 81 87 Lebanon 3 91 Luxembourg 1 95 Martinique 1 103 Morocco 2 107 Netherlands 43 111 Norway 3 115 Panama 1 117 Peru 9 118 Philippines 12 119 Poland 5 120 Portugal 1 130 San Marino 9 137 Slovenia 1 140 Spain 533 142 Sudan 1 145 Sweden 7 146 Switzerland 40 147 Taiwan* 1 149 Thailand 1 156 Turkey 1 158 US 108 159 Ukraine 2 161 United Kingdom 217
Let’s select subset Recovered=0 vs Country/Region
data = covid_data.groupby(‘Country/Region’)[‘Confirmed’, ‘Deaths’, ‘Recovered’].sum().reset_index()
result = data[data[‘Recovered’]==0][[‘Country/Region’, ‘Confirmed’, ‘Deaths’, ‘Recovered’]]
print(result)
Country/Region Confirmed Deaths Recovered 1 Albania 55 1 0 4 Antarctica 0 0 0 5 Antigua and Barbuda 1 0 0 8 Aruba 3 0 0 14 Barbados 2 0 0 17 Benin 1 0 0 18 Bhutan 1 0 0 19 Bolivia 11 0 0 22 Brunei 56 0 0 23 Bulgaria 67 2 0 24 Burkina Faso 15 0 0 26 Cameroon 10 0 0 28 Central African Republic 1 0 0 29 Chile 201 0 0 32 Congo (Brazzaville) 1 0 0 33 Congo (Kinshasa) 3 0 0 34 Costa Rica 41 0 0 38 Cuba 5 0 0 39 Cyprus 40 0 0 42 Dominican Republic 21 1 0 43 Ecuador 58 2 0 45 Equatorial Guinea 1 0 0 47 Eswatini 1 0 0 48 Ethiopia 5 0 0 51 French Guiana 11 0 0 52 Gabon 1 0 0 55 Ghana 7 0 0 57 Greenland 1 0 0 58 Guadeloupe 18 0 0 59 Guam 3 0 0 60 Guatemala 6 1 0 61 Guernsey 0 0 0 62 Guinea 1 0 0 63 Guyana 7 1 0 64 Holy See 1 0 0 65 Honduras 8 0 0 67 Iceland 220 1 0 77 Jersey 0 0 0 79 Kazakhstan 33 0 0 80 Kenya 3 0 0 81 Kiribati 0 0 0 82 Korea, North 0 0 0 84 Kosovo 2 0 0 88 Liberia 5 0 0 89 Liechtenstein 19 0 0 91 Luxembourg 140 1 0 92 Malaysia 0 0 0 93 Maldives 13 0 0 95 Martinique 16 1 0 96 Mauritania 1 0 0 97 Mayotte 1 0 0 100 Monaco 7 0 0 101 Mongolia 4 0 0 102 Montenegro 2 0 0 104 Namibia 2 0 0 105 Nauru 0 0 0 108 New Zealand 12 0 0 109 Nigeria 3 0 0 114 Palau 0 0 0 115 Panama 69 1 0 116 Paraguay 11 0 0 121 Puerto Rico 0 0 0 123 Republic of the Congo 0 0 0 124 Reunion 9 0 0 127 Rwanda 7 0 0 128 Saint Lucia 2 0 0 129 Saint Vincent and the Grenadines 1 0 0 134 Seychelles 8 0 0 136 Slovakia 72 0 0 137 Slovenia 275 1 0 138 Somalia 1 0 0 139 South Africa 62 0 0 142 Sudan 1 1 0 143 Summer Olympics 2020 0 0 0 144 Suriname 1 0 0 148 Tanzania 1 0 0 150 The Bahamas 1 0 0 151 The Gambia 1 0 0 152 Togo 1 0 0 153 Tonga 0 0 0 154 Trinidad and Tobago 5 0 0 155 Tunisia 24 0 0 156 Turkey 47 1 0 157 Tuvalu 0 0 0 159 Ukraine 14 2 0 162 Uruguay 29 0 0 163 Uzbekistan 10 0 0 164 Venezuela 33 0 0 166 Winter Olympics 2022 0 0 0 167 occupied Palestinian territory 0 0 0
Let’s check the condition data[data[‘Confirmed’]==data[‘Deaths’]]
data = covid_data.groupby(‘Country/Region’)[‘Confirmed’, ‘Deaths’, ‘Recovered’].sum().reset_index()
result = data[data[‘Confirmed’]==data[‘Deaths’]]
result = result[[‘Country/Region’, ‘Confirmed’, ‘Deaths’]]
result = result.sort_values(‘Confirmed’, ascending=False)
result = result[result[‘Confirmed’]>0]
result = result.reset_index(drop=True)
print(result)
Country/Region Confirmed Deaths 0 Sudan 1 1
Let’s check the condition data[data[‘Confirmed’]==data[‘Recovered’]]
data = covid_data.groupby(‘Country/Region’)[‘Confirmed’, ‘Deaths’, ‘Recovered’].sum().reset_index()
result = data[data[‘Confirmed’]==data[‘Recovered’]]
result = result[[‘Country/Region’, ‘Confirmed’, ‘Recovered’]]
result = result.sort_values(‘Confirmed’, ascending=False)
result = result[result[‘Confirmed’]>0]
result = result.reset_index(drop=True)
print(result)
Country/Region Confirmed Recovered 0 Nepal 1 1
Let’s select top 10 Confirmed countries
result = covid_data.groupby(‘Country/Region’).max().sort_values(by=’Confirmed’, ascending=False)[:10]
pd.set_option(‘display.max_column’, None)
print(result)
Last Update Confirmed Deaths Recovered Latitude \
Country/Region
China 2020-03-17T12:13:13 67799 3111 56003 47.8620
Italy 2020-03-17T18:33:02 31506 2503 2941 41.8719
Iran 2020-03-17T15:13:09 16169 988 5389 32.4279
Spain 2020-03-17T20:53:02 11748 533 1028 40.4637
Germany 2020-03-17T18:53:02 9257 24 67 51.1657
Korea, South 2020-03-17T10:33:03 8320 81 1407 35.9078
France 2020-03-17T19:13:08 7652 148 12 46.2276
Switzerland 2020-03-17T16:33:04 2700 40 4 46.8182
United Kingdom 2020-03-17T15:13:09 1950 198 52 56.4907
US 2020-03-17T23:53:03 1706 55 6 61.3707
Longitude Active
Country/Region
China 127.7615 8685
Italy 12.5674 26062
Iran 53.6880 9792
Spain -3.7492 10187
Germany 10.4515 9166
Korea, South 127.7669 6832
France 55.2471 7492
Switzerland 8.2275 2656
United Kingdom -1.1743 1896
US 144.7937 1693
Let’s plot Total Deaths(>150), Confirmed, Recovered and Active Cases by Country (top 10 countries)
import pandas as pd
import matplotlib.pyplot as plt
covid_data= pd.read_csv(‘https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/03-19-2020.csv’, usecols = [‘Last Update’, ‘Country/Region’, ‘Confirmed’, ‘Deaths’, ‘Recovered’])
covid_data[‘Active’] = covid_data[‘Confirmed’] – covid_data[‘Deaths’] – covid_data[‘Recovered’]
r_data = covid_data.groupby([“Country/Region”])[“Deaths”, “Confirmed”, “Recovered”, “Active”].sum().reset_index()
r_data = r_data.sort_values(by=’Deaths’, ascending=False)
r_data = r_data[r_data[‘Deaths’]>50]
plt.figure(figsize=(15, 5))
plt.plot(r_data[‘Country/Region’], r_data[‘Deaths’],color=’red’)
plt.plot(r_data[‘Country/Region’], r_data[‘Confirmed’],color=’green’)
plt.plot(r_data[‘Country/Region’], r_data[‘Recovered’], color=’blue’)
plt.plot(r_data[‘Country/Region’], r_data[‘Active’], color=’black’)
plt.title(‘Total Deaths(>150), Confirmed, Recovered and Active Cases by Country’)
plt.show()

Let’s select US and plot Deaths per State
import pandas as pd
import plotly.express as px
covid_data= pd.read_csv(‘https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/03-17-2020.csv’)
us_data = covid_data[covid_data[‘Country/Region’]==’US’].drop([‘Country/Region’,’Latitude’, ‘Longitude’], axis=1)
us_data = us_data[us_data.sum(axis = 1) > 0]
us_data = us_data.groupby([‘Province/State’])[‘Deaths’].sum().reset_index()
us_data_death = us_data[us_data[‘Deaths’] > 0]
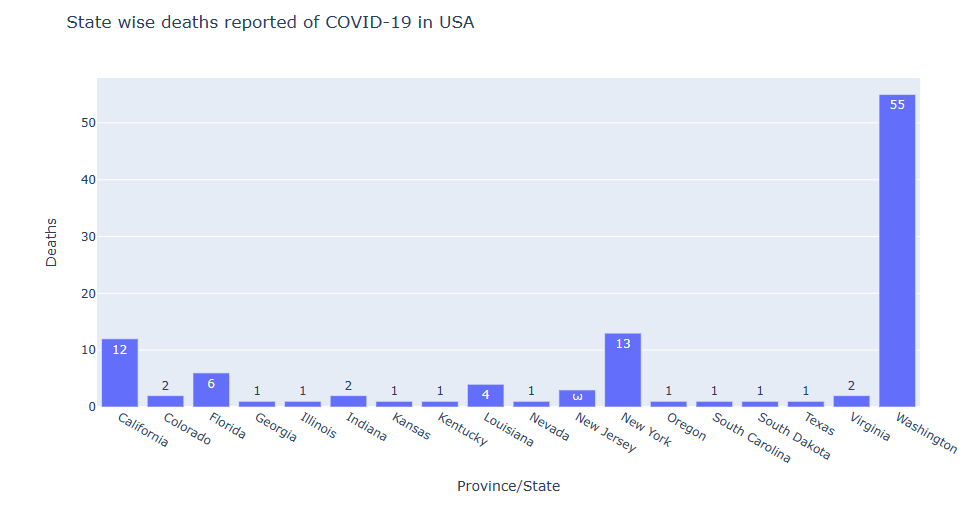
state_fig = px.bar(us_data_death, x=’Province/State’, y=’Deaths’, title=’State wise deaths reported of COVID-19 in USA’, text=’Deaths’)
state_fig.show()
Let’s plot US states Recovery
import pandas as pd
import plotly.express as px
covid_data= pd.read_csv(‘https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/03-18-2020.csv’)
covid_data[‘Active’] = covid_data[‘Confirmed’] – covid_data[‘Deaths’] – covid_data[‘Recovered’]
us_data = covid_data[covid_data[‘Country/Region’]==’US’].drop([‘Country/Region’,’Latitude’, ‘Longitude’], axis=1)
us_data = us_data[us_data.sum(axis = 1) > 0]
us_data = us_data.groupby([‘Province/State’])[‘Active’].sum().reset_index()
us_data_death = us_data[us_data[‘Active’] > 0]
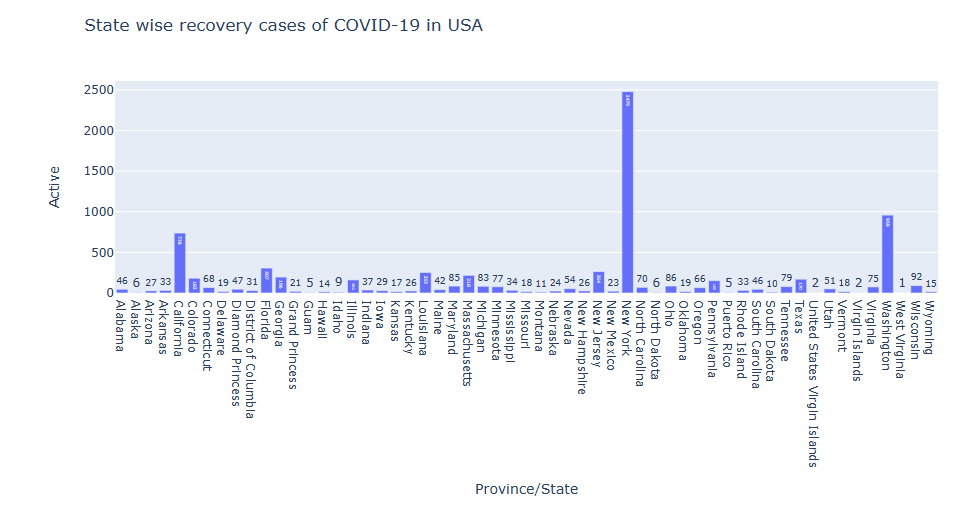
state_fig = px.bar(us_data_death, x=’Province/State’, y=’Active’, title=’State wise recovery cases of COVID-19 in USA’, text=’Active’)
state_fig.show()
Let’s plot Confirmed, Deaths, Active, and Recovered for US states
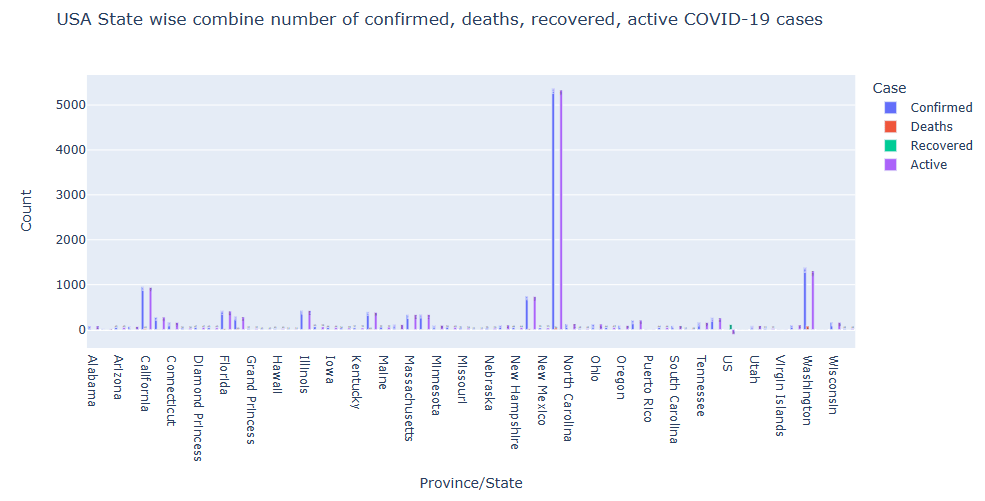
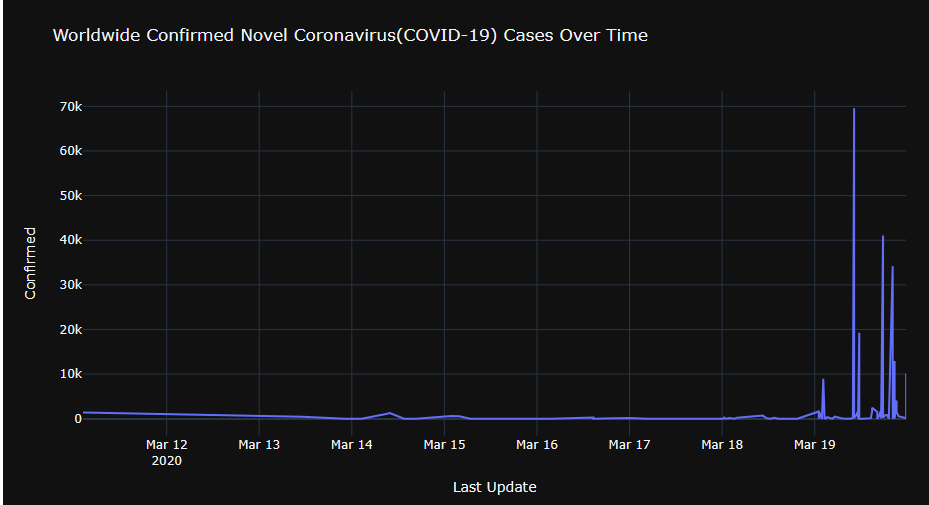
Let’s plot Worldwide Confirmed Novel Coronavirus(COVID-19) Cases Over Time
import pandas as pd
import plotly.express as px
import plotly.io as pio
pio.templates.default = “plotly_dark”
covid_data= pd.read_csv(‘https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/03-19-2020.csv’)
grouped = covid_data.groupby(‘Last Update’)[‘Last Update’, ‘Confirmed’, ‘Deaths’].sum().reset_index()
fig = px.line(grouped, x=”Last Update”, y=”Confirmed”,
title=”Worldwide Confirmed Novel Coronavirus(COVID-19) Cases Over Time”)
fig.show()
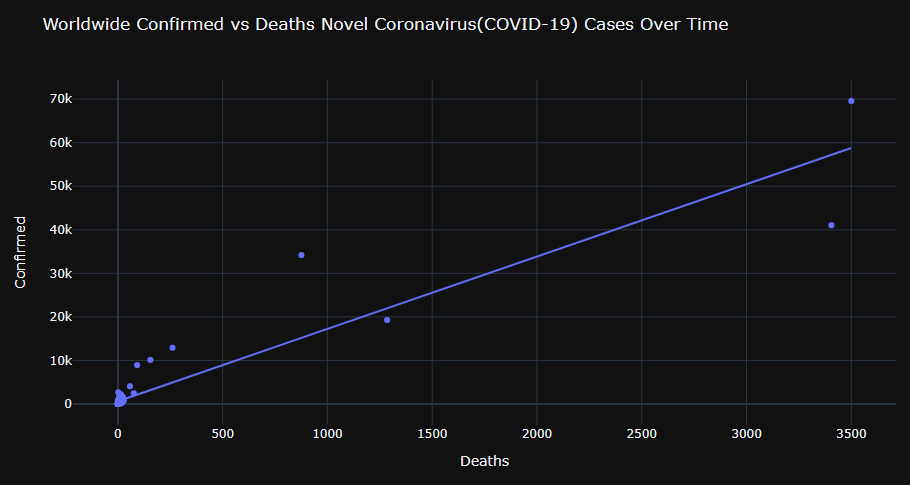
Let’s look at the scatter plot Worldwide Confirmed vs Deaths Novel Coronavirus(COVID-19) Cases Over Time (with the linear trend)
fig = px.scatter(grouped, x=”Deaths”, y=”Confirmed”,
title=”Worldwide Confirmed vs Deaths Novel Coronavirus(COVID-19) Cases Over Time”,trendline=”ols”)
fig.show()
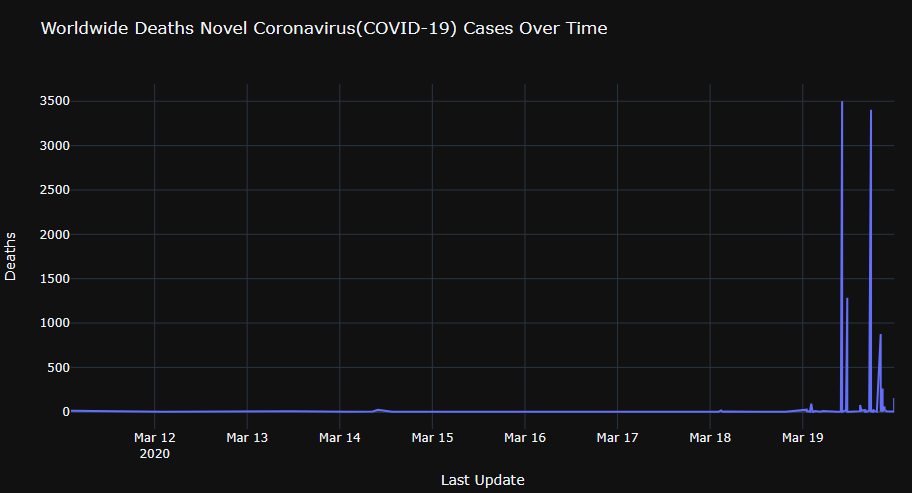
Let’s plot Worldwide Deaths Novel Coronavirus(COVID-19) Cases Over Time
fig = px.line(grouped, x=”Last Update”, y=”Deaths”,
title=”Worldwide Deaths Novel Coronavirus(COVID-19) Cases Over Time”)
fig.show()
Bokeh Plots
Let’s invoke the Bokeh library to see how Asia is doing against COVID-19
We begin with importing the key libraries
import gc
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from bokeh.plotting import figure
from bokeh.models import LabelSet, ColumnDataSource
from bokeh.io import show, output_notebook
from bokeh.io import output_notebook
from bokeh.resources import INLINE
from bokeh.layouts import row, column
output_notebook(INLINE)
import warnings
warnings.filterwarnings(‘ignore’)
Loading BokehJS …
Importing the dataset
df = pd.read_csv(‘AsiaCases_.csv’)
df.head().style.background_gradient(cmap=’RdGy’)

Checking for any missing values
df.isnull().sum()
ID 0 Country 0 TotalCases 0 TotalDeaths 1 TotalRecovered 0 ActiveCases 0 TotalCasesPerMillion 0 TotalDeathsPerMillion 1 TotalTests 1 TotalTestsPerMillion 1 TotalPopulation 0 dtype: int64
Dropping the countries with missing data
df.dropna(axis=0, inplace=True)
and checking the descriptive statistics
df.describe().T

df.info()
<class 'pandas.core.frame.DataFrame'> Int64Index: 47 entries, 0 to 48 Data columns (total 11 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 ID 47 non-null int64 1 Country 47 non-null object 2 TotalCases 47 non-null int64 3 TotalDeaths 47 non-null float64 4 TotalRecovered 47 non-null int64 5 ActiveCases 47 non-null int64 6 TotalCasesPerMillion 47 non-null int64 7 TotalDeathsPerMillion 47 non-null float64 8 TotalTests 47 non-null float64 9 TotalTestsPerMillion 47 non-null float64 10 TotalPopulation 47 non-null int64 dtypes: float64(4), int64(6), object(1) memory usage: 4.4+ KB
Checking the number of rows and columns
print(‘Number of rows:’, df.shape[0])
print(‘Number of columns:’, df.shape[1])
Number of rows: 47 Number of columns: 11
Let’s drop the ID column
df.drop(columns=’ID’, axis=1, inplace=True)
Let’s plot top 10 countries with the highest number of cases
df_high_cases = df[[‘Country’,’TotalCases’,’TotalPopulation’]].sort_values(by=’TotalCases’, ascending=False)
df_high_cases = df_high_cases.iloc[:10,:]
df_high_cases[‘perc’] = round((df_high_cases[‘TotalCases’] / df_high_cases[‘TotalPopulation’]) * 100, 1)
df_high_cases[‘perc’] = df_high_cases[‘perc’].apply(str)
df_high_cases[‘perc’] = df_high_cases[‘perc’]+’%’
Our x and y axis
country = list(df_high_cases[‘Country’].values)
pop = list(df_high_cases[‘TotalPopulation’].values)
case = list(df_high_cases[‘TotalCases’].values)
perc = list(df_high_cases[‘perc’].values)
For the Cases
p1 = figure(x_range=country, y_range=[0,40000000], height=500, width=1000, title=”Top 10 Countries in terms of Covid-19 Cases”)
p1.background_fill_color = “#efefef”
p1.vbar(x=country, top=case, width=0.9, color=’#db0000′,
alpha=[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.1],
line_color=”black”,line_width=3)
source = ColumnDataSource(data=dict(y=case,
x=country,
names=perc,
text_font_size=[’20px’],
text_alpha=[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.2]))
labels = LabelSet(x=’x’, y=’y’, text=’names’, source=source, text_font_size=’text_font_size’, text_alpha=’text_alpha’, x_offset=-18)
p1.ygrid.grid_line_color = None
p1.xgrid.grid_line_color = None
p1.title.text_font_size = ’18pt’
p1.add_layout(labels, place=’center’)
p1.xaxis.major_label_text_font_size = “12pt”
Let’s call show(column(p1, p2))
gc.collect()
show(p1)

Let’s plot top 10 countries with highest number of deaths
df_high_death = df[[‘Country’,’TotalDeaths’,’TotalPopulation’]].sort_values(by=’TotalDeaths’, ascending=False)
df_high_death = df_high_death.iloc[:10,:]
df_high_death[‘perc’] = round((df_high_death[‘TotalDeaths’] / df_high_death[‘TotalPopulation’]) * 100, 3)
df_high_death[‘perc’] = df_high_death[‘perc’].apply(str)
df_high_death[‘perc’] = df_high_death[‘perc’]+’%’
Our x and y axis
country = list(df_high_death[‘Country’].values)
pop = list(df_high_cases[‘TotalPopulation’].values)
case = list(df_high_death[‘TotalDeaths’].values)
perc = list(df_high_death[‘perc’].values)
For the Cases
p1 = figure(x_range=country, y_range=[0,500000], height=500, width=1000, title=”Top 10 Countries in terms of Covid-19 Deaths”)
p1.background_fill_color = “#efefef”
p1.vbar(x=country, top=case, width=0.9, color=’#db0000′,
alpha=[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.1],
line_color=”black”,line_width=3)
source = ColumnDataSource(data=dict(y=case,
x=country,
names=perc,
text_font_size=[’20px’],
text_alpha=[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.2]))
labels = LabelSet(x=’x’, y=’y’, text=’names’, source=source, text_font_size=’text_font_size’, text_alpha=’text_alpha’, x_offset=-25)
p1.ygrid.grid_line_color = None
p1.xgrid.grid_line_color = None
p1.title.text_font_size = ’18pt’
p1.add_layout(labels, place=’center’)
p1.xaxis.major_label_text_font_size = “12pt”
gc.collect()
show(p1)

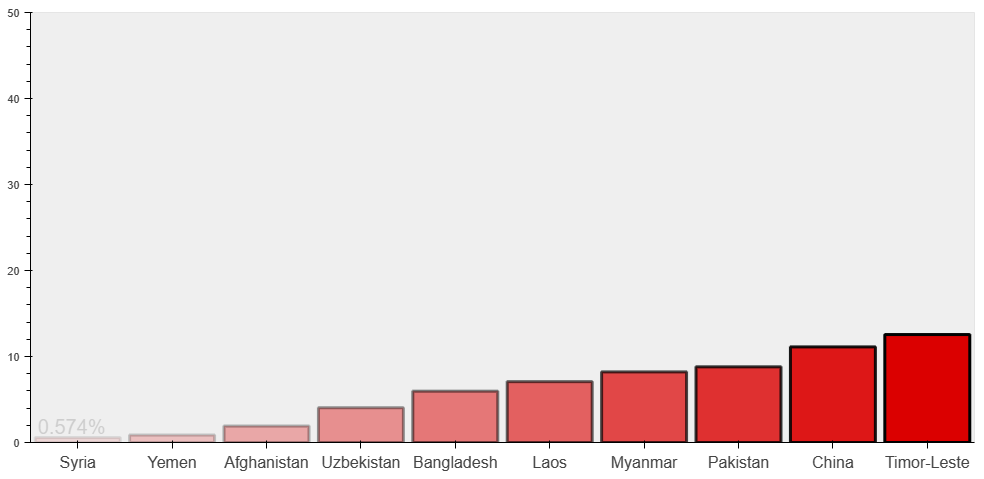
Let’s plot top 10 Asian countries with the lowest number of tests
df_low_test = df[[‘Country’,’TotalTests’,’TotalPopulation’]]
df_low_test[‘perc’] = round((df_low_test[‘TotalTests’] / df_low_test[‘TotalPopulation’]) * 100, 3)
df_low_test = df_low_test.sort_values(by=’perc’, ascending=True).iloc[:10,:]
df_low_test[‘perc_’] = df_low_test[‘perc’].apply(str)
df_low_test[‘perc_’] = df_low_test[‘perc_’]+’%’
Our x and y axis
country = list(df_low_test[‘Country’].values)
pop = list(df_high_cases[‘TotalPopulation’].values)
case = list(df_low_test[‘TotalTests’].values)
perc = list(df_low_test[‘perc’].values)
perc_ = list(df_low_test[‘perc_’].values)
For the Cases
p1 = figure(x_range=country, y_range=[0,50], height=500, width=1000, title=”Bottom 10 Countries in terms of Covid-19 Test/Population”)
p1.background_fill_color = “#efefef”
p1.vbar(x=country, top=perc, width=0.9, color=’#db0000′,
alpha=[0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1],
line_color=”black”,line_width=3)
source = ColumnDataSource(data=dict(y=perc,
x=country,
names=perc_,
text_font_size=[’20px’],
text_alpha=[0.2,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1]))
labels = LabelSet(x=’x’, y=’y’, text=’names’, source=source, text_font_size=’text_font_size’, text_alpha=’text_alpha’, x_offset=-40)
p1.ygrid.grid_line_color = None
p1.xgrid.grid_line_color = None
p1.title.text_font_size = ’18pt’
p1.add_layout(labels, place=’center’)
p1.xaxis.major_label_text_font_size = “12pt”
gc.collect()
show(p1)
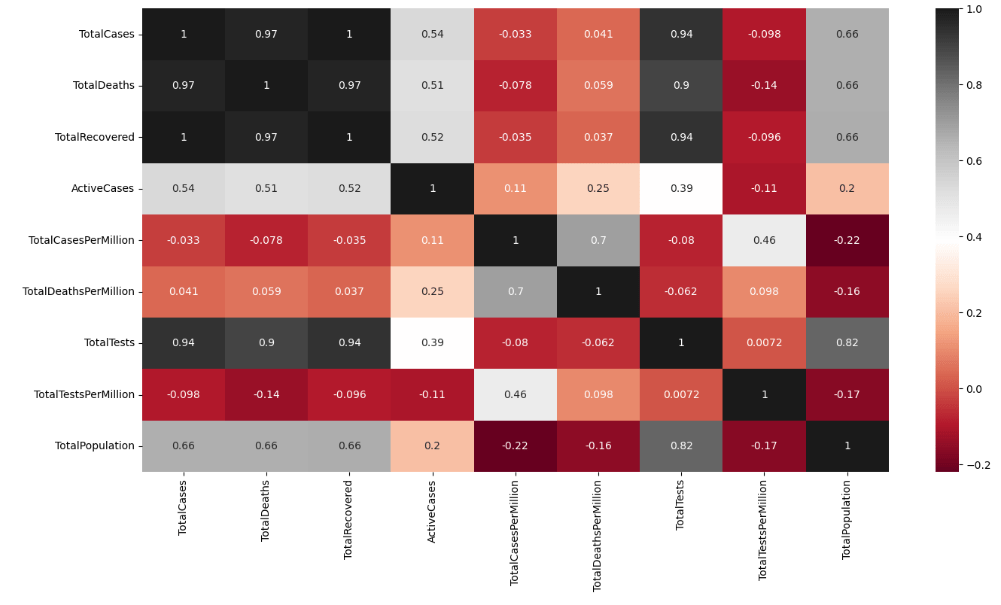
Checking if there are any data correlations
heat = df.corr()
plt.figure(figsize=[16,8])
sns.heatmap(heat, annot=True, cmap=’RdGy’)
gc.collect()
plt.show()
Let’s plot Total Deaths to Recovery ratio
df_ded_reco = df[[‘Country’,’TotalDeaths’,’TotalRecovered’]]
df_ded_reco[‘DeadRecovRatio’] = round(df_ded_reco[‘TotalDeaths’]/df_ded_reco[‘TotalRecovered’], 4)
df_ded_reco = df_ded_reco.sort_values(by=’DeadRecovRatio’, ascending=False).iloc[:10,:]
Our x and y axis
country = list(df_ded_reco[‘Country’].values)
case = list(df_ded_reco[‘DeadRecovRatio’].values)
For the Cases
p1 = figure(x_range=country, y_range=[0,0.5], height=500, width=1000, title=”Top 10 Countries in terms of Death/Recovery Ratio”)
p1.background_fill_color = “#efefef”
p1.vbar(x=country, top=case, width=0.9, color=’#db0000′,
alpha=[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.2],
line_color=”black”,line_width=3)
p1.ygrid.grid_line_color = None
p1.xgrid.grid_line_color = None
p1.title.text_font_size = ’18pt’
gc.collect()
show(p1)

Plotly Impact Analysis
Let’s implement the COVID-19 impact analysis using Plotly.
Let’s set the working directory YOURPATH, import key libraries and input COVID-19 data
import os
os.chdir(‘C:/Users/adrou/OneDrive/Documents/COVIDAMAN’) # Set working directory
os. getcwd()
import pandas as pd
import plotly.express as px
import plotly.graph_objects as go
data = pd.read_csv(“transformed_data.csv”)
data2 = pd.read_csv(“raw_data.csv”)
print(data)
CODE COUNTRY DATE HDI TC TD STI \
0 AFG Afghanistan 2019-12-31 0.498 0.000000 0.000000 0.000000
1 AFG Afghanistan 2020-01-01 0.498 0.000000 0.000000 0.000000
2 AFG Afghanistan 2020-01-02 0.498 0.000000 0.000000 0.000000
3 AFG Afghanistan 2020-01-03 0.498 0.000000 0.000000 0.000000
4 AFG Afghanistan 2020-01-04 0.498 0.000000 0.000000 0.000000
... ... ... ... ... ... ... ...
50413 ZWE Zimbabwe 2020-10-15 0.535 8.994048 5.442418 4.341855
50414 ZWE Zimbabwe 2020-10-16 0.535 8.996528 5.442418 4.341855
50415 ZWE Zimbabwe 2020-10-17 0.535 8.999496 5.442418 4.341855
50416 ZWE Zimbabwe 2020-10-18 0.535 9.000853 5.442418 4.341855
50417 ZWE Zimbabwe 2020-10-19 0.535 9.005405 5.442418 4.341855
POP GDPCAP
0 17.477233 7.497754
1 17.477233 7.497754
2 17.477233 7.497754
3 17.477233 7.497754
4 17.477233 7.497754
... ... ...
50413 16.514381 7.549491
50414 16.514381 7.549491
50415 16.514381 7.549491
50416 16.514381 7.549491
50417 16.514381 7.549491
[50418 rows x 9 columns]
Let’s check value_counts() per country
data[“COUNTRY”].value_counts()
Afghanistan 294
Indonesia 294
Macedonia 294
Luxembourg 294
Lithuania 294
...
Tajikistan 172
Comoros 171
Lesotho 158
Hong Kong 51
Solomon Islands 4
Name: COUNTRY, Length: 210, dtype: int64
The total number of countries is
data[“COUNTRY”].value_counts().mode()
0 294 Name: COUNTRY, dtype: int64
Aggregating the data
code = data[“CODE”].unique().tolist()
country = data[“COUNTRY”].unique().tolist()
hdi = []
tc = []
td = []
sti = []
population = data[“POP”].unique().tolist()
gdp = []
for i in country:
hdi.append((data.loc[data[“COUNTRY”] == i, “HDI”]).sum()/294)
tc.append((data2.loc[data2[“location”] == i, “total_cases”]).sum())
td.append((data2.loc[data2[“location”] == i, “total_deaths”]).sum())
sti.append((data.loc[data[“COUNTRY”] == i, “STI”]).sum()/294)
population.append((data2.loc[data2[“location”] == i, “population”]).sum()/294)
aggregated_data = pd.DataFrame(list(zip(code, country, hdi, tc, td, sti, population)),
columns = [“Country Code”, “Country”, “HDI”,
“Total Cases”, “Total Deaths”,
“Stringency Index”, “Population”])
print(aggregated_data.head())
Country Code Country HDI Total Cases Total Deaths \ 0 AFG Afghanistan 0.498000 5126433.0 165875.0 1 ALB Albania 0.600765 1071951.0 31056.0 2 DZA Algeria 0.754000 4893999.0 206429.0 3 AND Andorra 0.659551 223576.0 9850.0 4 AGO Angola 0.418952 304005.0 11820.0 Stringency Index Population 0 3.049673 17.477233 1 3.005624 14.872537 2 3.195168 17.596309 3 2.677654 11.254996 4 2.965560 17.307957
Sorting Data According to Total Cases
data = aggregated_data.sort_values(by=[“Total Cases”], ascending=False)
print(data.head())
Country Code Country HDI Total Cases Total Deaths \
200 USA United States 0.92400 746014098.0 26477574.0
27 BRA Brazil 0.75900 425704517.0 14340567.0
90 IND India 0.64000 407771615.0 7247327.0
157 RUS Russia 0.81600 132888951.0 2131571.0
150 PER Peru 0.59949 74882695.0 3020038.0
Stringency Index Population
200 3.350949 19.617637
27 3.136028 19.174732
90 3.610552 21.045353
157 3.380088 18.798668
150 3.430126 17.311165
Let’s select top 10 Countries with Highest Covid Cases
data = data.head(10)
print(data)
Country Code Country HDI Total Cases Total Deaths \
200 USA United States 0.924000 746014098.0 26477574.0
27 BRA Brazil 0.759000 425704517.0 14340567.0
90 IND India 0.640000 407771615.0 7247327.0
157 RUS Russia 0.816000 132888951.0 2131571.0
150 PER Peru 0.599490 74882695.0 3020038.0
125 MEX Mexico 0.774000 74347548.0 7295850.0
178 ESP Spain 0.887969 73717676.0 5510624.0
175 ZAF South Africa 0.608653 63027659.0 1357682.0
42 COL Colombia 0.581847 60543682.0 1936134.0
199 GBR United Kingdom 0.922000 59475032.0 7249573.0
Stringency Index Population
200 3.350949 19.617637
27 3.136028 19.174732
90 3.610552 21.045353
157 3.380088 18.798668
150 3.430126 17.311165
125 3.019289 18.674802
178 3.393922 17.660427
175 3.364333 17.898266
42 3.357923 17.745037
199 3.353883 18.033340
Let’s compare country GDP before/during COVID
data[“GDP Before Covid”] = [65279.53, 8897.49, 2100.75,
11497.65, 7027.61, 9946.03,
29564.74, 6001.40, 6424.98, 42354.41]
data[“GDP During Covid”] = [63543.58, 6796.84, 1900.71,
10126.72, 6126.87, 8346.70,
27057.16, 5090.72, 5332.77, 40284.64]
print(data)
Country Code Country HDI Total Cases Total Deaths \
200 USA United States 0.924000 746014098.0 26477574.0
27 BRA Brazil 0.759000 425704517.0 14340567.0
90 IND India 0.640000 407771615.0 7247327.0
157 RUS Russia 0.816000 132888951.0 2131571.0
150 PER Peru 0.599490 74882695.0 3020038.0
125 MEX Mexico 0.774000 74347548.0 7295850.0
178 ESP Spain 0.887969 73717676.0 5510624.0
175 ZAF South Africa 0.608653 63027659.0 1357682.0
42 COL Colombia 0.581847 60543682.0 1936134.0
199 GBR United Kingdom 0.922000 59475032.0 7249573.0
Stringency Index Population GDP Before Covid GDP During Covid
200 3.350949 19.617637 65279.53 63543.58
27 3.136028 19.174732 8897.49 6796.84
90 3.610552 21.045353 2100.75 1900.71
157 3.380088 18.798668 11497.65 10126.72
150 3.430126 17.311165 7027.61 6126.87
125 3.019289 18.674802 9946.03 8346.70
178 3.393922 17.660427 29564.74 27057.16
175 3.364333 17.898266 6001.40 5090.72
42 3.357923 17.745037 6424.98 5332.77
199 3.353883 18.033340 42354.41 40284.64
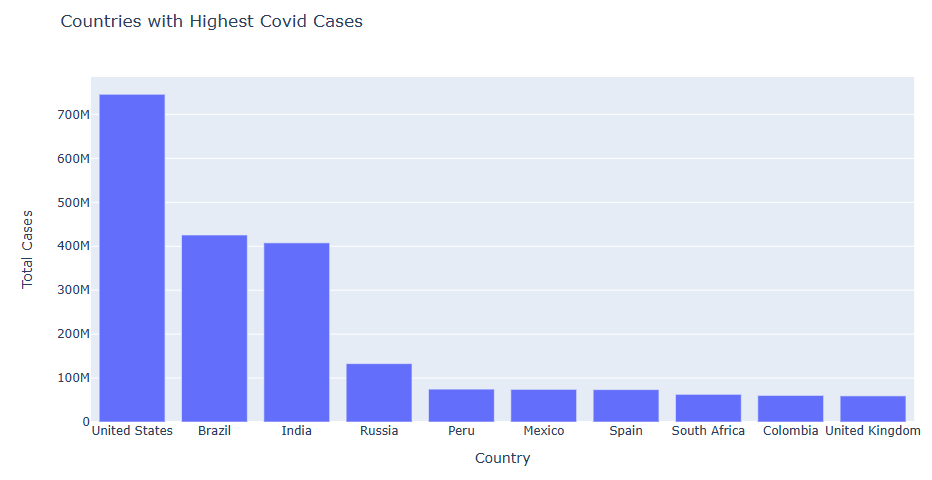
Let’s plot top 10 Total Cases vs Countries
import plotly.express as px
figure = px.bar(data, y=’Total Cases’, x=’Country’,
title=”Countries with Highest Covid Cases”)
figure.show()
Let’s plot top 10 Countries with Highest Deaths
figure = px.bar(data, y=’Total Deaths’, x=’Country’,
title=”Countries with Highest Deaths”)
figure.show()

Let’s compare Total Cases vs Total Deaths for top 10 countries
fig = go.Figure()
fig.add_trace(go.Bar(
x=data[“Country”],
y=data[“Total Cases”],
name=’Total Cases’,
marker_color=’indianred’
))
fig.add_trace(go.Bar(
x=data[“Country”],
y=data[“Total Deaths”],
name=’Total Deaths’,
marker_color=’lightsalmon’
))
fig.update_layout(barmode=’group’, xaxis_tickangle=-45)
fig.show()

let’s plot the Percentage of Total Cases and Deaths
cases = data[“Total Cases”].sum()
deceased = data[“Total Deaths”].sum()
labels = [“Total Cases”, “Total Deaths”]
values = [cases, deceased]
fig = px.pie(data, values=values, names=labels,
title=’Percentage of Total Cases and Deaths’, hole=0.5)
fig.show()

The death rate is given by
death_rate = (data[“Total Deaths”].sum() / data[“Total Cases”].sum()) * 100
print(“Death Rate = “, death_rate)
Death Rate = 3.6144212045653767
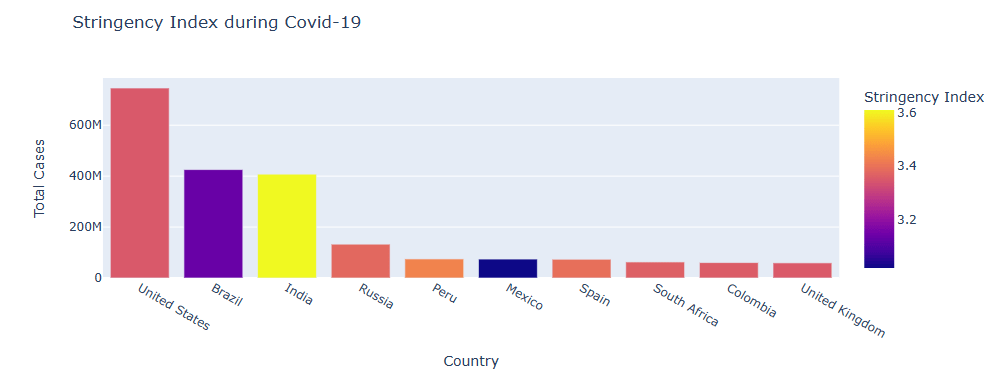
Let’s plot the Stringency Index during COVID-19 for top 10 countries
fig = px.bar(data, x=’Country’, y=’Total Cases’,
hover_data=[‘Population’, ‘Total Deaths’],
color=’Stringency Index’, height=400,
title= “Stringency Index during Covid-19”)
fig.show()
Let’s compare GDP Per Capita Before Covid-19
fig = px.bar(data, x=’Country’, y=’Total Cases’,
hover_data=[‘Population’, ‘Total Deaths’],
color=’GDP Before Covid’, height=400,
title=”GDP Per Capita Before Covid-19″)
fig.show()

Let’s compare GDP Per Capita During Covid-19
fig = px.bar(data, x=’Country’, y=’Total Cases’,
hover_data=[‘Population’, ‘Total Deaths’],
color=’GDP During Covid’, height=400,
title=”GDP Per Capita During Covid-19″)
fig.show()

Let’s compare GDP per Capita before/during COVID-19
fig = go.Figure()
fig.add_trace(go.Bar(
x=data[“Country”],
y=data[“GDP Before Covid”],
name=’GDP Per Capita Before Covid-19′,
marker_color=’indianred’
))
fig.add_trace(go.Bar(
x=data[“Country”],
y=data[“GDP During Covid”],
name=’GDP Per Capita During Covid-19′,
marker_color=’lightsalmon’
))
fig.update_layout(barmode=’group’, xaxis_tickangle=-45)
fig.show()

Let’s plot Human Development Index during Covid-19
fig = px.bar(data, x=’Country’, y=’Total Cases’,
hover_data=[‘Population’, ‘Total Deaths’],
color=’HDI’, height=400,
title=”Human Development Index during Covid-19″)
fig.show()

Vaccine Sentiment Analysis
Finally, let’s turn our attention to the COVID-19 vaccine sentiment analysis.
Let’s set the working directory YOURPATH
import os
os.chdir(‘YOURPATH’)
os. getcwd()
Let’s install and import the key NLP libraries
!pip install nltk
!pip install wordcloud
!pip install statsmodels
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import re
import string
import nltk
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style(‘darkgrid’)
import plotly.express as ex
import plotly.graph_objs as go
import plotly.offline as pyo
from plotly.subplots import make_subplots
pyo.init_notebook_mode()
nltk.download(‘vader_lexicon’)
from nltk.sentiment.vader import SentimentIntensityAnalyzer as SIA
from wordcloud import WordCloud,STOPWORDS
from pandas.plotting import autocorrelation_plot
from statsmodels.graphics.tsaplots import plot_acf
from statsmodels.graphics.tsaplots import plot_pacf
from statsmodels.tsa.seasonal import seasonal_decompose
from nltk.util import ngrams
from nltk import word_tokenize
from nltk.stem import PorterStemmer
from nltk.stem import WordNetLemmatizer
import random
plt.rc(‘figure’,figsize=(17,13))
Let’s read and edit the input dataset
f_data = pd.read_csv(‘vaccination_tweets.csv’)
f_data.text =f_data.text.str.lower()
Remove twitter handlers
f_data.text = f_data.text.apply(lambda x:re.sub(‘@[^\s]+’,”,x))
Remove hashtags
f_data.text = f_data.text.apply(lambda x:re.sub(r’\B#\S+’,”,x))
Remove URLs
f_data.text = f_data.text.apply(lambda x:re.sub(r”http\S+”, “”, x))
Remove all the special characters
f_data.text = f_data.text.apply(lambda x:’ ‘.join(re.findall(r’\w+’, x)))
Remove all single characters
f_data.text = f_data.text.apply(lambda x:re.sub(r’\s+[a-zA-Z]\s+’, ”, x))
Substituting multiple spaces with single space
f_data.text = f_data.text.apply(lambda x:re.sub(r’\s+’, ‘ ‘, x, flags=re.I))
Let’s invoke SentimentIntensityAnalyzer
sid = SIA()
f_data[‘sentiments’] = f_data[‘text’].apply(lambda x: sid.polarity_scores(‘ ‘.join(re.findall(r’\w+’,x.lower()))))
f_data[‘Positive Sentiment’] = f_data[‘sentiments’].apply(lambda x: x[‘pos’]+1(10-6)) f_data[‘Neutral Sentiment’] = f_data[‘sentiments’].apply(lambda x: x[‘neu’]+1(10-6))
f_data[‘Negative Sentiment’] = f_data[‘sentiments’].apply(lambda x: x[‘neg’]+1(10*-6))
f_data.drop(columns=[‘sentiments’],inplace=True)
Let’s get the f_data info
f_data.shape
(11020, 19)
f_data.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 11020 entries, 0 to 11019 Data columns (total 19 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 id 11020 non-null int64 1 user_name 11020 non-null object 2 user_location 8750 non-null object 3 user_description 10341 non-null object 4 user_created 11020 non-null object 5 user_followers 11020 non-null int64 6 user_friends 11020 non-null int64 7 user_favourites 11020 non-null int64 8 user_verified 11020 non-null bool 9 date 11020 non-null object 10 text 11020 non-null object 11 hashtags 8438 non-null object 12 source 11019 non-null object 13 retweets 11020 non-null int64 14 favorites 11020 non-null int64 15 is_retweet 11020 non-null bool 16 Positive Sentiment 11020 non-null float64 17 Neutral Sentiment 11020 non-null float64 18 Negative Sentiment 11020 non-null float64 dtypes: bool(2), float64(3), int64(6), object(8) memory usage: 1.5+ MB
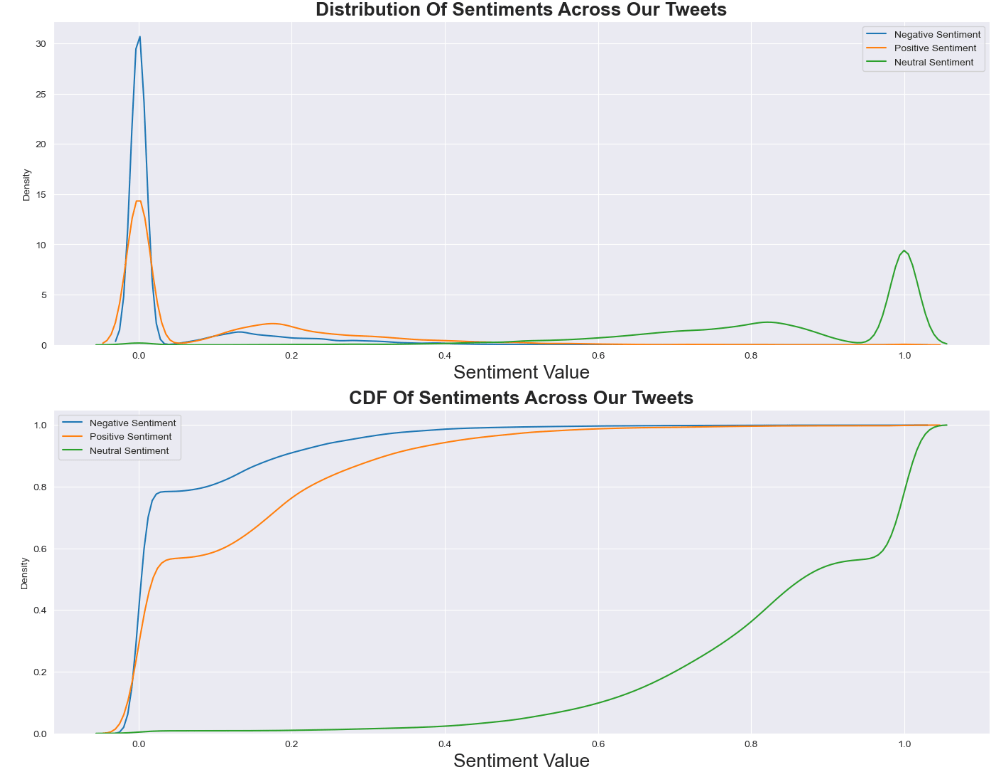
Let’s plot the Distribution and CDF Of Sentiments Across Our Tweets
plt.subplot(2,1,1)
plt.title(‘Distribution Of Sentiments Across Our Tweets’,fontsize=19,fontweight=’bold’)
sns.kdeplot(f_data[‘Negative Sentiment’],bw=0.1)
sns.kdeplot(f_data[‘Positive Sentiment’],bw=0.1)
sns.kdeplot(f_data[‘Neutral Sentiment’],bw=0.1)
plt.legend(labels=[‘Negative Sentiment’, ‘Positive Sentiment’, ‘Neutral Sentiment’])
plt.xlabel(‘Sentiment Value’,fontsize=19)
plt.subplot(2,1,2)
plt.title(‘CDF Of Sentiments Across Our Tweets’,fontsize=19,fontweight=’bold’)
sns.kdeplot(f_data[‘Negative Sentiment’],bw=0.1,cumulative=True)
sns.kdeplot(f_data[‘Positive Sentiment’],bw=0.1,cumulative=True)
sns.kdeplot(f_data[‘Neutral Sentiment’],bw=0.1,cumulative=True)
plt.legend(labels=[‘Negative Sentiment’, ‘Positive Sentiment’, ‘Neutral Sentiment’])
plt.xlabel(‘Sentiment Value’,fontsize=19)
plt.show()
Data Sorting, Feature Engineering and Selecting A Cut-Off For Most Positive/Negative Tweets
f_data = f_data.sort_values(by=’date’)
ft_data=f_data.copy()
ft_data[‘date’] = pd.to_datetime(f_data[‘date’]).dt.date
ft_data[‘year’] = pd.DatetimeIndex(ft_data[‘date’]).year
ft_data[‘month’] = pd.DatetimeIndex(ft_data[‘date’]).month
ft_data[‘day’] = pd.DatetimeIndex(ft_data[‘date’]).day
ft_data[‘day_of_year’] = pd.DatetimeIndex(ft_data[‘date’]).dayofyear
ft_data[‘quarter’] = pd.DatetimeIndex(ft_data[‘date’]).quarter
ft_data[‘season’] = ft_data.month%12 // 3 + 1
plt.subplot(2,1,1)
plt.title(‘Selecting A Cut-Off For Most Positive/Negative Tweets’,fontsize=19,fontweight=’bold’)
ax0 = sns.kdeplot(f_data[‘Negative Sentiment’],bw=0.1)
kde_x, kde_y = ax0.lines[0].get_data()
ax0.fill_between(kde_x, kde_y, where=(kde_x>0.25) ,
interpolate=True, color=’b’)
plt.annotate(‘Cut-Off For Most Negative Tweets’, xy=(0.25, 0.5), xytext=(0.4, 2),
arrowprops=dict(facecolor=’red’, shrink=0.05),fontsize=16,fontweight=’bold’)
ax0.axvline(f_data[‘Negative Sentiment’].mean(), color=’r’, linestyle=’–‘)
ax0.axvline(f_data[‘Negative Sentiment’].median(), color=’tab:orange’, linestyle=’-‘)
plt.legend({‘PDF’:f_data[‘Negative Sentiment’],r’Mean: {:.2f}’.format(f_data[‘Negative Sentiment’].mean()):f_data[‘Negative Sentiment’].mean(),
r’Median: {:.2f}’.format(f_data[‘Negative Sentiment’].median()):f_data[‘Negative Sentiment’].median()})
plt.subplot(2,1,2)
ax1 = sns.kdeplot(f_data[‘Positive Sentiment’],bw=0.1,color=’green’)
plt.annotate(‘Cut-Off For Most Positive Tweets’, xy=(0.4, 0.43), xytext=(0.4, 2),
arrowprops=dict(facecolor=’red’, shrink=0.05),fontsize=16,fontweight=’bold’)
kde_x, kde_y = ax1.lines[0].get_data()
ax1.fill_between(kde_x, kde_y, where=(kde_x>0.4) ,
interpolate=True, color=’green’)
ax1.set_xlabel(‘Sentiment Strength’,fontsize=18)
ax1.axvline(f_data[‘Positive Sentiment’].mean(), color=’r’, linestyle=’–‘)
ax1.axvline(f_data[‘Positive Sentiment’].median(), color=’tab:orange’, linestyle=’-‘)
plt.legend({‘PDF’:f_data[‘Positive Sentiment’],r’Mean: {:.2f}’.format(f_data[‘Positive Sentiment’].mean()):f_data[‘Positive Sentiment’].mean(),
r’Median: {:.2f}’.format(f_data[‘Positive Sentiment’].median()):f_data[‘Positive Sentiment’].median()})
plt.show()

Let’s look at the Common Words Among Most Positive/Negative Tweets
Most_Positive = f_data[f_data[‘Positive Sentiment’].between(0.4,1)]
Most_Negative = f_data[f_data[‘Negative Sentiment’].between(0.25,1)]
Most_Positive_text = ‘ ‘.join(Most_Positive.text)
Most_Negative_text = ‘ ‘.join(Most_Negative.text)
pwc = WordCloud(width=600,height=400,collocations = False).generate(Most_Positive_text)
nwc = WordCloud(width=600,height=400,collocations = False).generate(Most_Negative_text)
plt.subplot(1,2,1)
plt.title(‘Common Words Among Most Positive Tweets’,fontsize=16,fontweight=’bold’)
plt.imshow(pwc)
plt.axis(‘off’)
plt.subplot(1,2,2)
plt.title(‘Common Words Among Most Negative Tweets’,fontsize=16,fontweight=’bold’)
plt.imshow(nwc)
plt.axis(‘off’)
plt.show()
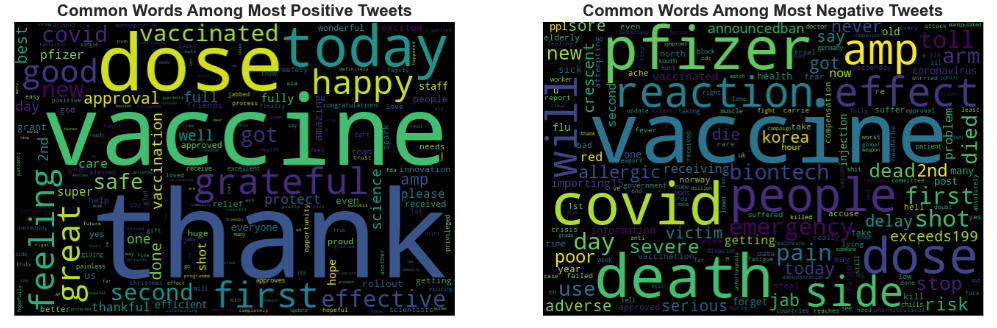
Let’s compare Top 10 Words In Most Positive/Negative Tweets
l_t = Most_Positive_text
w1_dict = dict()
for word in l_t.split():
w= word.strip()
if w in STOPWORDS:
continue
else:
w1_dict[w] = w1_dict.get(w,0)+1
w1_dict = {k: v for k, v in sorted(w1_dict.items(), key=lambda item: item[1],reverse=True)}
l_t = Most_Negative_text
w2_dict = dict()
for word in l_t.split():
w= word.strip()
if w in STOPWORDS:
continue
else:
w2_dict[w] = w2_dict.get(w,0)+1
w2_dict = {k: v for k, v in sorted(w2_dict.items(), key=lambda item: item[1],reverse=True)}
top_10_pos = list(w1_dict.keys())[:10]
top_10_neg = list(w2_dict.keys())[:10]
plt.subplot(1,2,1)
w_c = WordCloud(width=600,height=400,collocations = False,colormap=’nipy_spectral’).generate(‘ ‘.join(top_10_pos))
plt.title(‘Top 10 Words In Most Positive Tweets’,fontsize=19,fontweight=’bold’)
plt.imshow(w_c)
plt.axis(‘off’)
plt.subplot(1,2,2)
w_c = WordCloud(width=600,height=400,collocations = False,colormap=’nipy_spectral’).generate(‘ ‘.join(top_10_neg))
plt.title(‘Top 10 Words In Most Negative Tweets’,fontsize=19,fontweight=’bold’)
plt.imshow(w_c)
plt.axis(‘off’)
plt.show()

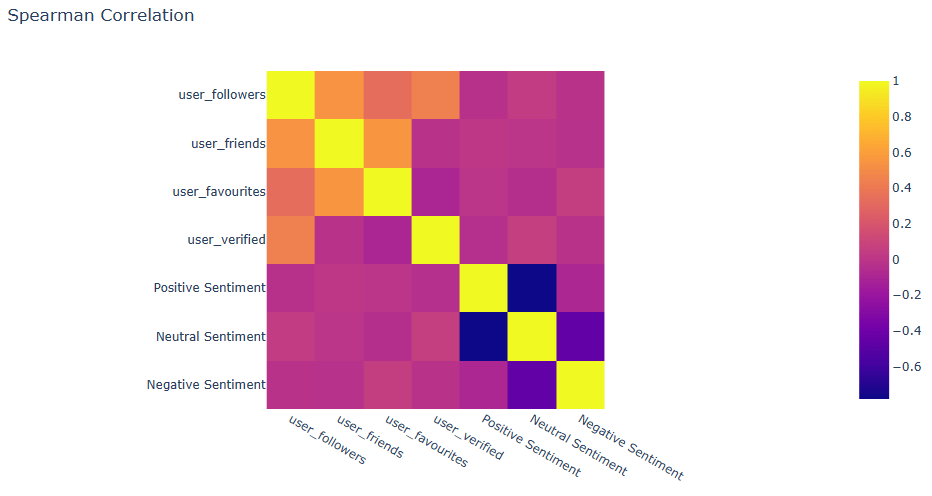
Let’s examine the Spearman correlation matrix
ex.imshow(f_data[[‘user_followers’,’user_friends’,’user_favourites’,’user_verified’,’Positive Sentiment’,
‘Neutral Sentiment’,’Negative Sentiment’]].corr(‘spearman’),title=’Spearman Correlation’)
Conclusions
- We implemented and tested a Python program to get the latest number of confirmed deaths and recovered people of Novel Coronavirus (COVID-19) cases Country/Region – Province/State wise.
- We analyzed the impact of COVID-19 on the global economy using the Kaggle dataset.
- We performed the COVID-19 vaccine sentiment analysis using the Twitter dataset.
Explore More
COVID-19 Geospatial Data Visualization with Plotly, Geopandas, and Folium
50 Coronavirus COVID-19 Free APIs
Interactive Global COVID-19 Data Visualization with Plotly
Comparing 4 Python Libraries for Interactive COVID-19 Data Science Visualization
Embed Socials
https://youtube.com/shorts/F5vSYgIwtAY
Infographic
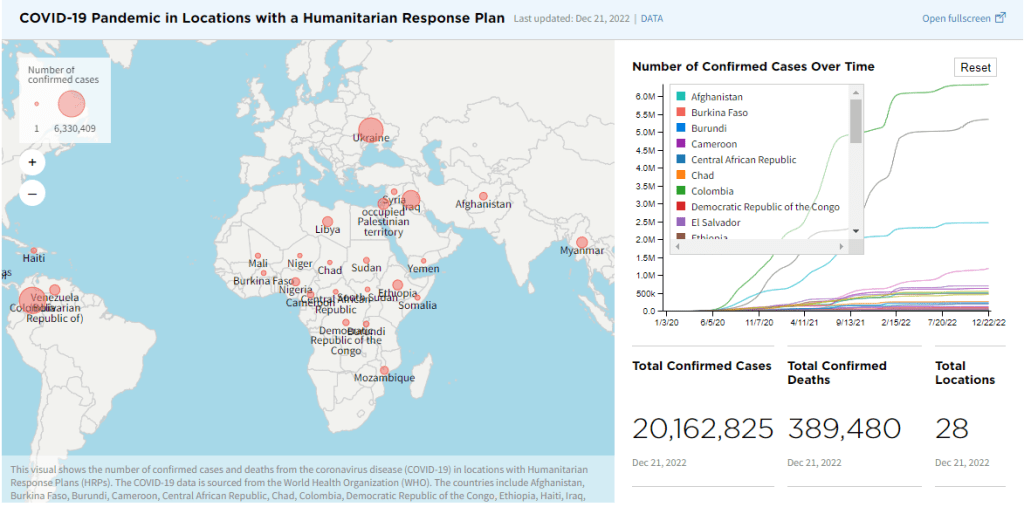

Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.


Leave a comment